Abstract
A mortality rate higher than 90 % was observed in a larva-rearing facility for Pacific cod, Gadus macrocephalus, in China. Larvae showing clinical signs of infection were collected. Initial suspicion of nervous necrosis virus (NNV) infection was confirmed by sequencing, absolute quantification real-time PCR (A-qPCR), and electron microscopy. The nucleotide sequence of RNA2 was 1,375 bases long (GenBank no. KM576685), coding for a single ORF corresponding to the capsid protein from residues 21 to 1034. Phylogenetic analysis of the capsid protein sequence showed that PCNNV belongs to the barfin flounder NNV (BFNVV) genotype. An amino acid sequence alignment revealed 39 differences between the cold- and warm-resistant viral groups, suggesting that PCNNV evolved under temperature selection. The 3-D structure of the predicted capsid protein was modeled to identify potential epitopes, and the gene was expressed in Escherichia coli, yielding a protein with a molecular mass of 55 kDa. During PCNNV outbreaks, the viral copy number was found to reach 107 per ng of total RNA, which could be considered the lethal copy number of NNV in cod. The gonads, eggs, fertilized eggs and asymptomatic cod fry were all positive for PCNNV, indicating viral vertical transmission as the main source of the viral load. The amount of virus in the apparent healthy fry or survivors seemed to decrease gradually with development. These results might lead to efficient diagnostic methods to help farmers select NNV-free broodfish for cod breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nervous necrosis viruses (NNVs) of the genus Betanodavirus, family Nodaviridae, are pathogens that cause viral nervous necrosis (VNN), a disease observed in marine aquaculture hatcheries worldwide. VNN can cause 100 % mortality and affect more than 30 species of fish [30], such as barramundi (Lates calcarifer) [13], Japanese sea bass (Lateolabrax japonicas) [21], sea bass (Dicentrarchus labrax) [6], red-spotted grouper (Epinephelus akaara) [29], striped jack (Pseudocaranx dentex), sea bream (Sparus aurata) [9], halibut (Hippoglossus hippoglossus) [15], turbot (Scophthalmus maximus) [5], and tiger puffer (Takifugu rubripes) [32].
Betanodaviruses have a linear, bipartite (RNA1 and RNA2 transcripts), positive-sense, single-stranded RNA genome. They are non-enveloped viruses with an isosahedral capsid (triangulation number = 3) ranging from 29 to 35 nm in diameter. The capsid consists of 32 capsomers [16, 28]. Betanodaviruses are divided into four genotypes based on the nucleotide sequence of the capsid protein gene [33]. These include red-spotted grouper (RGNNV), striped jack (SJNNV), barfin flounder (BFNNV), and tiger puffer (TPNNV) NNV. NNV carriers are identified by their abnormal swimming behavior, consisting of looping or spiral swimming in a belly-up position and loss of coordination [31]. Viral particles with a diameter in the range of 25-34 nm can be visualized in the brain by electron microscopy [36]. Vacuolation has been observed in tissue sections of the brain and eyes [3]. The use of molecular methods such as sequencing and PCR seems to be the most precise and efficient way to detect the presence of virus. In addition, bioinformatics provides useful information about the molecular epidemiology of VNN [15] and helps in the prediction of epitopes.
Pacific cod, Gadus macrocephalus, as an important commercial food species, is considered a potential culture target (FAO). Farming of G. macrocephalus has been carried out for 8 years in our lab (Key Laboratory of Mariculture & Stock Enhancement in North China’s Sea, Ministry of Agriculture, Dalian) [11, 14, 24, 45], but the farming work is still in the laboratory stage. During breeding, G. macrocephalus with looping or spiral swimming in a belly-up position and loss of coordination could be observed every year, especially at 20 to 45 days post-hatching (dph). Usually, over 90 % mortality is observed in cod with the clinical signs. We attempted to isolate bacteria from the abnormal cod, but we found no evidence of the presence of bacteria, and this conclusion was supported by the lack of effect of antibiotics (data not shown). The high morbidity of hatchery-reared G. macrocephalus limits their production. We designed a set of primers based on known fish bacteria and viruses, and the results of PCR showed that the cod sample was NNV positive.
In the present study, we report the sequence of NNV capsid from farmed cod larvae showing typical clinical signs. Absolute quantification PCR (A-qPCR) and electron microscopy were used to confirm that the pathogen was NNV and to investigate its mode of transmission. In addition, a prokaryotic expression system was established to produce the coat protein for further vaccine research.
Materials and methods
Cloning and expression of the NNV capsid gene
Cloning and sequencing: Total RNA was extracted from cod larvae with clinical signs using TriPure Isolation Reagent (ROCHE, Switzerland). Reverse transcription (PrimerScript™ RT-PCR kit, TaKaRa, Japan) was then used to make cDNA, which was then used as a template to amplify partial NNV cDNA sequences by polymerase chain reaction (PCR) using the specific primers NNVF1 (5′-GTGGTTACGTGGCTGGTTC-3′) and NNVR1 (5′-CAAATGGTGGGAAAGCAGAAC-3′), which have been published previously [42]. PCR amplification was done using the following cycling conditions: 94 °C for 5 min, 35 cycles of denaturation at 94 °C for 30 s, annealing at 56 °C for 30 s, and elongation at 72 °C for 1 min, and a final 10-min elongation step at 72 °C. PCR products were analyzed for purity and size by electrophoresis in a 1 % agarose gel, purified from the gel using a QIAquick Gel Extraction Kit (QIAGEN, Germany), and ligated into the pMD18-T vector according to the manufacturer’s protocol (TaKaRa, Japan). After overnight incubation at 4 °C, the ligation mix was introduced into competent E. coli DH5α cells (TransGen, China) by transformation. Clones were sequenced (Invitrogen) to verify the partial cDNA sequences using BLAST. The full-length sequence of NNV RNA2 were obtained by using the primers NNVF2 (5′-AACACTGCTTTGCCATCA-3′) and NNVR2 (5′-GCTGACTCGGGAAACTAAC-3′), which were designed to correspond to conserved regions of known fish NNV sequences. PCR products were detected and purified using a gel extraction kit (QIAGEN, Germany) and ligated into the vector pMD18-T (TaKaRa, Japan). The recombinant plasmid, named pMD-NNV, was introduced into E. coli by transformation and sequenced (Invitrogen) to verify the full-length cDNA sequences of the Pacific cod NNV (PCNNV).
Prokaryotic expression: A pair of specific primers was designed based on the Tripure Isolation Reagent newly obtained sequence of PCNNV. The sequences of the forward (F4) and reverse primers (R4) were 5′-GGCCATGGTACGCAAAGGGAATAAGAAATTG-3′ and 5′-CATGCGGCCGCTTAATGGTGATGGTGATGATGGTTTCCCGAGTCAGCCCGGGTGAAG-3′, respectively. The primers included protective bases as well as NcoI and NotI restriction sites for digestion. The primers were synthesized by Shanghai Invitrogen Biotechnology Co., Ltd. PCR was conducted using F4 and R4. The recovered PCR products and plasmid pET32a were digested with NcoI and NotI, and the expression vector pEVCP was constructed by fusing a fragment encoding the major capsid protein with the linearized plasmid pET32a using T4 DNA ligase (New England Biolabs, Beijing). DNA sequences were verified at each step. E. coli BL21(DE23) cells were transformed with pEVCP and grown at 37 °C in LB medium supplemented with 100 µg of ampicillin per mL. Recombinant protein expression was induced at an OD600 of 0.4-1.0 by treatment with 1 mM isopropyl β-D-1-thiogalactopyranoside (IPTG, TransGen, China) for 3 h. A 1-mL aliquot was taken before induction to monitor protein expression. The cells were collected by centrifugation and resuspended in 100 μL of PBS buffer and 1× SDS-PAGE loading buffer, boiled for 5-10 minutes, and centrifuged, and the supernatants were analyzed by SDS-PAGE to examine protein expression.
Sequence analysis
The sequence of PCNNV obtained in this study was submitted to the National Center for Biotechnology Information (NCBI) to perform multiple alignments (http://www.ncbi.nlm.nih.gov/). A multiple sequence alignment including the deduced amino acid sequence was made using ClustalW 1.81. A sequence alignment was made using BioEdit software. A phylogenetic tree was constructed by the maximum-likelihood (ML) method, using MEGA version 5 [40]. A 3-D model was constructed using Phyre2 [22]. A Ramachandran plot was generated using PROCHECK (http://www.ebi.ac.uk/~roman).
Establishment of an absolute quantitative real-time PCR (A-qPCR)
Plasmid pMD-NNV was extracted and purified from transfected E. coli cells using a Plasmid Kit (TransGen, China). The plasmid copy number was calculated using the following formula: number of copies = (amount × 6.022 × 1023) / (length × 1 109 × 650) as described by Staroscik (http://cels.uri.edu/gsc/cndna.html). Purified plasmid DNA was stored in aliquots at -20 °C. A decreasing series of tenfold dilutions was made from purified plasmid DNA to make a standard curve of the target gene. A SYBR Green real-time qPCR assay was performed using an ABI 7500 Real Time PCR System, and Ct values and a standard curve were obtained using the 7500 software version 2.0.6. The gene-specific primers for qPCR were NNVF3 (5′-GTGGTTACGTGGCTGGTTC-3′) and NNVR3 (5′-CAAATGGTGGGAAAGCAGAAC-3′). The qPCR conditions were as follows: 50 °C for 2 min, 95 °C for 10 min, and 40 cycles of denaturing at 95 °C for 15 s, followed by annealing and primer extension at 60 °C for 1 min. PCR products were also purified and sequenced (Invitrogen) to confirm the specificity of the primers. In each assay, a standard curve was generated and used to estimate the copy number of target sequences in unknown samples.
PCNNV detection using A-qPCR
PCNNV in apparent healthy larvae and larvae showing clinical signs
Five pieces of apparent healthy larvae were sampled in 1.5-mL tubes at 5, 10, 20, 25, 30, and 45 days post-hatching (dph). Larvae/juveniles showing clinical signs of NNV-like disease (uncoordinated, corkscrew-like swimming behavior) were sampled from the farming facility at 37, 46, 64, 76, 82, 85 dph.
Route of PCNNV transmission
During farming, the food source of brine shrimp (Artemia franciscana), chlorella (Chlorella pyrenoidosa) and nutrition enhancer (Schizochytrium sp.) were collected separately for NNV detection. All samples were frozen in liquid nitrogen and stored at –80 °C for molecular analysis.
Fertilized eggs were collected from an artificial breeding field at Dalian Ocean University during the winter of - 2013 and 2014. While in the process of hatching, 15 to 20 eggs were collected in 1.5-mL tubes.
Total RNAs of the above samples were extracted using an RNAprep pure Tissue Kit (TIANGEN, China) according to the manufacturer’s instructions. Total RNA was used as template for reverse transcription to synthesize the first chain of cDNA (TaKaRa, Japan). The absolute quantification of PCNNV transcripts was done using the relative standard curve method described above.
Electron microscopy
Brain samples from dying larvae or young cod were collected and prepared according to the manual of the transmission electron microscope JEM-2000EX (JEOL). Micrographs were taken using a JEM-2000EX microscope at magnifications ranging from 30000× to 200,000×.
Results
RNA2 sequencing and in vitro expression of the PCNNV capsid protein
Larvae exhibiting typical VNN-like behaviour were collected for total RNA extraction. The symptoms included uncoordinated corkscrew-like swimming behaviour, hypersensitivity to stimuli, darkening of the body and assembly into large groups. Primers were designed based on the conserved region of BFNNV RNA2. The nucleotide sequence and translated amino acid sequence of the PCNNV coat protein gene were determined (Supplementary data). The nucleotide sequence was 1375 bases long (GenBank accession no. KM576685) and contained a single ORF spanning positions 21-1034. Thus, there were 20 bases in the 5′-non-coding region and 338 bases in the 3′-non-coding region. The ORF encoded a polypeptide of 338 amino acids with a calculated molecular mass of 36.906 kDa. The plasmid construct pEVPC, which included the ORF, was constructed in a pET32a vector with an N-terminal Trx tag and a C-terminal 8X-His tag, resulting in a fused protein molecular weight of 55 kDa (Fig. 1). A 3-D model of the capsid protein was constructed based on the crystal structure of an N-terminally truncated capsid protein mutant2 of Orsay virus (PDB: 4nww) [16]. A structure consisting of 168 residues (50 % of the capsid protein sequence) was modelled with 100 % confidence using the template (Fig. 2A). The secondary structure of the 168-amino-acid fragment was generated by PDBsum Generate (Fig. 2C), and the 3-D model was analyzed using PROCHECK. A Ramachandran Plot showed that 79.7 % of the residues were in most-favored regions, and none were in disallowed regions (Fig. 2B). The overall average of R-factors generated by PROCHECK statistics was -0.30, which indicates a normal structure.
Features of the PCNNV capsid protein. (A) 3D model of the PCNNV capsid protein. The model was generated using Phyre2 [22]. The image was colored using a rainbow gradient from the N- to the C-terminus. The model was made using c4nwwC as a template. Information of c4nwwC: PDB header, virus; PDB molecule, capsid protein; PDB title, Crystal structure of an N-terminally truncated capsid protein mutant2 of Orsay virus. In this model, 168 residues (50 % of the capsid protein sequence) were modelled with 100 % confidence using the single highest-scoring template. (B) Ramachandran Plot. A Ramachandran plot was generated using PROCHECK. Plot statistics: residues in most-favoured regions [A, B, L], 114 (79.7 %); residues in additional allowed regions [a, b, l, p], 28 (19.6 %); residues in generously allowed regions [~a, ~b, ~l, ~p], 1 (0.7 %); residues in disallowed regions 0 (0.0 %). (C) Secondary structure predictions were made using the software PDBsum Generate. Motifs: β, beta turn; γ, gamma turn;  , beta hairpin.
, beta hairpin.  Helices are labeled H1, H2, etc., and strands according to their sheets (A, B, etc.)
Helices are labeled H1, H2, etc., and strands according to their sheets (A, B, etc.)
Alignment and phylogenetic analysis
Sequence alignment showed that the PCNNV coat protein was closely related to Atlantic cod NNV (ABU95413.1). The most closely related sequences were from Atlantic cod NNV (98.8 %), barfin flounder NNV (ACF36170; 98.5 %), Atlantic halibut NNV (CAB53257.1; 95.9 %) and haddock NNV (AAT28125; 95.6 %). Phylogenetic analysis was carried out using capsid protein gene sequences obtained from NCBI. The coat protein sequences of fish NNVs were obtained between 1994 and 2014 from different countries. There were five branches, representing the five genotypes, e.g., RGNNB, BFNNV, TPNNV, SJNNV and TNNV. The analysis showed that the recent PCNNV was grouped within the BFNNV genotype (Fig. 3). Sequence alignment highlighted differences in NNV genotypes based on 39 amino acid substitutions generating a dual genotype classification (Fig. 4): NNV virus infecting cold-and warm-water species. However, striped jack and turbot were excluded.
Phylogenetic analysis of NNV coat proteins from various fish species. The phylogenetic tree was constructed by the maximum-likelihood (ML) method. The scale bars show the number of substitutions as a proportion of branch length. The genotypes of RGNNV, BFNNV, TNNV, TPNNV and SJNNV were classified according to Nishizawa et al. [33]
A-qPCR protocol optimization
In this study, the method used for NNV detection is key to understanding the mechanism of virus transmission. Melting curve analysis and DNA sequencing confirmed the amplification specificity. The results showed a sharp peak for the PCR product of the quantitative standard sample, pMD-NNV, at 82.0 °C (Fig. 5A). A single melting peak at the same melting temperature was produced for the PCR product of the total DNA sample. The identities of amplified products were confirmed by sequencing.
Absolute quantification of PCNNV by real-time PCR. (A) Melting curve. The melting temperature and the amplicon lengths were 82 °C and 113 bp for PCNNV detection. (B) A standard curve was constructed using serial tenfold dilutions of the pMD-VNN plasmid, ranging from 106-102 copies/μL from left to right. Each standard dilution was amplified by real-time qPCR in triplicate (slope = -3.416; R2 = 0.994; Eff % = 96.234)
Serial dilutions of the pMD-NNV plasmid DNA were made, and amplification plots for qPCR assays were generated using the primers NNVF3 and NNVR3. The log-linear standard curve (Fig. 5B) from these plots showed a fit of R2 = 0.994, indicating high accuracy over a wide range of concentrations ranging from 102 to 106 copies of pMD-NNV plasmid. The formula generated based on the standard curve is “y = -3.1756x + 36.171”. The highest dilution at which reliable amplification occurred was defined as the detection limit, which was 20 copies of pMD-NNV.
PCNNV detection in symptomatic carriers
Dying cod with evident clinical signs of infection (floating and erratic swimming) were sampled separately at 37, 46, 64, 76 and 85 dph, and their brains were collected for total RNA extraction. Reverse transcription was performed using 500 ng total RNA for each sample. Ct values of the PCNNV in each sample were obtained by qPCR, and the viral copy number (VCN) was calculated using the formula “y = -3.1756x + 36.171” for the standard curve. The result showed that the VCN was up to 107 (Fig. 6A), using a dilution series of the plasmid pMD-NNV for calibration. In order to confirm the qPCR result, we imaged brain tissue using TEM (Fig. 7). Viral particles of 25 nm in diameter were observed in the brain samples, and these correlated with the presence of many vacuoles, in accordance with previous reports [7, 20, 41]. The evidence provided by qPCR and electron microscopy indicated that the dying young cod were suffering from a massive NNV infection during the first trimester after hatching.
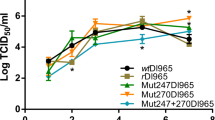
Detection of PCNNV using A-qPCR. (A) A-qPCR analysis of symptomatic cod during NVV outbreaks. Absolute number of PCNNV copies per ng of total RNA at different developmental stages of farmed cod during PCNNV outbreaks. Samples at different stages were plotted against the logarithm of the PCNNV copy number (n = 3). (B) A-qPCR analysis of the developmental viral load for asymptomatic cod. Samples at different stages were plotted against the logarithm of the PCNNV copy number (n ≥ 3)
PCNNV detection in potential asymptomatic carriers
Asymptomatic hatchery-reared cods without clinical signs were collected at 1, 5, 10, 20, 25, 30 and 45 dph for PCNNV detection. Copy numbers of each sample were calculated by the formula ‘y=-3.1756x + 36.171’. The mean VCN per total RNA decreased gradually from 1 dph to 45 dph as shown in Fig. 6B. However, even after 45 dph, samples were NNV positive (data not shown).
In order to evaluate the possibility of vertically transmitted infection, female gonads, ova, and fertilized eggs were collected from several wild Pacific cod and tested using A-qPCR. Gonads, eggs and fertilized eggs were found to be NNV positive (Fig. 8), and the VCN ranged from 103 per total RNA in some eggs (ovum-2) compared to less than 10 per total RNA in others (ovum-1). The viral content of PCNNV was quite different among broodfish. Viral horizontal transmission was also examined, especially in the feed. However, brine shrimp, chlorella and enhancer limacinum all tested NNV negative (data not shown).
Discussion
This is the first report of an outbreak of NNV in farmed Pacific cod in China. The clinical observations (floating and erratic swimming) are consistent with other reports of nodavirus infections in cod [2, 39, 43]. The nucleic acid sequence of RNA2 and the amino acid sequence of the capsid protein were determined. The ORF encoding the putative coat protein is 993 nt long, and the calculated molecular mass of 36,906 Da was confirmed by recombinant protein expression (Fig. 2). In a Ramachandran plot, 79.7 % of the residues in our 3-D model of the coat protein were in the most-favoured regions, which indicated that this model needs to be modified further. As R-factors below -0.5 are unusual and those below -1.0 are highly unusual [23], an R-factor of -0.30 is within the normal range. This model will help in identifying epitopes that are relevant for vaccine development.
NNV is indeed a worldwide viral disease. The virus infects not only warm-water species but also causes outbreaks in cold-water species. Phylogenetic analysis based on the nucleotide sequence of the capsid protein showed that it was related to a betanodaviruses within the genotype BFNNV, with high homology. Five genotypes of NNV (RGNNB, BFNNV, TPNNV, SJNNV and TNNV) have been classified in accordance with classical phylogenetic groups [44], and these show remarkable temperature specificity [4]. Interestingly, some fish species such as seabass (European and Asian subspecies) may show susceptibility to infection by both NNV isoforms (Fig. 4). NNV virus shows a tendency to infect warm-water fish species, but not exclusively. The question arises how NNV infects cold-water fish species, even fish living in sea water near the freezing point, e.g., cod [34]. It has been reported that the polymerase gene of RNA1 is subject to stronger positive selection pressure than RNA2, which has been shown to be important in determining the temperature sensitivity of betanodaviruses [35]. The possible reason was that the viral polymerase gene of RNA1 evolved significantly more rapidly than the coat protein gene [32]. One possible explanation for this difference in evolutionary rates is that there has been adaptation to local temperature conditions, which is known to modulate viral RNA replication and which is under the control of RNA1 [17]. Interestingly, the present study indicated that this virus strain was also grouped within the cluster of cold-resistant strains according to the results of amino acid alignments of coat protein sequences (Fig. 5), and all of the cold strains belonged to the BFNNV genotype. No correlation was found between the viral strain and specificity of host tropism except for sea bass, since the viruses with the same genotype are capable of infecting species belonging to different taxonomic orders [18].
Traditionally, severe vacuolation in the brain, spinal cord and retinal tissues of the eye is the most important and reliable diagnostic feature of VNN in fish [1, 8, 31]. Although immunohistochemistry (IHC) has been the most frequently used method, it requires specific antibodies, and it is expensive and time-consuming. In this study, A-qPCR was implemented as an efficient, low-cost, high-throughput, and reliable method to detect NNV infection based on the calibration of the system using the plasmid pMD-NNV. The standard curve was reliable, as R2 was close to 1, and the slope was close to −3.32, as shown in Fig. 6D. However, it has to be pointed out that there is a significant risk of laboratory contamination when working with plasmid standards.
When high mortality occurred at 37, 46, 64, 76 and 85 dph in 2014, the dying cod were sampled randomly to confirm virus infection. The results of A-qPCR showed that a very high copy number, greater than 106, was detected in the dying cods, which was beyond the range of the standard curve. This suggests that NNV occupies the nervous system, leading to fish death. The brains of the diseased cod were also analyzed by scanning electron microscopy to confirm NNV infection, and the results were in accordance with those of qPCR. Several particles around 25 nm in diameter were found in the imaged tissue (Fig. 7D).
In order to track the mechanism of PCNNV infection, asymptomatic cod without clinical evidence of infection were examined during the early developmental stages. Our previous study showed that NNV carrier larvae can survive, due to a sufficiently strong balance between innate and adaptive immunity [37]. When the maternal antibody pool is exhausted but the adaptive immune system is still immature, there is a “crisis period” in which there are not enough anti-NNV antibodies [25]. Interestingly, this study showed that the VCN in the first 20 days was much higher than that in the following days (Fig. 7B). Accordingly, virus content decreased significantly as developmental progressed, which was consistent with earlier studies of cod immune system ontogenesis [27]. Our previous work revealed that the immunity of surviving cod increased 2 months after hatching (Fig. 9). It has been reported that ontogenesis of the immune system was first tested on 16 dpf (days post-fertilization) in E. akaara [26], 4 dpf in zebrafish [47], common carp [19] and goldfish [35], 8 dpf in Japanese flounder [46] and 21 dpf in gadoid haddock [10]. From that time point, fish immunity increases gradually.
Analysis of Rag1 (A) and Igμ (B) expression during developmental stages in Pacific cod using A-qPCR [27]
Vertical transmission of NNV allows virus spread from generation to generation, which is the most challenging issue in fish farming. In this study, we confirmed the presence of NNV in wild-caught broodfish gonads, eggs, and fertilized eggs (Fig. 8). However, horizontal transmission cannot be ruled out, as it is difficult to distinguish the NNV carriers. In this study, the feed given to the fish was shown to be NNV negative, ruling out contamination of food as the cause of infection. Nodavirus has been shown to be very stable under extreme environmental conditions [12]. Once the virus has entered a farm or rearing unit, it may be very difficult to eliminate [38]. The phylogenetic tree for NNV showed high homology (~99 % sequence identity) among members of the genus Gadus, e.g., Pacific cod, Atlantic cod and haddock, which may account for inter-species virus spread, although this may depend on species habitat and interaction.
In conclusion, our findings demonstrate the presence of nodavirus in Pacific cod in China and confirm the genotype of BFNNV based on the capsid gene sequences. Amino acid alignment reveals that there are significant differences between cold- and warm-water species. Using an A-qPCR-based method, we confirmed that vertical transmission is the main mode of propagation of the virus. This assay can help farmers select NNV-negative broodfish. In addition, we have successfully expressed the viral capsid protein as a step towards producing a protective NNV vaccine, even though immunization seems to have a limited effect [38].
References
Aranguren R, Tafalla C, Novoa B, Figueras A (2002) Experimental transmission of encephalopathy and retinopathy induced by nodavirus to sea bream, Sparus aurata L., using different infection models. J Fish Dis 25:317–324
Azad IS, Shekhar MS, Thirunavukkarasu AR, Jithendran KP (2006) Viral nerve necrosis in hatchery-produced fry of Asian seabass Lates calcarifer: sequential microscopic analysis of histopathology. Dis Aquat Organ 73:123–130
Bigarré L, Cabon J, Baud M, Heimann M, Body A, Lieffrig F, Castric J (2009) Outbreak of betanodavirus infection in tilapia, Oreochromis niloticus (L.), in fresh water. J Fish Dis 32:667–673
Binesh CP, Renuka K, Malaichami N, Greeshma C (2013) First report of viral nervous necrosis-induced mass mortality in hatchery-reared larvae of clownfish, Amphiprion sebae Bleeker. J Fish Dis 36:1017–1020
Bloch B, Gravningen K, Larsen JL (1991) Encephalomyelitis among turbot associated with a picornavirus-like agent. Dis Aquat Organ 10:65–70
Breuil G, Bonami JR, Pepin JF, Pichot Y (1991) Viral infection (picorna-like virus) associated with mass mortalities in hatchery-reared sea-bass (Dicentrarchus labrax) larvae and juveniles. Aquaculture 97:109–116
Chi SC, Lo BJ, Lin SC (2001) Characterization of grouper nervous necrosis virus (GNNV). J Fish Dis 24:3–13
Chi SC, Lo CF, Kou GH, Chang PS, Peng SE, Chen SN (1997) Mass Mortalities associated with viral nervous necrosis (VNN) disease in two species of hatchery-reared grouper, Epinephelus fuscogutatus and Epinephelus akaara (Temminck & Schlegal). J Fish Dis 20:185–193
Comps M, Raymond JC (1996) Virus-like particles in the retina of the sea-bream, Sparus aurata. Bull Eur Assoc Fish Pathol 16:151–153
Corripio-Miyar Y, Bird S, Treasurer JW, Secombes CJ (2007) RAG-1 and IgM genes, markers for early development of the immune system in the gadoid haddock, Melanogrammus aeglefinus, L. Fish Shellfish Immun 23:71–85
Fan R, Jiang ZQ, Li YJ, Gao XQ, Qian C, Gao M (2014) Chromosome karyotypic analysis and Ag-NORs of Pacific cod Gadus macrocephalus. Acta Hydrobiol Sin 38:115–120
Frerichs GN, Tweedie A, Starkey WG, Richards RH (2000) Temperature, pH and electrolyte sensitivity, and heat, UV and disinfectant inactivation of sea bass (Dicentrarchus labrax) neuropathy nodavirus. Aquaculture 185:13–24
Glazebrook JS, Heasman MP, Beer SW (1990) Picornalike viral particles associated with mass mortalities in larval barramundi, Lates calcarifer Bloch. J Fish Dis 13:245–249
Gong X, Fan R, Jiang ZQ, Meng XK, Sun Y, Yang JJ (2013) Effects of pH values on digestive enzyme activity in various day old larvae of Pacific cod Gadus macrocephalus. Fish Sci 12:730–733
Grotmol S, Totland GK, Kvellestad A, Fjell K, Olsen AB (1995) Mass mortality of larval and juvenile hatchery-reared halibut (Hippoglossus hippoglossus L.) associated with the presence of virus-like particles in vacuolated lesions in the central nervous system and retina. Bull Eur Assoc Fish Pathol 15:176–180
Guo YR, Hryc C, Jakana J, Jiang H, Wang D, Chiu W, Zhong W, Tao YJ (2014) Crystal structure of an N-terminally truncated capsid protein mutant of Orsay virus. Proc Natl Acad Sci USA (in press)
Hata N, Okinaka Y, Iwamoto T, Kawato Y, Mori K, Nakai T (2010) Identification of RNA regions that determine temperature sensitivities in betanodaviruses. Arch Virol 155:1597–1606
Hick P, Gore K, Whittington R (2013) Molecular epidemiology of betanodavirus-sequence analysis strategies and quasispecies influence outbreak source attribution. Virology 436:15–23
Huttenhuis HBT, Huising MO, Meulen T, Oosterhoud CN, Sanchez NA, Taverne-Thiele AJ, Stroband HWJ, Rombout JHWM (2005) Rag expression identifies B and T cell lymphopoietic tissues during the development of common carp (Cyprinus carpio). Dev Comp Immunol 29:1033–1047
Iwamoto T, Mise K, Mori K, Arimoto M, Nakai T, Okuno T (2001) Establishment of an infectious RNA transcription system for Striped jack nervous necrosis virus, the type species of the betanodaviruses. J Gen Virol 82:2653–2662
Jung SJ, Miyazaki T, Miyata M, Oishi T (1996) Histopathological studies on viral nervous necrosis in a new host, Japanese sea bass Lateolabrax japonicus. Bull Fac Bioresour Mie Univ (Jpn) 16:9–16
Kelley LA, Sternberg MJE (2009) Protein structure prediction on the web: a case study using the Phyre server. Nat Protoc 4:363–371
Laskowski RA, Rullmannn JAC, MacArthur MW, Kaptein R, Thornton JM (1996) AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J Biomol NMR 8:477–486
Li YQ, Jang ZQ, Sun Y, Mao MG, Meng XK (2014) Experimental starvation on Gadus macrocephalus and definition of the point of no return. Acta Ecol Sin 34:3873–3878
Mao MG, Jiang JL, Perálvarez-Marín A, Wang KJ, Lei JL (2013) Characterization of the Mx and hepcidin genes in Epinephelus akaara asymptomatic carriers of the nervous necrosis virus. Aquaculture 408:175–183
Mao MG, Lei JL, Alex PM, Hong WS, Wang KJ (2012) Characterization of RAG1 and IgM (mu chain) marking development of the immune system in red-spotted grouper (Epinephelus akaara). Fish Shellfish Immun 33:725–735
Mao MG, Li X, Perálvarez-Marín Alejandro, Jiang JL, Jiang ZQ, Wen SH, Lü HQ (2015) Transcriptomic analysis and biomarkers (Rag1 and Igμ) for probing the immune system development in Pacific cod, Gadus macrocephalus. Fish Shellfish Immun 44:622–632
Mori K, Nakai T, Muroga K, Arimoto M, Mushiake K, Furusawa I (1992) Properties of a new virus belonging to Nodaviridae found in larval striped jack (Pseudocaranx dentex) with nervous necrosis. Virology 187:368–371
Mori K, Nakai T, Nagahara M, Muroga K, Mekuchi T, Kanno T (1991) A viral disease in hatchery-reared larvae and juveniles of redspotted grouper. Fish Pathol 26:209–210
Munday BL, Kwang J, Moody N (2002) Betanodavirus infections of teleost fish: a review. J Fish Dis 25:127–142
Munday BL, Nakai T (1997) Nodaviruses as pathogens in larval and juvenile marine finfish. World J Microb Biot 13:375–381
Nakai T, Nguyen HD, Nishizawa T, Muroga K, Arimoto M, Ootsuki K (1994) Occurrence of viral nervous necrosis in kelp grouper and tiger puffer. Fish Pathol 29:211–212
Nishizawa T, Furuhashi M, Nagai T, Nakai T, Muroga K (1997) Genomic classification of fish nodaviruses by molecular phylogenetic analysis of the coat protein gene. Appl Environ Microbiol 63:1633–1636
Nylund A, Karlsbakk E, Nylund S, Isaksen TE, Karlsen M, Korsnes K, Handeland S, Martinsen R, Mork Pedersen T, Ottem KF (2008) New clade of betanodaviruses detected in wild and farmed cod (Gadus morhua) in Norway. Arch Virol 153:541–547
Panzarin V, Fusaro A, Monne I, Cappellozza E, Patarnello P, Bovo G, Capua I, Holmes EC, Cattoli G (2012) Molecular epidemiology and evolutionary dynamics of betanodavirus in southern Europe. Infect Genet Evol 12:63–70
Parameswaran V, Kumar SR, Ahmed VPI, Hameed ASS (2008) A fish nodavirus associated with mass mortality in hatchery-reared Asian sea bass, Lates calcarifer. Aquaculture 275:366–369
Patel S, Korsnes K, Bergh O, Vik-Mo F, Pedersen J, Nerland AH (2007) Nodavirus in farmed Atlantic cod Gadus morhua in Norway. Dis Aquat Organ 77:169–173
Patel S, Nerland AH (2014) Vaccination against diseases caused by betanodavirus. Fish Vaccin (in press)
Renault T, Haffner P, Baudin Laurencien F, Breuil G, Bonami JR (1991) Mass mortalities in hatchery-reared sea bass (Lates calcarifer) larvae associated with the presence in the brain and retina of virus-like particles. Bull Eur Assoc Fish Pathol 11:68–73
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tan C, Huang B, Chang SF, Ngoh GH, Munday B, Chen SC, Kwang J (2001) Determination of the complete nucleotide sequences of RNA1 and RNA2 from greasy grouper (Epinephelus tauvina) nervous necrosis virus, Singapore strain. J Gen Virol 82:647–653
Toffolo V, Negrisolo E, Maltese C, Bovo G, Belvedere P, Colombo L, Valle LD (2007) Phylogeny of betanodaviruses and molecular evolution of their RNA polymerase and coat proteins. Mol Phylogenet Evol 43:298–308
Ucko M, Colorni A, Diamant A (2004) Nodavirus infections in Israeli mariculture. J Fish Dis 27:459–469
Valle LD, Negrisolo E, Patarnello P, Zanella L, Maltese C, Bovo G, Colombo L (2001) Sequence comparison and phylogenetic analysis of fish nodaviruses based on the coat protein gene. Arch Virol 146:1125–1137
Wang W, Jiang ZQ, Meng FP, Li Y, Wang ZY (2012) The effects of sharply changes in temperature on survival and indices of physiology and biochemistry in Pacific cod Gadus macrocephalus. Fish Sci 31:463–466
Wang XL (2006) Characterization and expression of flounder recombination activating gene (rag). Chinese Academy of Sciences, Qingdao
Willett CE, Cherry JJ, Steiner LA (1997) Characterization and expression of the recombination activating genes (rag1 and rag2) of zebrafish. Immunogenetics 45:394–404
Acknowledgments
This work was supported by the National Natural Science Foundation of China (NSFC-31302202), the National High Technology Research and Development Programme of China (2012AA10A413), the Open Project of Key Laboratory of Mariculture and Stock Enhancement in North Chin’s Sea, Ministry of Agriculture (2014-MSENC-KF-12), the General Project of the Education Department of Liaoning Province (L2013276), and Dalian Talents Introduction Project (DLYJ201311). All fieldwork in this study complied with the current laws of China, where it was performed.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mao, MG., Wen, SH., Perálvarez-Marín, A. et al. Evidence for and characterization of nervous necrosis virus infection in Pacific cod (Gadus macrocephalus). Arch Virol 160, 2237–2248 (2015). https://doi.org/10.1007/s00705-015-2484-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-015-2484-1