Abstract
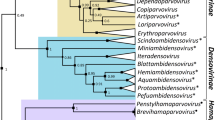
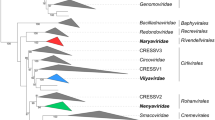
The nucleocytoplasmic large DNA viruses (NCLDVs) comprise a monophyletic group of viruses that infect animals and diverse unicellular eukaryotes. The NCLDV group includes the families Poxviridae, Asfarviridae, Iridoviridae, Ascoviridae, Phycodnaviridae, Mimiviridae and the proposed family “Marseilleviridae”. The family Mimiviridae includes the largest known viruses, with genomes in excess of one megabase, whereas the genome size in the other NCLDV families varies from 100 to 400 kilobase pairs. Most of the NCLDVs replicate in the cytoplasm of infected cells, within so-called virus factories. The NCLDVs share a common ancient origin, as demonstrated by evolutionary reconstructions that trace approximately 50 genes encoding key proteins involved in viral replication and virion formation to the last common ancestor of all these viruses. Taken together, these characteristics lead us to propose assigning an official taxonomic rank to the NCLDVs as the order “Megavirales”, in reference to the large size of the virions and genomes of these viruses.

Similar content being viewed by others
References
Iyer LM, Aravind L, Koonin EV (2001) Common origin of four diverse families of large eukaryotic DNA viruses. J Virol 75:11720–11734. doi:10.1128/JVI.75.23.11720-11734.2001
Iyer LM, Balaji S, Koonin EV, Aravind L (2006) Evolutionary genomics of nucleo-cytoplasmic large DNA viruses. Virus Res 117:156–184. doi:10.1016/j.virusres.2006.01.009
Yutin N, Wolf YI, Raoult D, Koonin EV (2009) Eukaryotic large nucleo-cytoplasmic DNA viruses: clusters of orthologous genes and reconstruction of viral genome evolution. Virol J 6:223. doi:10.1186/1743-422X-6-223
Raoult D, Audic S, Robert C, Abergel C, Renesto P, Ogata H, La Scola B, Suzan M, Claverie JM (2004) The 1.2-megabase genome sequence of Mimivirus. Science 306:1344–1350. doi:10.1126/science.1101485
Claverie JM, Abergel C, Ogata H (2009) Mimivirus. Curr Top Microbiol Immunol 328:89–121
La Scola B, Campocasso A, N’Dong R, Fournous G, Barrassi L, Flaudrops C, Raoult D (2010) Tentative characterization of new environmental giant viruses by MALDI-TOF mass spectrometry. Intervirology 53:344–353. doi:10.1159/000312919
Yoosuf N, Yutin N, Colson P, Shabalina SA, Pagnier I, Robert C, Azza S, Klose T, Wong J, Rossmann MG, La SB, Raoult D, Koonin EV (2012) Related giant viruses in distant locations and different habitats: Acanthamoeba polyphaga moumouvirus represents a third lineage of the Mimiviridae that is close to the megavirus lineage. Genome Biol Evol 4:1324–1330. doi:10.1093/gbe/evs109
Boyer M, Yutin N, Pagnier I, Barrassi L, Fournous G, Espinosa L, Robert C, Azza S, Sun S, Rossmann MG, Suzan-Monti M, La SB, Koonin EV, Raoult D (2009) Giant Marseillevirus highlights the role of amoebae as a melting pot in emergence of chimeric microorganisms. Proc Natl Acad Sci USA 106:21848–21853. doi:10.1073/pnas.0911354106
Thomas V, Bertelli C, Collyn F, Casson N, Telenti A, Goesmann A, Croxatto A, Greub G (2011) Lausannevirus, a giant amoebal virus encoding histone doublets. Environ Microbiol 13:1454–1466. doi:10.1111/j.1462-2920.2011.02446.x
Colson P, Pagnier I, Yoosuf N, Fournous G, La Scola B, Raoult D (2012) Marseilleviridae, a new family of giant viruses infecting amoebae. Arch Virol 158(4):915–920. doi:10.1007/s00705-012-1537-y
Claverie JM, Ogata H, Audic S, Abergel C, Suhre K, Fournier PE (2006) Mimivirus and the emerging concept of “giant” virus. Virus Res 117:133–144. doi:10.1016/j.virusres.2006.01.008
Arslan D, Legendre M, Seltzer V, Abergel C, Claverie JM (2011) Distant Mimivirus relative with a larger genome highlights the fundamental features of Megaviridae. Proc Natl Acad Sci USA 108:17486–17491
Fischer MG, Allen MJ, Wilson WH, Suttle CA (2010) Giant virus with a remarkable complement of genes infects marine zooplankton. Proc Natl Acad Sci USA 107:19508–19513. doi:10.1073/pnas.1007615107
Monier A, Claverie JM, Ogata H (2008) Taxonomic distribution of large DNA viruses in the sea. Genome Biol 9:R106. doi:10.1186/gb-2008-9-7-r106
Monier A, Larsen JB, Sandaa RA, Bratbak G, Claverie JM, Ogata H (2008) Marine mimivirus relatives are probably large algal viruses. Virol J 5:12. doi:10.1186/1743-422X-5-12
Yamada T (2011) Giant viruses in the environment: their origins and evolution. Curr Opin Virol 1:58–62
Kristensen DM, Mushegian AR, Dolja VV, Koonin EV (2010) New dimensions of the virus world discovered through metagenomics. Trends Microbiol 18:11–19. doi:10.1016/j.tim.2009.11.003
Colson P, de Lamballerie X, Fournous G, Raoult D (2012) Reclassification of giant viruses composing a fourth domain of life in the new order Megavirales. Intervirology 55(5):321–332. doi:10.1159/000336562
Yutin N, Koonin EV (2012) Hidden evolutionary complexity of nucleo-cytoplasmic large DNA viruses of eukaryotes. Virol J 9:161
Koonin EV, Yutin N (2010) Origin and evolution of eukaryotic large nucleo-cytoplasmic DNA viruses. Intervirology 53:284–292. doi:10.1159/000312913
Krupovic M, Bamford DH (2008) Virus evolution: how far does the double beta-barrel viral lineage extend? Nat Rev Microbiol 6:941–948
Bahar MW, Graham SC, Stuart DI, Grimes JM (2011) Insights into the evolution of a complex virus from the crystal structure of vaccinia virus D13. Structure 19:1011–1020
Bigot Y, Asgari S, Bideshi D, Cheng XW, Federici BA et al. (2011) Family Ascoviridae. In: CM Fauquet, Mayo MA, Maniloff J, Desselberger U, Ball LA (eds) Virus taxonomy. Ninth report of the international committee on taxonomy of viruses, San Diego, pp 147–152
Szajner P, Weisberg AS, Lebowitz J, Heuser J, Moss B (2005) External scaffold of spherical immature poxvirus particles is made of protein trimers, forming a honeycomb lattice. J Cell Biol 170:971–981
Seet BT, Johnston JB, Brunetti CR, Barrett JW, Everett H, Cameron C, Sypula J, Nazarian SH, Lucas A, McFadden G (2003) Poxviruses and immune evasion. Annu Rev Immunol 21:377–423
Werden SJ, Rahman MM, McFadden G (2008) Poxvirus host range genes. Adv Virus Res 71:135–171. doi:10.1016/S0065-3527(08)00003-1
Dixon LK, Abrams CC, Bowick G, Goatley LC, Kay-Jackson PC, Chapman D, Liverani E, Nix R, Silk R, Zhang F (2004) African swine fever virus proteins involved in evading host defence systems. Vet Immunol Immunopathol 100:117–134. doi:10.1016/j.vetimm.2004.04.002
Condit RC (2007) Vaccinia, Inc.—probing the functional substructure of poxviral replication factories. Cell Host Microbe 2:205–207
Mutsafi Y, Zauberman N, Sabanay I, Minsky A (2010) Vaccinia-like cytoplasmic replication of the giant Mimivirus. Proc Natl Acad Sci USA 107:5978–5982. doi:10.1073/pnas.0912737107
Netherton CL, Wileman TE (2013) African swine fever virus organelle rearrangements. Virus Res 173(1):76–86. doi:10.1016/j.virusres.2012.12.014
Novoa RR, Calderita G, Arranz R, Fontana J, Granzow H, Risco C (2005) Virus factories: associations of cell organelles for viral replication and morphogenesis. Biol Cell 97:147–172. doi:10.1042/BC20040058
Netherton CL, Wileman T (2011) Virus factories, double membrane vesicles and viroplasm generated in animal cells. Curr Opin Virol 1:381–387
Keeling PJ (2007) Genomics. Deep questions in the tree of life. Science 317:1875–1876
Keeling PJ, Burger G, Durnford DG, Lang BF, Lee RW, Pearlman RE, Roger AJ, Gray MW (2005) The tree of eukaryotes. Trends Ecol Evol 20:670–676
Koonin EV (2010) The origin and early evolution of eukaryotes in the light of phylogenomics. Genome Biol 11:209–211
Acknowledgments
Natalya Yutin and Eugene V. Koonin are supported by intramural funds of the US Department of Health and Human Services (to the National Library of Medicine). James Van Etten is partially supported by NIH Grant P20 RR15635 from the COBRE Program of the National Center for Research Resources.
Conflict of interest
There are no potential conflicts of interest or financial disclosures for any of the authors.
Author information
Authors and Affiliations
Corresponding author
Additional information
S. Asgari, Y. Bigot, D. K. Bideshi, X.-W. Cheng and B. A. Federici belong to the Ascoviridae study group of the International Committee on Taxonomy of Viruses (ICTV).
Rights and permissions
About this article
Cite this article
Colson, P., De Lamballerie, X., Yutin, N. et al. “Megavirales”, a proposed new order for eukaryotic nucleocytoplasmic large DNA viruses. Arch Virol 158, 2517–2521 (2013). https://doi.org/10.1007/s00705-013-1768-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-013-1768-6