Abstract
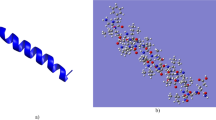
Antimicrobial peptides (AMPs) constitute part of a broad range of bioactive compounds present on diverse organisms, including frogs. Peptides, produced in the granular glands of amphibian skin, constitute a component of their innate immune response, providing protection against pathogenic microorganisms. In this work, two novel cruzioseptins peptides, cruzioseptin-16 and -17, extracted from the splendid leaf frog Cruziohyla calcarifer are presented. These peptides were identified using molecular cloning and tandem mass spectrometry. Later, peptides were synthetized using solid-phase peptide synthesis, and their minimal inhibitory concentration and haemolytic activity were tested. Furthermore, these two cruzioseptins plus three previously reported (CZS-1, CZS-2, CZS-3) were computationally characterized. Results show that cruzioseptins are 21–23 residues long alpha helical cationic peptides, with antimicrobial activity against E. coli, S. aureus, and C. albicans and low haemolytic effect. Docking results agree with the principal action mechanism of cationic AMPs that goes through cell membrane disruption due to electrostatic interactions between cationic residues in the cruzioseptins and negative phosphate groups in the pathogen cell membrane. An action mechanism through enzymes inhibition was also tried, but no conclusive results about this mechanism were obtained.












Similar content being viewed by others
References
Ali M, Jamil A, Alosaimi M et al (2020) Disease-a-Month Antimicrobial resistance, mechanisms and its clinical significance. Dis Mon. https://doi.org/10.1016/j.disamonth.2020.100971
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Bachem. Peptide calculator. https://www.bachem.com/service-support/peptide-calculator/. Accessed 14 Jan 2020
Bahar AA, Ren D (2013) Antimicrobial peptides. Pharmaceuticals 6:1543–1575
Berman HM, Battistuz T, Bhat TN et al (2002) The protein data bank. Acta Crystallogr Sect D Biol Crystallogr 28:235–242. https://doi.org/10.1107/S0907444902003451
Bio-Synthesis (2018) Peptide property calculator. https://www.biosyn.com/peptidepropertycalculatorlanding.aspx. Accessed 14 Jan 2020
Burton MG, Huang QM, Hossain MA et al (2013) Long-time-scale interaction dynamics between a model antimicrobial peptide and giant unilamellar vesicles. Langmuir 29:14613–14621. https://doi.org/10.1021/la403083m
Che Q, Zhou Y, Yang H et al (2008) A novel antimicrobial peptide from amphibian skin secretions of Odorrana grahami. Peptides 29:529–535. https://doi.org/10.1016/j.peptides.2008.01.004
Chen T, Bjourson AJ, Orr DF, Kwok H, Rao P, Ivanyi C, Shaw C (2002) Unmasking venom gland transcriptomes in reptile venoms. Anal Biochem 311:152–156
Chen T, Farragher S, Bjourson AJ, Orr DF, Rao P, Shaw C (2003) Granular gland transcriptomes in stimulated amphibian skin secretions. Biochem J 371:125–130
Ciumac D, Gong H, Hu X, Lu JR (2019) Membrane targeting cationic antimicrobial peptides. J Colloid Interface Sci 537:163–185
Conlon JM (2011) Structural diversity and species distribution of host-defense peptides in frog skin secretions. Cell Mol Life Sci 68:2303–2315
Conlon JM (2012) The potential of frog skin antimicrobial peptides for development into therapeutically valuable anti-infective agents. In: ACS symposium series. American Chemical Society, pp 47–60
Conlon JM, Mechkarska M, Lukic ML, Flatt PR (2014) Potential therapeutic applications of multifunctional host-defense peptides from frog skin as anti-cancer, anti-viral, immunomodulatory, and anti-diabetic agents. Peptides 57:67–77
Cuesta S, Gallegos F, Arias J et al (2019) Molecular modeling of four Dermaseptin-related peptides of the gliding tree frog Agalychnis spurrelli. J Mol Model. https://doi.org/10.1007/s00894-019-4141-1
Dathe M, Meyer J, Beyermann M et al (2002) General aspects of peptide selectivity towards lipid bilayers and cell membranes studied by variation of the structural parameters of amphipathic helical model peptides. Biochim Biophys Acta Biomembr 1558:171–186. https://doi.org/10.1016/S0005-2736(01)00429-1
Demori I, El RZ, Corradino V et al (2019) Peptides for skin protection and healing in amphibians. Molecules. https://doi.org/10.3390/molecules24020347
Di Tommaso P, Moretti S, Xenarios I et al (2011) T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res 39:W13–W17. https://doi.org/10.1093/nar/gkr245
Drozdetskiy A, Cole C, Procter J, Barton GJ (2015) JPred4: a protein secondary structure prediction server. Nucleic Acids Res 43:W389–W394. https://doi.org/10.1093/nar/gkv332
Eisenberg D, Weiss RM, Terwilliger TC (1982) The helical hydrophobic moment: a measure of the amphiphilicity of a helix. Nature 299:371–374. https://doi.org/10.1038/299371a0
Fauchere J-L, Pliska V (1983) Hydrophobic parameters pi of amino-acid side chains from the partitioning of N-acetyl-amino-acid amides. Eur J Med Chem 18:369–375
Fieser TM, Tainer JA, Geysen HM et al (1987) Influence of protein flexibility and peptide conformation on reactivity of monoclonal anti-peptide antibodies with a protein alpha-helix. Proc Natl Acad Sci U S A 84:8568–8572. https://doi.org/10.1073/pnas.84.23.8568
Frieri M, Kumar K, Boutin A (2017) Antibiotic resistance. J Infect Public Health 10:369–378. https://doi.org/10.1016/j.jiph.2016.08.007
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenber DJ (2009) Gaussian 09. Gaussian, Inc., Wallingford
Frost DR (2019) Amphibian species of the world: an online reference. Version 6.0. American Museum of Natural History, New York, USA. http://research.amnh.org/herpetology/amphibia/index.html
García MG, Rodríguez A, Alba A et al (2020) New antibacterial peptides from the freshwater mollusk Pomacea poeyana (Pilsbry, 1927). Biomolecules 10:1–17. https://doi.org/10.3390/biom10111473
Gautier R, Douguet D, Antonny B, Drin G (2008) HELIQUEST: a web server to screen sequences with specific α-helical properties. Bioinformatics 24:2101–2102. https://doi.org/10.1093/bioinformatics/btn392
Geourjon C, Deléage G (1995) Sopma: significant improvements in protein secondary structure prediction by consensus prediction from multiple alignments. Bioinformatics 11:681–684. https://doi.org/10.1093/bioinformatics/11.6.681
Guruprasad K, Reddy BVB, Pandit MW (1990) Correlation between stability of a protein and its dipeptide composition: a novel approach for predicting in vivo stability of a protein from its primary sequence. Protein Eng Des Sel 4:155–161. https://doi.org/10.1093/protein/4.2.155
Hetényi C, van der Spoel D (2009) Efficient docking of peptides to proteins without prior knowledge of the binding site. Protein Sci 11:1729–1737. https://doi.org/10.1110/ps.0202302
Hollmann A, Martínez M, Noguera ME et al (2016) Role of amphipathicity and hydrophobicity in the balance between hemolysis and peptide-membrane interactions of three related antimicrobial peptides. Colloids Surf B Biointerfaces 141:528–536. https://doi.org/10.1016/j.colsurfb.2016.02.003
Huang F, Nau WM (2003) A conformational flexibility scale for amino acids in peptides. Angew Chem Int Ed 42:2269–2272. https://doi.org/10.1002/anie.200250684
Innovagen (2015) Peptide property calculator. https://pepcalc.com/. Accessed 14 Jan 2020
Jiang Z, Vasil AI, Hale JD et al (2008) Effects of net charge and the number of positively charged residues on the biological activity of amphipathic α-helical cationic antimicrobial peptides. Biopolym Pept Sci Sect 90:369–383. https://doi.org/10.1002/bip.20911
Jones DT (1999) Protein secondary structure prediction based on position-specific scoring matrices. J Mol Biol 292:195–202. https://doi.org/10.1006/jmbi.1999.3091
Juretić D, Vukičević D, Ilić N et al (2009) Computational design of highly selective antimicrobial peptides. J Chem Inf Model 49:2873–2882. https://doi.org/10.1021/ci900327a
Karplus PA, Schulz GE (1985) Prediction of chain flexibility in proteins - a tool for the selection of peptide antigens. Naturwissenschaften 72:212–213. https://doi.org/10.1007/BF01195768
König E, Bininda-Emonds ORP, Shaw C (2015) The diversity and evolution of anuran skin peptides. Peptides 63:96–117. https://doi.org/10.1016/j.peptides.2014.11.003
Kumar S, Stecher G, Li M et al (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Ladram A, Nicolas P (2016) Antimicrobial peptides from frog skin: biodiversity and therapeutic promises 3. 1. Intraspecies diversity of frog skin AMPs: each species of frog has a customized arsenal of peptide weapons 3. Biodivers Antimicrob 21:1341–1371
Leite JR, Silva LP, Rodrigues MI et al (2005) Phylloseptins: a novel class of anti-bacterial and anti-protozoan peptides from the Phyllomedusa genus. Peptides 26:565–573. https://doi.org/10.1016/j.peptides.2004.11.002
Li RF, Lu ZF, Sun YN et al (2016) Molecular design, structural analysis and antifungal activity of derivatives of peptide CGA-N46. Interdiscip Sci Comput Life Sci 8:319–326. https://doi.org/10.1007/s12539-016-0163-x
Life Technologies (2012) Vector NTI. Thermo Fisher Scientific. Waltham MA
López Cascales JJ, Zenak S, García De La Torre J et al (2018) Small cationic peptides: influence of charge on their antimicrobial activity. ACS Omega 3:5390–5398. https://doi.org/10.1021/acsomega.8b00293
Malmsten M (2014) Antimicrobial peptides. Ups. J Med Sci 119:199–204
Marshall SH, Arenas G (2003) Antimicrobial peptides: a natural alternative to chemical antibiotics andapotential for applied biotechnology. Electron J Biotechnol 6:96–109. https://doi.org/10.2225/vol6-issue3-fulltext-1
Matsuzaki K (2009) Control of cell selectivity of antimicrobial peptides. BBA Biomembr 1788:1687–1692. https://doi.org/10.1016/j.bbamem.2008.09.013
Morris G, Huey R, Linkstrom W et al (2010) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 16:2785–2791. https://doi.org/10.1002/jcc
Nacif-Marçal L, Pereira GR, Abranches MV et al (2015) Identification and characterization of an antimicrobial peptide of Hypsiboas semilineatus (Spix, 1824) (Amphibia, Hylidae). Toxicon 99:16–22. https://doi.org/10.1016/j.toxicon.2015.03.006
Oleg T, Arthur JO (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461. https://doi.org/10.1002/jcc
PerkinElmer (2009) ChembioDraw. PerkinElmer Informatics. Waltham MA
Pirtskhalava M, Amstrong AA, Grigolava M et al (2021) DBAASP v3: database of antimicrobial/cytotoxic activity and structure of peptides as a resource for development of new therapeutics. Nucleic Acids Res 49:D288–D297. https://doi.org/10.1093/nar/gkaa991
Powers JS, Hancock REW (2003) The relationship between peptide structure and antibacterial activity. Peptides 24:1681–1691. https://doi.org/10.1016/j.peptides.2003.08.023
Proaño-Bolaños C, Zhou M, Wang L et al (2016) Peptidomic approach identifies cruzioseptins, a new family of potent antimicrobial peptides in the splendid leaf frog, Cruziohyla calcarifer. J Proteomics 146:1–13. https://doi.org/10.1016/j.jprot.2016.06.017
Proaño-Bolaños C, Blasco-Zúñiga A, Almeida JR et al (2019a) Unravelling the skin secretion peptides of the gliding leaf frog, Agalychnis spurrelli (hylidae). Biomolecules 9:667. https://doi.org/10.3390/biom9110667
Proaño-Bolaños C, Zhou M, Chen T, Shaw C (2019b) Skin secretion transcriptome remains in chromatographic fractions suitable for molecular cloning. Anal Biochem 564–565:13–15
Romero SM, Cardillo AB, Martínez Ceron MC et al (2019) Temporins: an approach of potential pharmaceutic candidates. Surg Infect 20:1–14. https://doi.org/10.1089/sur.2019.266
Ron SR, Read M, Pazmiño-Armijos G (2018) Anfibios del Ecuador. In: Mus. Zool. Pontif. Univ. Católica del Ecuador. https://bioweb.bio/faunaweb/amphibiaweb/FichaEspecie/Cruziohylacalcarifer
Roy M, Lebeau L, Chessa C, Damour A, Ladram A, Oury B, Boutolleau D, Bodet C, Lévêque N (2019) Comparison of anti-viral activity of frog skin anti-microbial peptides Temporin-Sha and [K3]SHa to LL-37 and Temporin-Tb against Herpes Simplex Virus Type 1. Viruses. https://doi.org/10.3390/v11010077
Rozenski (1999) Peptide Mass Calculator v3.2. http://rna.rega.kuleuven.be/masspec/pepcalc.htm. Accessed 14 Jan 2020
Schrödinger L (2015) The PyMol molecular graphics system, Versión 1.8. Thomas Hold
Shang D, Liu Y, Jiang F et al (2019) Synergistic antibacterial activity of designed Trp-containing antibacterial peptides in combination with antibiotics against multidrug-resistant Staphylococcus epidermidis. Front Microbiol 10:1–15. https://doi.org/10.3389/fmicb.2019.02719
Vanhoye D, Bruston F, Nicolas P, Amiche M (2003) Antimicrobial peptides from hylid and ranin frogs originated from a 150-million-year-old ancestral precursor with a conserved signal peptide but a hypermutable antimicrobial domain. Eur J Biochem 270:2068–2081. https://doi.org/10.1046/j.1432-1033.2003.03584.x
Varga JFA, Bui-Marinos MP, Katzenback BA (2019) Frog skin innate immune defences: sensing and surviving pathogens. Front Immunol 10:1–21. https://doi.org/10.3389/fimmu.2018.03128
Wechselberger C (1998) Cloning of cDNAs encoding new peptides of the dermaseptin-family. Biochim Biophys Acta Protein Struct Mol Enzymol 1388:279–283. https://doi.org/10.1016/S0167-4838(98)00202-7
WHO (2014) Antimicrobial resistance. Global report on surveillance. World Health Organ 61:12–28. https://doi.org/10.1007/s13312-014-0374-3
Wilkins MR, Gasteiger E, Bairoch A et al (1999) Protein identification and analysis tools in the ExPASy server. In: Methods in molecular biology (Clifton, N.J.). pp 531–552
Xi X, Li R, Jiang Y et al (2013) Medusins: a new class of antimicrobial peptides from the skin secretions of phyllomedusine frogs. Biochimie 95:1288–1296. https://doi.org/10.1016/j.biochi.2013.02.005
Xu X, Lai R (2015) The chemistry and biological activities of peptides from amphibian skin secretions. Chem Rev 115:1760–1846. https://doi.org/10.1021/cr4006704
Zasloff M (2002) Antimicrobial peptides of multicellular organisms. Nature 415:389–395. https://doi.org/10.1038/415389a
Acknowledgements
This research was done under the framework contract for genetic resources access between MAE and IKIAM (MAE-DNB-CM-2016-0051) and is part of the project “Conservation of Ecuadorian amphibian diversity and sustainable use of its genetic resources,” which involves MAE, IKIAM-Universidad Regional Amazónica, Queen’s University Belfast, and Centro Jambatu, and the help of the Global Environmental Facility (GEF) and “Programa de las Naciones Unidas para el Desarrollo” (PNUD). We thank the kindly donation of bacterial and fungal strains by Dr. Jorge Reyes (INSPI).
Funding
This research was funded by research grants to L.M. and MR of the Dirección General Académica of the Pontificia Universidad Católica del Ecuador, projects: QINV0180-IINV529020200 and QINV0209-IINV529010100. This research was partially funded by the GEF Project ID 5534 “Conservation of Ecuadorian Amphibian Diversity and Sustainable Use of its Genetic Resources”, Ministry of Environment of Ecuador and Universidad Regional Amazonica IKIAM.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Research involving human participants and/or animals
All of the animal procedures were approved by the Laboratorio de Investigación en Citogenética y Biomoléculas de Anfibios (LICBA) Animal Care, and the International Guidelines for Animal Research were strictly followed in this study.
Informed consent
All authors listed have contributed to conception, design, gathering, analysis or interpretation of data, and have contributed to the writing and intellectual content of the article. All authors gave informed consent to the submission of this manuscript.
Additional information
Handling editor: F. Albericio.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cuesta, S.A., Reinoso, C., Morales, F. et al. Novel antimicrobial cruzioseptin peptides extracted from the splendid leaf frog, Cruziohyla calcarifer. Amino Acids 53, 853–868 (2021). https://doi.org/10.1007/s00726-021-02986-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-021-02986-w