Abstract
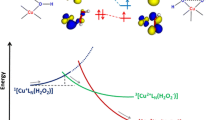
The octarepeat region of the prion protein can bind Cu2+ ions up to full occupancy (one ion per octarepeat) at neutral pH. While crystallographic data show that the HGGG octarepeat subdomain is the basic binding unit, multiple histidine coordination at lower Cu occupancy has been reported by X-ray absorption spectroscopy, EPR, and potentiometric experiments. In this paper we investigate, with first principles Car–Parrinello simulations, the first step for the formation of the Cu low-level binding mode, where four histidine side chains are coordinated to the same Cu2+ ion. This step involves the further binding of a second histidine to an already HGGG domain bonded Cu2+ ion. The influence of the pH on the ability of Cu to bind two histidine side chains was taken into account by simulating different protonation states of the amide N atoms of the two glycines lying nearest to the first histidine. Multiple histidine coordination is also seen to occur when glycine deprotonation occurs and the presence of the extra histidine stabilizes the Cu–peptide complex. Though the stabilization effect slightly decreases with the number of deprotonated glycines (reaching a minimum when both N atoms of the two nearest glycines are available as Cu ligands), the system is still capable of binding the second histidine in a 4N tetrahedral (though slightly distorted) coordination, whose energy is very near to that of the crystallographic square-planar 3N1O coordination. This result suggests that at low metal concentration the reorganization energy associated with Cu(II)/Cu(I) reduction is small also at pH ~ 7, when glycines are deprotonated.





Similar content being viewed by others
Notes
Actually the Makov–Payne correction also applies to systems with a nonvanishing dipole moment, irrespective of their charge. For the systems of interest here, this bit of the correction is, however, very small.
References
Prusiner SB (1982) Science 216:136–144
Prusiner SB (1998) Nobel Lect 74–129
Prusiner SB (1998) Proc Natl Acad Sci USA 95:13363–13383
Gasset M, Baldwin MA, Fletterick RJ, Prusiner SB (1993) Proc Natl Acad Sci USA 90:1–5
Pan KM, Baldwin M, Nguyen J, Gasset M, Serban A, Groth D, Mehlhorn I, Huang Z, Fletterick RJ, Cohen FE, Prusiner SB (1993) Proc Natl Acad Sci USA 90:10962–10966
Dominikus AL, Schorn C, Nivon LG, Esteve-Moya V, Christen B, Calzolai L, von Schroetter C, Fiorito F, Herrmann T, Güntert P, Wüthrich K (2005) Proc Natl Acad Sci USA 102:640–645
McKinley MP, Meyer RK, Kenaga L, Rahbar F, Serban RCA, Prusiner SB (1991) J Virol 65:1340–1351
Chiesa R, Drisaldi B, Quaglio E, Migheli A, Piccardo P, Ghetti B, Harris DA (2000) Proc Natl Acad Sci USA 97:5574–5579
Nunziante M, Gilch S, Schatzl HM (2003) J Biol Chem 278:3726–3734
Eghiaian F, Grosclaude J, Lesceu S, Debey P, Doublet B, Tréguer E, Rezaei H, Knossow M (2004) Proc Natl Acad Sci USA 101:10254–10259
Stockel J, Safar J, Wallace AC, Cohen FE, Prusiner SB (1998) Biochemistry 37:7185–7193
Miura T, Hori-i A, Mototani H, Takeuchi H (1999) Biochemistry 38:11560–11569
Jackson GS, Murray I, Hosszu LLP, Gibbs N, Waltho JP, Clarke AR, Colling J (2001) Proc Natl Acad Sci USA 98:8531–8535
Rachidi W, Mange A, Senator A, Guiraud P, Riondel J, Benboubetra M, Favier A, Lehmann S (2003) J Biol Chem 278:14595–14598
Gaggelli E, Kozlowski H, Valensin D, Valensin G (2006) Chem Rev 106:1995–2044
Hornshaw MP, McDermott JR, Candy JM, Lakey JH (1995) Biochem Biophys Res Commun 214:993–999
Whittal RM, Ball HL, Cohen FE, Burlingame AL, Prusiner SB, Baldwin MA (2000) Protein Sci 9:332–343
Wells MA, Jelinska C, Hosszu LLP, Craven CJ, Clarke AR, Collinge J, Waltho J, Jackson G (2006) Biochem J 400:501–510
Burns CS, Aronoff-Spencer E, Dunham CM, Lario P, Avdievich NI, Antholine WE, Olmstead MM, Vrielink A, Gerfen GJ, Peisach J, Scott WG, Millhauser GL (2002) Biochemistry 41:3991–4001
Qin K, Yang Y, Mastrangelo P, Westaway D (2002) J Biol Chem 277:1981–1990
Viles JH, Cohen FE, Prusiner SB, Goodin DB, Wright PE, Dyson HJ (1999) Proc Natl Acad Sci USA 96:2042–2047
Kramer ML, Kratzin HD, Schmidt B, Windl ARO, Liemann S, Hornemann S, Kretzschmar H (2001) J Biol Chem 276:16711–16719
Garnett AP, Viles JH (2003) J Biol Chem 278:6795–6802
Kozlowski H, Janicka-Klos A, Stanczak P, Valensin D, Valensin G, Kulon K (2008) Coord Chem Rev 252:1069–1078
Hasnain SS, Murphy LM, Grossmann RWSJG, Clarke AR, Jackson GS, Collinge J (2001) J Mol Biol 311:467–473
Cereghetti GM, Schweiger A, Glockshuber R, van Doorslaer S (2001) J Biophys 81:516–525
Burns CS, Aronoff-Spencer E, Legname G, Prusiner SB, Antholine WE, Gerfen GJ, Peisach J, Millhauser GL (2003) Biochemistry 42:6794–6803
Morante S, Gonzalez-Iglesias R, Potrich C, Meneghini C, Meyer-Klaucke W, Menestrina G, Gasset M (2004) J Biol Chem 279:11753–11759
Chattopadhyay M, Walter ED, Newell DJ, Jackson PJ, Aronoff-Spencer E, Peisach J, Gerfen GJ, Bennett B, Antholine WE, Millhauser GL (2005) J Am Chem Soc 127:12647–12656
Valensin D, Luczkowski FM, Mancini A, Gaggelli ELE, Valensin G, Rolka K, Kozlowski H (2004) Dalton Trans 1284–1293
Stanczak P, Kozlowski H (2007) Biochem Biophys Res Comm 352:198–202
Pradines V, Stroia AJ, Faller P (2008) New J Chem 32:1189
Sigel H, Martin RB (1982) Chem Rev 82:385–426
Furlan S, La Penna G, Guerrieri F, Morante S, Rossi GC (2007) J Biol Inorg Chem 12:571–583
Baroni S, Dal Corso A, de Gironcoli S, Giannozzi P, Cavazzoni C, Ballabio G, Scandolo S, Chiarotti G, Focher P, Pasquarello A, Laasonen K, Trave A, Car R, Marzari N, Kokalj A (2008) Quantum ESPRESSO. http://www.quantum-espresso.org
Vanderbilt D (1990) Phys Rev B 41:7892–7895
Perdew JP, Burke K, Ernzerhof M (1996) Phys Rev Lett 77:3865–3868
Nosé S (1984) Mol Phys 52:255–268
Frenkel D, Smit B (1996) Understanding molecular simulation. Academic Press, San Diego
Giannozzi P, De Angelis F, Car R (2004) J Chem Phys 120:5903–5915
La Penna G (2003) J Chem Phys 119:8162–8174
Muegge I, Qui PX, Wand AJ, Chu ZT, Warshel A (1997) J Phys Chem B 101:825–836
Acknowledgments
We thank P. Giannozzi and C. Cavazzoni for invaluable suggestions and constant help. The computations were performed on Linux clusters at the E. Fermi Research Center (Rome, Italy), at the BEN facility of the ECT* Institute (Trento, Italy), at Cineca (Bologna, Italy), and on locally assembled hardware. Financial support from Firb 2003 projects RBNE03PX83, PRIN05, and PRIN06 is acknowledged.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Guerrieri, F., Minicozzi, V., Morante, S. et al. Modeling the interplay of glycine protonation and multiple histidine binding of copper in the prion protein octarepeat subdomains. J Biol Inorg Chem 14, 361–374 (2009). https://doi.org/10.1007/s00775-008-0454-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-008-0454-8