Abstract
A competitive colorimetric assay has been established to detect chloramphenicol (CAP). It is based on the use of colloidal and electrostatically stabilized aptamer-modified gold nanoparticles (GNPs). The CAP aptamer is modified by a sequence of 5 adenosine groups to anchor it on the surface of GNPs. It can competitively capture two compounds, viz. D-(-)-threo-2-amino-1-(4-nitrophenyl)-1,3-propanediol (CAP-base, with a positive charge) and CAP (which is uncharged). The capture of the positively charged CAP-base triggers the aggregation of modified GNPs in salt-containing solution, and this causes a color change from red to purple. However, in the presence of CAP and CAP-base, the capture of the uncharged CAP weakens this color change by a competing process for capture. Thus, the concentration of CAP is associated with the degree of deaggregation of GNPs and can be quantified by the ratio of absorbances at 620 nm and 520 nm. The assay has a 22 nM limit of detection in acidic solution, and the response is linear in the range of 0.20 to 3.20 μM CAP concentration. This assay was successfully applied to the determination of CAP in spiked environmental water samples. Conceivably, this method has a wide scope in that it may be applied to a wide range of analytes if respective aptamers are available.

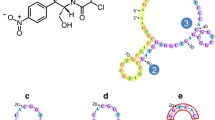
Schematic presentation of a competitive non-cross linking deaggregating method for detecting chloramphenicol. The surface charge of polyA-Apt@GNPs and its aggregation degree (purple) are determined by the charge of target. (CAP-base: precursor of CAP; PolyA-Apt@GNPs: 5′-polyA-modified DNA aptamer functionalized gold nanoparticles.)







Similar content being viewed by others
References
Lee JS, Ulmann PA, Han MS, Mirkin CA (2008) A DNA-gold nanoparticle-based colorimetric competition assay for the detection of cysteine. Nano Lett 8(2):529–533. https://doi.org/10.1021/nl0727563
Shahdordizadeh M, Yazdian-Robati R, Ansari N, Ramezani M, Abnous K, Taghdisi SM (2018) An aptamer-based colorimetric lead(II) assay based on the use of gold nanoparticles modified with dsDNA and exonuclease I. Microchim Acta 185(2):151. https://doi.org/10.1007/s00604-018-2699-4
Kim YS, Raston NH, Gu MB (2016) Aptamer-based nanobiosensors. Biosens Bioelectron 76:2–19. https://doi.org/10.1016/j.bios.2015.06.040
Mirkin CA, Letsinger RL, Mucic RC, Storhoff JJ (1996) A DNA-based method for rationally assembling nanoparticles into macroscopic materials. Nature 382(6592):607–609. https://doi.org/10.1038/382607a0
Wang G, Akiyama Y, Shiraishi S, Kanayama N, Takarada T, Maeda M (2017) Cross-linking versus non-cross-linking aggregation of gold nanoparticles induced by DNA hybridization: a comparison of the rapidity of solution color change. Bioconjug Chem 28(1):270–277. https://doi.org/10.1021/acs.bioconjchem.6b00410
Zhao W, Chiuman W, Lam JC, McManus SA, Chen W, Cui Y, Pelton R, Brook MA, Li Y (2008) DNA aptamer folding on gold nanoparticles: from colloid chemistry to biosensors. J Am Chem Soc 130(11):3610–3618. https://doi.org/10.1021/ja710241b
Pei H, Li F, Wan Y, Wei M, Liu H, Su Y, Chen N, Huang Q, Fan C (2012) Designed diblock oligonucleotide for the synthesis of spatially isolated and highly hybridizable functionalization of DNA-gold nanoparticle nanoconjugates. J Am Chem Soc 134(29):11876–11879. https://doi.org/10.1021/ja304118z
Li H, Rothberg L (2004) Colorimetric detection of DNA sequences based on electrostatic interactions with unmodified gold nanoparticles. Proc Natl Acad Sci U S A 101(39):14036–14039. https://doi.org/10.1073/pnas.0406115101
Sato K, Hosokawa K, Maeda M (2003) Rapid aggregation of gold nanoparticles induced by non-cross-linking DNA hybridization. J Am Chem Soc 125(27):8102–8103. https://doi.org/10.1021/ja034876s
Sai N, Chen Y, Liu N, Yu G, Su P, Feng Y, Zhou Z, Liu X, Zhou H, Gao Z, Ning BA (2010) A sensitive immunoassay based on direct hapten coated format and biotin-streptavidin system for the detection of chloramphenicol. Talanta 82(4):1113–1121. https://doi.org/10.1016/j.talanta.2010.06.018
Abnous K, Danesh NM, Ramezani M, Emrani AS, Taghdisi SM (2016) A novel colorimetric sandwich aptasensor based on an indirect competitive enzyme-free method for ultrasensitive detection of chloramphenicol. Biosens Bioelectron 78:80–86. https://doi.org/10.1016/j.bios.2015.11.028
Hanekamp JC, Bast A (2015) Antibiotics exposure and health risks: chloramphenicol. Environ Toxicol Pharmacol 39(1):213–220. https://doi.org/10.1016/j.etap.2014.11.016
Ding J, Li Q, Xu X, Zhang X, Su Y, Yue Q, Gao B (2018) A wheat straw cellulose-based hydrogel for Cu (II) removal and preparation copper nanocomposite for reductive degradation of chloramphenicol. Carbohydr Polym 190:12–22. https://doi.org/10.1016/j.carbpol.2018.02.032
Louie TJ, Tally FP, Bartlett JG, Gorbach SL (1976) Rapid microbiological assay for chloramphenicol and tetracyclines. Antimicrob Agents Chemother 9(6):874–878. https://doi.org/10.1128/aac.9.6.874
Chughtai MI, Maqbool U, Iqbal M, Shah MS, Fodey T (2017) Development of in-house ELISA for detection of chloramphenicol in bovine milk with subsequent confirmatory analysis by LC-MS/MS. J Environ Sci Health B 52(12):871–879. https://doi.org/10.1080/03601234.2017.1361771
Hussain A, Alajmi MF, Ali I (2016) Determination of chloramphenicol in biological matrices by solid-phase membrane micro-tip extraction and capillary electrophoresis. Biomed Chromatogr 30(12):1935–1941. https://doi.org/10.1002/bmc.3769
Luo L, Gu C, Li M, Zheng X, Zheng F (2018) Determination of residual 4-nitrobenzaldehyde in chloramphenicol and its pharmaceutical formulation by HPLC with UV/Vis detection after derivatization with 3-nitrophenylhydrazine. J Pharm Biomed Anal 156:307–312. https://doi.org/10.1016/j.jpba.2018.04.024
Chang GR, Chen HS, Lin FY (2016) Analysis of banned veterinary drugs and herbicide residues in shellfish by liquid chromatography-tandem mass spectrometry (LC/MS/MS) and gas chromatography-tandem mass spectrometry (GC/MS/MS). Mar Pollut Bull 113(1–2):579–584. https://doi.org/10.1016/j.marpolbul.2016.08.080
Zhang Y, Shao Y, Gao N, Gao Y, Chu W, Li S, Wang Y, Xu S (2018) Kinetics and by-products formation of chloramphenicol (CAP) using chlorination and photocatalytic oxidation. Chem Eng J 333:85–91. https://doi.org/10.1016/j.cej.2017.09.094
Miao YB, Gan N, Li TH, Zhang HR, Cao YT, Jiang QL (2015) A colorimetric aptasensor for chloramphenicol in fish based on double-stranded DNA antibody labeled enzyme-linked polymer nanotracers for signal amplification. Sensors Actuators B Chem 220:679–687. https://doi.org/10.1016/j.snb.2015.05.106
Chang CC, Wang G, Takarada T, Maeda M (2017) Iodine-mediated etching of triangular gold nanoplates for colorimetric sensing of copper ion and aptasensing of chloramphenicol. ACS Appl Mater Interfaces 9(39):34518–34525. https://doi.org/10.1021/acsami.7b13841
Duan Y, Gao Z, Wang L, Wang H, Zhang H, Li H (2016) Selection and identification of chloramphenicol-specific DNA Aptamers by Mag-SELEX. Appl Biochem Biotechnol 180(8):1644–1656. https://doi.org/10.1007/s12010-016-2193-6
Liu J, Lu Y (2006) Preparation of aptamer-linked gold nanoparticle purple aggregates for colorimetric sensing of analytes. Nat Protoc 1(1):246–252. https://doi.org/10.1038/nprot.2006.38
Liu X, Atwater M, Wang J, Huo Q (2007) Extinction coefficient of gold nanoparticles with different sizes and different capping ligands. Colloids Surf B: Biointerfaces 58(1):3–7. https://doi.org/10.1016/j.colsurfb.2006.08.005
Koo KM, Sina AAI, Carrascosa LG, Shiddiky MJA, Trau M (2015) DNA–bare gold affinity interactions: mechanism and applications in biosensing. Anal Methods UK 7(17):7042–7054. https://doi.org/10.1039/c5ay01479d
Li X, Cheng R, Shi H, Tang B, Xiao H, Zhao G (2016) A simple highly sensitive and selective aptamer-based colorimetric sensor for environmental toxins microcystin-LR in water samples. J Hazard Mater 304:474–480. https://doi.org/10.1016/j.jhazmat.2015.11.016
Sun Y, Wei T, Jiang M, Xu L, Xu Z (2018) Voltammetric sensor for chloramphenicol determination based on a dual signal enhancement strategy with ordered mesoporous carbon@polydopamine and β-cyclodextrin. Sensors Actuators B Chem 255:2155–2162. https://doi.org/10.1016/j.snb.2017.09.016
Liu S, Lai GS, Zhang HL, Yu AM (2017) Amperometric aptasensing of chloramphenicol at a glassy carbon electrode modified with a nanocomposite consisting of graphene and silver nanoparticles. Microchim Acta 184(5):1445–1451. https://doi.org/10.1007/s00604-017-2138-y
Wang Y, Bian F, Qin X, Wang Q (2018) Visible light photoelectrochemical aptasensor for chloramphenicol by using a TiO2 nanorod array sensitized with Eu(III)-doped CdS quantum dots. Microchim Acta 185(3):161. https://doi.org/10.1007/s00604-018-2711-z
Miao YB, Ren HX, Gan N, Zhou Y, Cao Y, Li T, Chen Y (2016) A homogeneous and “off-on” fluorescence aptamer-based assay for chloramphenicol using vesicle quantum dot-gold colloid composite probes. Anal Chim Acta 929:49–55. https://doi.org/10.1016/j.aca.2016.04.060
Li J, Shao B, Shen J, Wang S, Wu Y (2013) Occurrence of chloramphenicol-resistance genes as environmental pollutants from swine feedlots. Environ Sci Technol 47(6):2892–2897. https://doi.org/10.1021/es304616c
Liu H, Zhang G, Liu CQ, Li L, Xiang M (2009) The occurrence of chloramphenicol and tetracyclines in municipal sewage and the Nanming River, Guiyang City. J Environ Monit 11(6):1199–1205. https://doi.org/10.1039/b820492f
Gopinath SC, Lakshmipriya T, Awazu K (2014) Colorimetric detection of controlled assembly and disassembly of aptamers on unmodified gold nanoparticles. Biosens Bioelectron 51:115–123. https://doi.org/10.1016/j.bios.2013.07.037
Rayegan A, Allafchian A, Abdolhosseini Sarsari I, Kameli P (2018) Synthesis and characterization of basil seed mucilage coated Fe3O4 magnetic nanoparticles as a drug carrier for the controlled delivery of cephalexin. Int J Biol Macromol 113:317–328. https://doi.org/10.1016/j.ijbiomac.2018.02.134
Rimsza JM, Jones RE, Criscenti LJ (2018) Interaction of NaOH solutions with silica surfaces. J Colloid Interface Sci 516:128–137. https://doi.org/10.1016/j.jcis.2018.01.049
Acknowledgements
This work is sponsored by the National Natural Science Foundation of China (Grant No. 21707137), the Key Application and Development Program of Chongqing (No. cstc2017shmsA0159), Natural Science Foundation of Chongqing (Grant No. cstc2018jcyjAX0718), Research Funding Project of Yangtze Normal University (No. 2017KYQD23), Young Scientist Research Fund of Yangtze Normal University, Open Research Fund Program of Key Laboratory of Reservoir Aquatic Environment of CAS.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare that they have no competing interests.
Electronic supplementary material
ESM 1
(DOCX 4.68 kb)
Rights and permissions
About this article
Cite this article
Xie, Y., Huang, Y., Tang, D. et al. A competitive colorimetric chloramphenicol assay based on the non-cross-linking deaggregation of gold nanoparticles coated with a polyadenine-modified aptamer. Microchim Acta 185, 534 (2018). https://doi.org/10.1007/s00604-018-3067-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-018-3067-0