Abstract
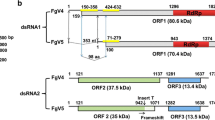
The complete genomes two different dsRNA mycoviruses, Fusarium graminearum virus 3 (FgV3) and Fusarium graminearum virus 4 (FgV4), was sequenced and analyzed. The viral genome of FgV3 is 9,098 base pairs (bp) long and contains two open reading frames (ORF) encoding a putative RNA-dependent RNA polymerase (RdRp) and a protein of unknown function. The FgV4 genome is composed of two dsRNA genome segments of 2,383 bp and 1,739 bp. FgV4 dsRNA-1 contains a single ORF, which has a conserved RdRp motif, while FgV4 dsRNA-2 contains two putative ORFs coding for products of unknown function. Both the genome organization and phylogenetic analysis indicated that FgV3 was closely related to members of the families Totiviriridae and Chrysoviridae, but it was placed outside of their main clusters, whereas FgV4 formed a distinct clade with the family Partitiviridae. This is the first report of the full-length nucleotide sequences of FgV3 and FgV4 infecting Fusarium graminearum.


Similar content being viewed by others
References
Aoki N, Moriyama H, Kodama M, Arie T, Teraoka T, Fukuhara T (2009) A novel mycovirus associated with four double-stranded RNAs affects host fungal growth in Alternaria alternata. Virus Res 140:179–187
Bruenn JA (1993) A closely related group of RNA-dependent RNA polymerases from double-stranded RNA viruses. Nucleic Acid Res 21:5667–5669
Chu Y-M, Jeon J-J, Yea S-J, Kim Y-H, Yun S–H, Lee Y-W, Kim K-H (2002) Double-stranded RNA mycovirus from Fusarium graminearum. Appl Environ Microbiol 68:2529–2534
Chu Y-M, Lim W-S, Yea S-J, Cho J-D, Lee Y-W, Kim K-H (2004) Complexity of dsRNA mycovirus isolated from Fusarium graminearum. Virus Genes 28:135–143
Cold spring harbor laboratory press (2005) Rapid amplification of 5’ complementary DNA ends (5’ RACE). Nat Methods 2:629–630
Cook RJ (1981) Fusarium diseases of wheat and other small grains in North America. In: Nelson PE, Toussoun TA, Cook RJ (eds) Fusarium diseases, biology, and taxonomy. The Pennsylvania State University Press, University Park, pp 39–52
Jiang D, Ghabrial SA (2004) Molecular characterization of Penicillium chrysogenum virus: reconsideration of the taxonomy of the genus Crysovirus. J Gen Virol 85:2111–2121
Kommedahl T, Windels CE (1981) Root-, stalk-, and ear-infecting Fusarium species on corn in the USA. In: Nelson PE, Toussoun TA, Cook RJ (eds) Fusarium diseases, biology, and taxonomy. The Pennsylvania State University Press, University Park, pp 94–103
Kumar S, Nei M, Dudley J, Tamura K (2008) MEGA: a biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief Bioinform 9:299–306
Kwon S-J, Cho S-Y, Lee K-M, Yu J, Sohn M-I, Kim K-H (2009) Proteomic analysis of fungal host factors differentially expressed by Fusarium graminearum infected with Fusarium graminearum virus-DK21. Virus Res 144:96–106
Kwon S-J, Lim W-S, Park S-H, Park M-R, Kim K-H (2007) Molecular characterization of a dsRNA Mycovirus, Fusarium graminearum virus-DK21, which is phylogenetically related to hypoviruses but has a genome organization and gene expression strategy resembling those of plant potex-like viruses. Mol Cells 23:304–315
Acknowledgments
This research was supported in part by grants from the Rural Development Administration (No. 20090101-080-092-001) and the Korea Science and Engineering Foundation grant (No. R11-2008-062-02003-0) funded by the Ministry of Education, Science, and Technology (MEST). JSY, KML, and MIS were supported by graduate research fellowships from the MEST through the Brain Korea 21 Project.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yu, J., Kwon, SJ., Lee, KM. et al. Complete nucleotide sequence of double-stranded RNA viruses from Fusarium graminearum strain DK3. Arch Virol 154, 1855–1858 (2009). https://doi.org/10.1007/s00705-009-0507-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-009-0507-5