Abstract
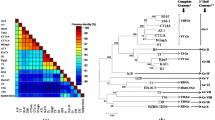
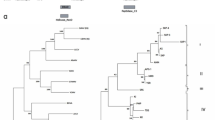
The complete genome sequence of a mandarin (Citrus reticulata) decline CTV isolate, Kpg3, of the Darjeeling hills of the Northeastern Himalayan region of India is reported for the first time. The complete Kpg3 genome has 19253 nt, and its nucleotide sequence identity ranged from 79% with the Florida CTV isolate T36 to 94% with the Israel isolate VT, whereas its identity to B165, the other Indian isolate, was 89%. Phylogenetic analysis indicated that the Kpg3 genome is closely related to isolate VT and distantly to T36 and B165. Recombination analysis indicated that Kpg3 is recombinant and originated through multiple recombination events in which parts of the genome were exchanged between divergent CTV sequences.

Similar content being viewed by others
References
Albiach-Marti MR, Mawassi M, Gowda S, Satyanayanana T, Hilf ME, Shanker S, Almira EC, Vives MC, Lopez C, Guerri J, Flores R, Moreno P, Garnsey SM, Dawson WO (2000) Sequences of Citrus tristeza virus separated in time and space are essentially identical. J Virol 74:6856–6865
Ahlawat YS (1997) Viruses, greening bacterium and viroids associated with citrus (Citrus species) decline in India. Indian J Agric Sci 67:51–57
Bar-Joseph M, Dawson WO (2008) Citrus tristeza virus. Encycl Virol 1:520–525
Bar-Joseph M, Marcus R, Lee RF (2002) The continuous challenge of citrus tristeza virus control. Ann Rev Phytopathol 27:291–316
Biswas KK (2010) Molecular characterization of Citrus tristeza virus isolates from the Northeastern Himalayan region of India. Arch Virol 155:959–963
Biswas KK (2008) Molecular diagnosis of Citrus tristeza virus in mandarin (Citrus reticulata) orchards of hills of West Bengal. Indian J Virol 19:26–31
Ghosh SP (2007) Citrus fruits. ICAR, New Delhi
Harper SJ, Dawson TE, Pearson MN (2010) Isolates of Citrus tristeza virus that overcome Poncirus trifoliata resistance comprise a novel strain. Arch Virol 155:471–480
Hilf ME, Mavrodieva VA, Garnsey SM (2005) Genetic marker analysis of a global collection of isolates of Citrus tristeza virus: characterization and distribution of CTV genotypes and association with symptoms. Phytopathology 95:909–917
Huson DH (1998) Splits tree: analyzing and visualizing evolutionary data. Bioinformatics 14:68–73
Karasev AV, Boyko VP, Gowda S, Nikolaeva OV, Hilf ME, Koonin EV, Cline K, Gumpf DJ, Lee RF, Garnsey SM, Lewandowski DJ, Dawson WO (1995) Complete sequences of the citrus tristeza virus RNA genome. Virology 208:511–520
Martin DP, Vander Walt E, Posada D (2005) RDP2: Recombination detection and analysis from sequence alignments. Bioinformatics 21:260–262
Martin S, Sambade A, Rubio L, Vives MC, Moya P, Guerri J, Elena SF, Moreno P (2009) Contribution of recombination and selection to molecular evolution of Citrus tristeza virus. J Gen Virol 90:1527–1538
Melzer MJ, Borth WB, Sether DM, Ferreira S, Gonsalves D, Hu JS (2010) Genetic diversity and evidence for recent modular recombination in Hawaiian Citrus tristeza virus. Virus Genes 40:111–118
Moreno P, Ambros S, Albiach-Marti MR, Guerri J, Pena L (2008) Citrus tristeza virus: a pathogen that changed the course of the citrus industry. Mol Plant Pathol 9:251–268
Mukhopadhyay S, Mukhopadhyay K, Mukherjee N, Ghosh MR, Bose TK, Choudhury AK, Chakroborty NK, Dutta S, Konar A, Gurung A (1986) The state of mandarin orange cultivation in Darjeeling District. In: Raychouduri SP, Verma JP (eds) Review of tropical plant pathology. vol II. pp 35–53
Nicholas KB, Nicholas HB Jr (1997) GeneDoc: a tool for editing and annotating multiple sequence alignments. http://www.psc.edu/biomed/genedoc
Roy A, Manjunath KL, Brlansky RH (2005) Assessment of sequence diversity in the 5′ terminal region of Citrus tristeza virus from India. Virus Res 113:132–142
Roy A, Brlansky RH (2010) Genome of an orange stem pitting Citrus tristeza virus isolate reveals a novel recombinant genotype. Virus Res. doi:101016/jvirusres201003017
Satyanayanana T, Gowda S, Mawassi M, Albiach-Marti MR, Ayllon MA, Robertson C, Garnsey SM, Dawson WO (2000) Closterovirus encoded HSP70 homolog and p61 in addition to both coat proteins function in efficient virion assembly. Virology 278:253–265
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The clustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–4882
Acknowledgments
The authors are grateful to Richard Lee and K.L. Manjunath, USDA-ARS, NCGRCD, Riverside, for teaching HMA analysis and for their valuable suggestions. This work was supported by Research Project (Code No. 24-33), Department of Biotechnology, Govt. of India. We are also thankful to Dr. R. K. Jain, Head, Division of Plant Pathology; Dr. V. G. Malathi, Ex-In-charge, ACPV; Director, Indian Agricultural Research Institute (IARI), New Delhi for providing financial and laboratory support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Biswas, K.K., Tarafdar, A. & Sharma, S.K. Complete genome sequence of mandarin decline Citrus tristeza virus of the Northeastern Himalayan hill region of India: comparative analyses determine recombinant. Arch Virol 157, 579–583 (2012). https://doi.org/10.1007/s00705-011-1165-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-011-1165-y