Abstract
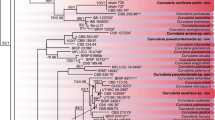
Lettuce necrotic yellows virus (LNYV) is the type member of the genus Cytorhabdovirus, family Rhabdoviridae, and causes a severe disease of lettuce (Lactuca sativa L.). This virus has been described as endemic to Australia and New Zealand, with sporadic reports of a similar virus in Europe. Genetic variability studies of plant-infecting rhabdoviruses are scarce. We have extended a previous study on the variability of the LNYV nucleocapsid gene, comparing sequences from isolates sampled from both Australia and New Zealand, as well as analysing symptom expression on Nicotiana glutinosa. Phylogenetic and BEAST analyses confirm separation of LNYV isolates into two subgroups (I and II) and suggest that subgroup I is slightly older than subgroup II. No correlation was observed between isolate subgroup and disease symptoms on N. glutinosa. The origin of LNYV remains unclear; LNYV may have moved between native and weed hosts within Australia or New Zealand before infecting lettuce or may have appeared as a result of at least two incursions, with the first coinciding with the beginning of European agriculture in the region. The apparent extinction of subgroup I in Australia may have been due to less-efficient dispersal than that which has occurred for subgroup II – possibly a consequence of suboptimal interactions with plant and/or insect hosts. Introduction of subgroup II to New Zealand appears to be more recent. More-detailed epidemiological studies using molecular tools are needed to fully understand how LNYV interacts with its hosts and to determine where the virus originated.



Similar content being viewed by others
References
Behncken GM (1983) A disease of chickpea caused by Lettuce necrotic yellows virus. Australas Plant Pathol 12:64–65
Blancard D, Lot H, Maisonneuve B (2006) A color atlas of diseases of lettuce and related salad crops: observation, biology and control. Manson Publishing, London
Boakye DB, Randles JW (1974) Epidemiology of lettuce necrotic yellows virus in South Australia. III: Virus transmission parameters and vector feeding behaviour in host and non-host plants. Aust J Agric Res 25:791–802
Bouckaert R, Heled J, Kühnert D, Vaughan T, Wu CH, Xie D, Suchard MA, Rambaut A, Drummond AJ (2014) BEAST 2: a software platform for Bayesian evolutionary analysis. PLoS Comput Biol 10:e1003537
Callaghan B (2005) Sequence analysis and variability study of Lettuce necrotic yellows virus. The University of Queensland, p 285
Callaghan B, Dietzgen RG (2005) Nucleocapsid gene variability reveals two subgroups of Lettuce necrotic yellows virus: brief report. Arch Virol 150:1661–1667
Dietzgen RG, Callaghan B, Wetzel T, Dale JL (2006) Completion of the genome sequence of Lettuce necrotic yellows virus, type species of the genus Cytorhabdovirus. Virus Res 118:16–22
Dietzgen RG, Callaghan B, Campbell PR (2007) Biology and genomics of Lettuce necrotic yellows virus. Plant Viruses. Global Science Books, pp 85–92
Fletcher J, France C, Butler R (2005) Virus surveys of lettuce crops and management of lettuce big-vein disease in New Zealand. N Z Plant Prot 58:239–244
Francki RIB, Randies JW, Dietzgen RG (1989) Lettuce necrotic yellows virus. Description of Plant Viruses No 343. Association of Applied Biologists, Wellesbourne
Fry P, Close R, Procter C, Sunde R (1973) Lettuce necrotic yellows virus in New Zealand. N Z J Agric Res 16:143–146
Fry PR, Close RC, Procter CH, Sunde R (1973) Lettuce necrotic yellows virus. Descriptions of Plant Viruses No 343. Association of Applied Biologists, Wellesbourne
Heim F, Lot H, Delecolle B, Bassler A, Krczal G, Wetzel T (2008) Complete nucleotide sequence of a putative new cytorhabdovirus infecting lettuce. Arch Virol 153:81–92
Ito T, Suzaki K, Nakano M (2013) Genetic characterization of novel putative rhabdovirus and dsRNA virus from Japanese persimmon. J Gen Virol 94:1917–1921
Kim SC, Lu CT, Lepschi BJ (2004) Phylogenetic positions of Actites megalocarpa and Sonchus hydrophilus (Sonchinae: Asteraceae) based on ITS and chloroplast non-coding DNA sequences. Aust Syst Bot 17:73–81
Kim SC, Chunghee L, Mejías JA (2007) Phylogenetic analysis of chloroplast DNA matK gene and ITS of nrDNA sequences reveals polyphyly of the genus Sonchus and new relationships among the subtribe Sonchinae (Asteraceae: Cichorieae). Mol Phylogenet Evol 44:578–597
Klerks MM, Lindner JL, Vaškova D, Špak J, Thompson JR, Jelkmann W, Schoen CD (2004) Detection and tentative grouping of Strawberry crinkle virus isolates. Eur J Plant Pathol 110:45–52
MacPhail M, Hope G, Anderson A (2001) Polynesian plant introductions in the Southwest Pacific: initial pollen evidence from Norfolk Island. Rec Aust Mus Suppl 27:123–134
Ragozzino A, Alioto D, Iengo C (1989) The yellowing virus and mycoplasma diseases of lettuce in Campana and Latium regions. Rivista di Patologia Vegetale 25:15–19
Randles JW, Carver M (1971) Epidemiology of lettuce necrotic yellows virus in South Australia. II: distribution of virus, host plants, and vectors. Aust J Agric Res 22:231–237
Rubio-Huertos M, Garcia-Hidalgo F (1982) A rhabdovirus resembling lettuce necrotic yellows from lettuce in Spain. Phytopathol Z 103:232–238
Stubbs LL, Grogan RG (1963) Necrotic yellows: a newly recognized virus disease of lettuce. Aust J Agric Res 14:439–459
Sward RJ (1990) Lettuce necrotic yellows rhabdovirus and other viruses infecting garlic. Australas Plant Pathol 19:46–51
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tanno F, Nakatsu A, Toriyama S, Kojima M (2000) Complete nucleotide sequence of Northern cereal mosaic virus and its genome organization. Arch Virol 145:1373–1384
Wetzel T, Dietzgen RG, Dale JL (1994) Genomic organization of lettuce necrotic yellows rhabdovirus. Virology 200:401–412
Acknowledgments
The authors would like to thank the following people for assistance in the collection of samples: Prof. John W. Randles (AU18), Ms. Smriti Nair and Ms. Gardette Valmonte (NZ1-6). Isolates AU13, 14 and 17 were kindly provided by Dr. John Thomas. Technical assistance by Mr. Roger Mitchell in the early stages of this work is gratefully acknowledged. The authors would like to thank Jade Hsu, Amy Chang and Ee Ren Tan for their assistance in sequencing isolates AU13-14, AU15-6 and AU17, respectively. CH would also like to thank Dr. Shane Lavery for useful discussions. CH received funding support from the Faculty of Health and Environmental Sciences, Auckland University of Technology. This research was supported by Australian Research Council grant LP110100047.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Higgins, C.M., Chang, WL., Khan, S. et al. Diversity and evolutionary history of lettuce necrotic yellows virus in Australia and New Zealand. Arch Virol 161, 269–277 (2016). https://doi.org/10.1007/s00705-015-2626-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-015-2626-5