Abstract
The Hawaiian Islands are home to a widespread and diverse population of Citrus tristeza virus (CTV), an economically important pathogen of citrus. In this study, we quantified the genetic diversity of two CTV genes and determined the complete genomic sequence for two strains of Hawaiian CTV. The nucleotide diversity was estimated to be 0.0565 ± 0.0022 for the coat protein (CP) gene (n = 137) and 0.0822 ± 0.0033 for the p23 gene (n = 30). The genome size and organization of CTV strains HA18-9 and HA16-5 were similar to other fully sequenced strains of CTV. The 3′-terminal halves of their genomes were nearly identical (98.5% nucleotide identity), whereas the 5′-terminal halves were more distantly related (72.3% nucleotide identity), suggesting a possible recombination event. Closer examination of strain HA16-5 indicated that it arose through recent recombination between the movement module of an HA18-9 genotype, and the replication module of an undescribed CTV genotype.


Similar content being viewed by others
References
H.R. Pappu, A.V. Karasev, E.J. Anderson, S.S. Pappu, M.E. Hilf, V. Febres, R.M.J. Eckloff, M. McCaffery, V. Boyko, S. Gowda, V.V. Dolja, E.V. Koonin, D.J. Gumpf, K. Cline, S.M. Garnsey, W.O. Dawson, R.F. Lee, C.L. Niblett, Virology 199, 35 (1994)
A.V. Karasev, V.P. Boyko, S. Gowda, O.V. Nikolaeva, M.E. Hilf, E.V. Koonin, C.L. Niblett, K. Cline, D.J. Gumpf, R.F. Lee, S.M. Garnsey, D.J. Lewandowski, W.O. Dawson, Virology 208, 511 (1995)
V.V. Dolja, J.F. Kreuze, J.P.T. Valkonen, Virus Res. 117, 38 (2006)
P. Moreno, S. Ambrós, M.R. Albiach-Martí, J. Guerri, L. Peña, Mol. Plant Pathol. 9, 251 (2008)
M.R. Albiach-Marti, M. Mawassi, S. Gowda, T. Satyanarayana, M.E. Hilf, S. Shanker, E.C. Almira, M.C. Vives, C. Lopez, J. Guerri, R. Flores, P. Moreno, S.M. Garnsey, W.O. Dawson, J. Virol. 74, 6856 (2000)
M.E. Hilf, A.V. Karasev, M.R. Albiach-Marti, W.O. Dawson, S.M. Garnsey, Phytopathology 89, 336 (1999)
M. Mawassi, E. Mietkiewska, R. Gofman, G. Yang, M. Bar-Joseph, J. Gen. Virol. 77, 2359 (1996)
D.C. Giacometti, W.B. Storey, Cal. Citrograph 37, 357 (1952)
M.J. Melzer, W.B. Borth, F. Zee, M.E. Hilf, S.M. Garnsey, J.S. Hu, in Proceedings of the 16th Conference of the International Organization of Citrus Virologists, ed. by M.E. Hilf, N. Duran-Vila, M.A. Rocha-Peña (IOCV, Riverside, CA, 2005), p. 179
M.E. Hilf, S.M. Garnsey, in Proceedings of the 14th Conference of the International Organization of Citrus Virologists, ed. by J.V. da Graça, R.F. Lee, R.K. Yokomi (IOCV, Riverside, CA, 2000), p. 18
P. Froussard, Nucleic Acids Res. 20, 2900 (1992)
H. Telenius, N.P. Carter, C.E. Bebb, M. Nordenskjöld, B.A. Ponder, A. Tunnacliffe, Genomics 13, 718 (1992)
J. Rozas, J.C. Sánchez-DelBarrio, X. Messeguer, R. Rozas, Bioinformatics 19, 2496 (2003)
C.N. Stewart Jr., L.E. Via, BioTechniques 14, 748 (1993)
X. Huang, A. Madan, Genome Res. 9, 868 (1999)
C. López, M.A. Ayllón, J. Navas-Castillo, J. Guerri, P. Moreno, R. Flores, Phytopathology 88, 685 (1998)
J.D. Thompson, T.J. Gibson, F. Plewniak, F. Jeanmougin, D.G. Higgins, Nucleic Acids Res. 24, 4876 (1997)
J.P. Huelsenbeck, F. Ronquist, Bioinformatics 17, 754 (2001)
F. Ronquist, J.P. Huelsenbeck, Bioinformatics 19, 1572 (2003)
M.E. Rott, W. Jelkmann, Eur. J. Plant Pathol. 107, 411 (2001)
W. Benthack, N. Mielke, C. Büttner, H.P. Muhlbach, Arch. Virol. 150, 37–53 (2004)
M.E. Hilf, in Proceedings of the 16th Conference of the International Organization of Citrus Virologists, ed. by M.E. Hilf, Duran-Vila, M.A. Rocha-Peña (IOCV, Riverside, CA, 2005), p. 52
L. Rubio, M.A. Ayllón, P. Kong, A. Fernandez, M. Polek, J. Guerri, B.W. Falk, J. Virol. 75, 8054 (2001)
M.C. Vives, L. Rubio, C. López, J. Navas-Castillo, M.R. Albiach-Marti, W.O. Dawson, J. Guerri, R. Flores, P. Moreno, J. Gen. Virol. 80, 811 (2005)
Z. Weng, R. Barthelson, S. Gowda, M.E. Hilf, W.O. Dawson, D.W. Galbraith, Z. Xiong, PLoS ONE 2 (2007). doi: https://doi.org/10.1371/journal.pone.0000917
P.D. Nagy, in Plant virus evolution, ed. by M. Roossinck (Springer, Berlin, 2008), p. 133
M.A. Ayllón, S. Gowda, T. Satyanarayana, W.O. Dawson, Virology 322, 41 (2004)
X. Che, D. Piestun, M. Mawassi, G. Yang, T. Satyanarayana, S. Gowda, W.O. Dawson, M. Bar-Joseph, Virology 283, 374 (2001)
M.R. Albiach-Marti, J.W. Grosser, S. Gowda, M. Mawassi, T. Satyanarayana, S.M. Garnsey, W.O. Dawson, Mol. Breed. 14, 117 (2004)
S.Y. Folimonova, A.S. Folimonov, S. Tatineni, W.O. Dawson, J. Virol. 82, 6546 (2008)
Acknowledgments
The research was funded, in part, by grants from USDA-CSREES T-STAR program agreement #2003-34135-13982 and a specific cooperative grant agreement #58-5320-5-785, between the USDA-ARS Pacific Basin Agricultural Research Center and the University of Hawaii at Manoa.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
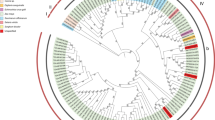
Fig S1
Neighbor-joining phylogram illustrating the genetic diversity of Citrus tristeza virus (CTV) in Hawaii and around the world. Trees were generated using the amino acid translation of 123 sequences from Hawaii and 28 foreign sequences. Hawaiian sequences are color-coordinated by island and foreign sequences are labeled by CTV isolate/clone, country of origin, and GenBank accession number (in brackets). Identical sequences are listed horizontally at the terminus of each branch. KA, Kauai; OA, Oahu; MO, Molokai; MA, Maui; HA, the Big Island of Hawaii (PDF 88 kb)
Rights and permissions
About this article
Cite this article
Melzer, M.J., Borth, W.B., Sether, D.M. et al. Genetic diversity and evidence for recent modular recombination in Hawaiian Citrus tristeza virus . Virus Genes 40, 111–118 (2010). https://doi.org/10.1007/s11262-009-0409-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-009-0409-3