Abstract
This paper presents a novel bio-inspired optimization algorithm called Coronavirus Optimization Algorithm (COVIDOA). COVIDOA is an evolutionary search strategy that mimics the mechanism of coronavirus when hijacking human cells. COVIDOA is inspired by the frameshifting technique used by the coronavirus for replication. The proposed algorithm is tested using 20 standard benchmark optimization functions with different parameter values. Besides, we utilized five IEEE Congress of Evolutionary Computation (CEC) benchmark test functions (CECC06, 2019 Competition) and five CEC 2011 real-world problems to prove the proposed algorithm's efficiency. The proposed algorithm is compared to eight of the most popular and recent metaheuristic algorithms from the state-of-the-art in terms of best cost, average cost (AVG), corresponding standard deviation (STD), and convergence speed. The results demonstrate that COVIDOA is superior to most existing metaheuristics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Nature is full of principles and mechanisms that inspire scientists to develop complex computational problems [15]. Researchers developed various renature-inspired algorithms such as Genetic Algorithm (GA) [26] and Differential Evolution (DE) [63] over the years. These algorithms are based on the theory of natural evolution. Another group of Algorithms mimics the behavior of birds, animals, insects, plants, or fish, such as Particle Swarm Optimization (PSO) [44], Artificial Bee Colony (ABC) [41], Chicken Swarm Optimization (CSO) [48], Flower Pollination Algorithm (FPA) [75], Grey Wolf Optimization (GWO) [51], Whale Optimization Algorithm (WOA) [50], Cuckoo Search (CS) [76], Bird Mating Optimizer [6], Social Spider Optimization (SSO) [38], Krill Herd [25], and Seagull Optimization Algorithm (SOA) [19]. Other algorithms based on physical phenomena such as Water Cycle Algorithm (WCA) [22], Central Force Optimization (CFO) [24], Gravitational Search Algorithm (GSA) [60], Water Wave Optimization (WWO) [78], and Gradient-based Optimizer (GBO) (Ahmadianfar et al. 2020). Many other optimization algorithms are proposed by [5, 7, 10, 11, 13, 17, 28, 42, 47, 53, 55,56,57, 61, 62, 64, 66, 69].
Generally speaking, optimization algorithms are classified into three categories: swarm-based, physics-based, and evolutionary algorithms. Swarm-based algorithms such as ABC, PSO, CSO, and CS, mimic how a group of agents would behave with each other and their environment [1]. Based on Newton's gravitational law, physics-based algorithms are based on a mathematical idea or physical processes, such as CFO and GSA [3]. On the other hand, evolutionary algorithms are search methods inspired by biological evolution mechanisms, such as reproduction and mutation [77]. The most popular evolutionary algorithm is GA, inspired by Darwin's theory of biological evolution. As mentioned in [23], evolutionary algorithms have some advantages over other types of optimization algorithms, such as:
-
(1)
They are conceptually simple: all evolutionary algorithms have similar necessary steps: initialization, fitness evaluation, selection, crossover, and mutation.
-
(2)
In evolutionary algorithms, the individuals with the highest fitness are selected for reproduction, leading to new individuals' production closer to the optimum solution.
-
(3)
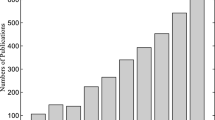
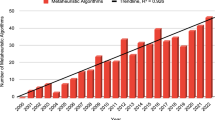
Broad applicability: researchers can apply evolutionary algorithms to any problem formulated in the form of an optimization function. A list of the most popular nature-inspired algorithms is shown in Fig. 1.
Since 2020, the world has suffered from the pandemic of coronavirus disease 2019 (COVID-19). Researchers worldwide are doing their best to understand this novel virus's mechanism and find an effective therapy for this disease [40, 73]. More than one researcher discussed the mechanism of the novel Coronavirus from different perspectives in the optimization field. The authors in [47] proposed a bio-inspired metaheuristic algorithm based on the propagation model of coronavirus, and the experimental results showed quite remarkable performance of the algorithm. Al-Betal et al. [4] proposed an optimization algorithm based on herd immunity's effect in tackling the COVID pandemic. The comparative analysis showed that the proposed algorithm yields very competitive results compared to other well-established methods. Another algorithm [27] models the coronavirus distribution process as an optimization problem to minimize the number of COVID-19 infected countries and slow the epidemic.
Once the virus is inside the human body, the most severe problem is replication and transcription, in which new copies of the virus are created and target new healthy cells [49, 71]. This paper presents a novel evolutionary optimization algorithm named Coronavirus Disease Optimization Algorithm (COVIDOA).
COVIDOA mimics the attacking behavior of coronavirus inside human cells. It is worth mentioning that almost all kinds of viruses have the same general steps for replication: entry, uncoating, replication, assembly, and virion release. However, replication between viruses greatly varies depending on the genes involved [20].
In addition to the advantages of evolutionary algorithms, COVIDOA has several advantages when compared with other similar mechanisms:
-
1.
Based on the virus's novelty and the lack of research on its various aspects, the reported numerical data about the coronavirus lifecycle may be inaccurate. Therefore, the proposed algorithm parameters, such as the number of virus particles in each generation and the number of viral proteins generated by each particle, haven't been set at fixed values. These reasons give the researchers’ flexibility to use the extendable values for the controlling parameters that most fit according to their problem.
-
2.
As mentioned in [8], the mutation rate of coronavirus is 1 × 10–6, which is very low; however, the mutation rate in the proposed algorithm is set at a larger value in the range [0.1 0.001], which helps in exploring new promising regions and avoid getting stuck in a local minimum.
-
3.
This study simulates a different virus replication technique known as the frameshifting technique [12, 43]. The virus uses frameshifting to create more copies of itself, leading to large-scale changes to polypeptide length and chemical composition. It is considered the most harmful to the molecular evolution of human cell proteins resulting in a non-functional protein that often disrupts the biochemical processes of a cell [59]. Applying the frameshifting technique in the proposed algorithm helps update solutions so that the solutions in each generation will not become too similar, which would allow the algorithm to converge to the global minimum.
The rest of the paper is structured as follows. Section 2 describes the inspiration and mathematical model of the proposed algorithm (COVIDOA). Experiments using test benchmark functions and the obtained results are discussed in Sect. 3. Finally, this study's conclusion and future work are presented in Sect. 4.
2 Proposed algorithm
In this section, the inspiration and mathematical model of COVIDOA are presented.
2.1 Inspiration
The new Coronavirus disease (COVID-19) is an infectious respiratory disease caused by Ssevere Aacute Respiratory Syndrome-CoronaVirus-2 (SARS-CoV-2). SARS-CoV-2 belongs to the coronaviruses family, named for crown-like spikes on their surface [34, 52], as shown in Fig. 2.
Structural proteins of COVID-19 (https://commons.wikimedia.org/wiki/File:3D_medical_animation_corona_virus.jpg) [30]
The coronavirus consists of a set of genetic instructions inside an oily membrane. These instructions are encoded in 30,000 letters of Ribonucleic Acid (RNA)—a, c, g, and u—then read by the infected cell and translated into many types of virus proteins [8]. Like other Coronaviruses, SARS-CoV-2 (COID-19) has four structural proteins, including the spike (S), envelope (E), and membrane (M) that constitute the viral coat, and the nucleocapsid (N) protein, which encapsulates the viral RNA [12]. Human-to-human transmission of SARS-CoV-2 occurs primarily via respiratory droplets from coughs and sneezes. Complications may include acute respiratory distress syndrome (ARDS), multi-organ failure, septic shock, and death [43].
The most serious problem of the virus is rapid replication, where it creates millions of copies of itself and sends it out to damage as many as possible human healthy cells. The replication mechanism of coronavirus inspires the proposed algorithm. For the virus to replicate, it passes through several stages as follows:
2.1.1 Virus entry and uncoating
For replication, coronavirus needs to use the human cell's protein-making machinery. So, it first needs to gain entry into the cell. The virus contains a set of spike (S) proteins; it uses its spike proteins as a key to getting inside a human cell [9, 72]. One spike of the virus binds to a protein called angiotensin-converting enzyme 2 (ACE-2) [67] on the surface of some human cells, as shown in Fig. 3. Coronavirus has a sort of membrane that hides its genetic material from the outside world; human cells have the same membrane that hides their material from the outside world. So, when those two things come together, the virus must find a way to get inside the host cell [65]. Once inside, all structural proteins are removed, and the virus contents, the genomic RNA, will be released into the host cell cytoplasm. This process is called virus uncoating [74], as shown in Fig. 4.
Virus attachment to human cell through spike protein (https://time.com/5839932/how-remdesivir-works-coronavirus/) [31]
2.1.2 Virus replication
Suppose the virus is getting fused in the cell membrane. Its small genetic material must hijack big cellular machinery in the next step. It will be tedious if the virus has few proteins to hijack the cell. The virus genome starts to find something in the host cell called a ribosome [79], a ribosome turns the virus RNA into many virus proteins through the ribosomal frameshifting technique [36], as shown in Fig. 5.
2.1.2.1 Ribosomal frameshifting during genome translation
Ribosomal frameshifting is also known as translational frameshifting, a biological phenomenon that occurs during translation [36, 54]. This phenomenon creates multiple unique proteins from a single messenger RNA (mRNA) molecule [14]. The translation is when the mRNA (messenger Ribonucleic Acid) molecule provides information to ribosomes, leading to protein molecules' formation [36, 37]. At the same time, frameshifting is when a specific reading frame of RNA molecule shifts to another reading frame to provide a new protein sequence [67, 72, 74]. To understand this, we need to understand translation and frameshifting separately.
The frameshifting technique is presented in Fig. 6. As shown in the figure, in the replication process, the virus's mRNA is translated into viral proteins by reading tri-nucleotides (e.g., ACG). Each tri-nucleotide is translated into single amino acid [52]. Thus, shifting (backward or forward) the reading frame of the nucleotides sequence by any number (not divisible by 3) will create different sequences that will be translated into different viral proteins [68].
Each group of the newly created viral proteins is merged to form a new virion. According to this technique, the virus can create millions of new particles than will damage millions of human cells.
In a translating ribosome, a frameshifting can result in either a nonsense mutation [68, 72] or a new protein after the frameshift. The most common types of frameshifting are − 1 frameshifting and + 1 frameshifting [58].
-
A.
− 1 Frameshifting
In − 1 frameshifting, the ribosome slips back one nucleotide (RNA letter) and continues translation in the − 1 frame, as shown in Fig. 7a.
-
B.
+ 1 Frameshifting
Different examples of frameshifting technique a − 1 frameshifting, b + 1 frameshifting, where E, P, and A, are the first, second, and third binding sites for RNA in the ribosome [14]
The ribosome starts translation from the + 1 frame when 0 is the initial position, as shown in Fig. 7b. Because of shifting, the sequences are read differently and translated into different proteins.
2.1.2.2 Synthesis of both genomic and subgenomic RNA species
The ribosomal frameshifting technique results in two types of RNAs, genomic RNA, and subgenomic RNAs. Genomic RNA is produced through the replication process and becomes the genome of the new virus particle. At the same time, Subgenomic RNAs are translated into many structural proteins (S: spike protein, E: envelope protein, M: membrane protein, and N: nucleocapsid protein). The genomic RNA and subgenomic RNAs are combined to form a viral particle [45, 58]. Finally, the new virion is released, trying to hijack new healthy cells, Fig. 8.
Release of the new virion. (https://time.com/5839932/how-remdesivir-works-coronavirus/)
2.1.3 Virus mutation
As coronaviruses spread from person to person, they randomly accumulate more mutations to escape from the immune system [45]. Mutations involve changing one or more letters that represent the virus genome. As mentioned in [8], coronavirus has lower mutation rates (≈10−6 per site per cycle) in comparison with influenza (≈3 × 10−5 per site per cycle). The replication stages of coronavirus are summarized in Fig. 9.
2.2 Mathematical model of COVIDOA
In this section, the mathematical model of COVIDOA is provided. COVIDOA is summarized in the following steps:
-
1.
Initialization population of solutions is randomly initialized, and the cost is evaluated for each solution. The solutions are then ordered ascendingly according to the fitness function, and the first solution is considered the best solution.
-
2.
Virus replication phase through frameshifting technique for each solution in the population, a parent is selected using roulette wheel selection [46] then,
-
a.
The frameshifting technique is applied to produce several proteins from the selected parent as follows:
-
b.
For each protein:
-
i.
If the + 1 frameshifting technique is used, the parent solution's values are shifted in the right direction by 1, and the value in the first position is set at a random value in the range [minVal maxVal] as follows.
$$S_{k} \left( 1 \right) = {\text{rand}}\left( {{\text{minVal}},{\text{maxVal}}} \right),$$(1)$$S_{k} \left( {2:D} \right) = P\left( {1:D - 1} \right),$$(2)where minVal and maxVal are the minima and maximum values for the variables in each solution.
-
ii.
If the − 1 frameshifting technique is used, the parent solution values are shifted backward by 1, and the value in the last position is to set a random value in the range [minVal, maxVal].
$$S_{k} \left( D \right) = {\text{rand}}\left( {{\text{minVal}},{\text{maxVal}}} \right),$$(3)$$S_{k} \left( {1:D - 1} \right) = P\left( {2:D} \right),$$(4)The symbol Sk refers to the kth generated protein, P is the parent solution, and D is the problem dimension (number of variables in each solution). The result of frameshifting represents a new protein sequence.
-
i.
-
c.
New virion formation a uniform crossover is applied to the generated sub-proteins to produce a new virion (new solution).
-
a.
-
3.
Mutation a mutation operator is applied to the solution created in the previous step to generate a new mutated solution as follows:
$$Z\left( i \right) = \left\{ {\begin{array}{*{20}l} r \hfill & {{\text{if}}\;{\text{rand}}\left( {0,1} \right) < {\text{MR}}} \hfill \\ {X\left( i \right)} \hfill & {{\text{otherwise}}} \hfill \\ \end{array} } \right.$$(5)X is the solution before mutation. Z is the mutated solution, \(X\left( i \right)\) and \(Z\left( i \right)\) are the ith element in the old and new solutions, respectively, i = 1, …, D, and r is a random value in the range [minVal, maxVal]. MR is the mutation rate.
-
4.
The objective function is evaluated for the new solution, and the population is updated for the next generation (the solutions with the highest fitness remain, and the others are removed).
-
5.
Repeat steps (2–4) for the new population until termination criteria are achieved. For example, the maximum number of iterations is reached.
-
6.
Output the best solution found so far.
The flowchart of the proposed algorithm is shown in Fig. 10.
2.3 Parameters of the proposed algorithm
The parameters of the proposed algorithm are suggested as follows:
-
Max_Iter maximum number of iterations.
-
PopNo number of solutions in the population.
-
MinVal and MaxVal minimum and maximum value of variables in a solution.
-
D problem dimension (number of variables in each solution).
-
CostFunction objective function;
-
MR Mutation Rate, MR is set at a value in the range [0.1 0.001].
The pseudocode of the proposed algorithm is as follows:

-
Shifting a number that represents the type of frameshifting used. For example, shiftingNo = 1 means that the + 1 frameshifting technique is used. We noticed that the + 1 frameshifting technique yields the best results.
-
numOfProtiens number of proteins generated during virus replication in the proposed algorithm, numOfProteins is 2.
3 Experimental results and discussion
3.1 Benchmark functions
To test the efficiency of the proposed COVIDOA, we utilized 30 benchmark functions. The first 20 are classical standard benchmark test functions (http://benchmarkfcns.xyz) [29]. We selected five functions from IEEE CEC 2019 Competition (https://www.mathworks.com/matlabcentral/fileexchange/72123-cec-06-2019-matlabimplementation) [33], while the remaining five were selected from CEC 2011 Competition on Testing Evolutionary Algorithms on Real-World Optimization Problems [16] as follows:
-
I.
Classical benchmark problems
Twenty standard optimization functions from the literature are discussed and used to test the proposed algorithm's efficiency. These functions are classified into four groups: unimodal, multimodal, fixed-dimension, and n-dimensional functions [35, 39]. In fixed-dimension problems, the number of design variables (problem dimension) is fixed, while the other n-dimension problems use any design variables. A multimodal function has multiple (at least locally optimum) solutions instead of a unimodal function with a single optimum solution [35]. As in “Table 11 in the Appendix”, the chosen optimization functions are described in terms of the function name, formula, problem dimension (D), range of possible values, the global optimum, and the group of benchmark functions to which it belongs.
-
II.
IEEE CEC 2019 benchmark problems
In addition to the classical benchmark functions, five CEC benchmark functions are utilized for evaluation. These are a group of modern test functions known as "The 100-Digit Challenge" intended to be used in single objective numerical optimization IEEE competitions [2]. As shown in “Table 12 in the Appendix”, these functions are described in terms of problem dimension, range of possible values, and the global optimum (https://www.mathworks.com/) [32].
-
III.
CEC 2011 Real-World Problems
For further evaluation, COVIDOA was applied to five real-world optimization problems. These are bound-constrained real-world optimization problems selected from the CEC 2011 Competition on Testing Evolutionary Algorithms on Real-World Optimization. These problems are as follows [16]:
-
1.
Lennard–Jones Potential Problem.
-
2.
Transmission Network Expansion Planning (TNEP) problem.
-
3.
Tersoff Potential Function Minimization Problem for model Si(B).
-
4.
Tersoff Potential Function Minimization Problem for model Si(C).
-
5.
Spread spectrum radar polyphase problem.
A detailed description of these real-world problems is discussed in the 2011 IEEE-Congress on Evolutionary Computation (IEEE-CEC 2011) [16].
3.2 Experimental results
COVIDOA is utilized to solve the previously mentioned test problems. COVIDOA is implemented in MATLAB R2016a software. The results are compared with eight well-known and recent optimization algorithms: GA [26], DE [63], PSO [44], FPA [75], GWO [51], WOA [50], SOA [19], and CHIO [4]. We selected this group of algorithms for many reasons:
-
(1)
Most of them are recent and published in reputable sources.
-
(2)
All of them have high performance in single-objective optimization on various benchmark functions.
-
(3)
Their MATLAB implementations are publicly available on the MATLAB website (https://www.mathworks.com/) [32].
-
(4)
Some of them are evolutionary algorithms, such as GA and DE, in the same category as COVIDOA. CHIO algorithm simulates coronavirus, as is COVIDOA, but each has its inspiration.
The obtained results change at each run in optimization algorithms due to the random process. The commonly used number of runs is 30, which would give acceptable statistical precision. So, the proposed algorithm and the state-of-the-art algorithms are run 30 times.
The proposed and state-of-the-art algorithms use Max_Iter = 500 and PopNo = 1000 for the classical benchmark functions. The comparison is made regarding optimum cost, average cost, standard deviation (STD), and convergence speed. The authors downloaded the source code of the state-of-the-art optimization algorithms from the MATLAB website.
Tables 1, 2, 3 and 4 show the results of the best cost, average cost, standard deviation, and convergence speed, respectively, for the 20 classical benchmark functions. The best-obtained results in all the following tables are highlighted in bold. Table 1 shows that the proposed algorithm reaches the optimum global cost in 18 of 20 problems and gets very close to the global optimum in the two remaining problems. Table 2 proves the COVIDOA algorithm's efficiency in terms of the average cost. It reaches the minimum average cost in 17 from 20 problems and the second minimum average cost in three. The third criterion is STD, which shows how the cost values are far from the average cost. Low STD values mean the cost values over the iterations are clustered closely around the average cost. Table 3 shows that the COVIDOA algorithm reaches the minimum STD values in 17 of 20 problems, the second minimum in two, and the third minimum in two, which means that the results of COVIDOA are more reliable than the other algorithms with higher STD values.
Compared with the recently proposed algorithm, CHIO, which simulates herd immunity's effect in tackling the COVID pandemic, COVIDOA is the best. As shown in Tables 1, 2, 3 and 4 and Figs. 11, 12, 13 and 14, CHIO reaches the minimum optimum cost in seven benchmark functions only from 25; in contrast, COVIDOA reaches the minimum optimum cost in 21 from 25 test functions. This indicates that COVIDOA has robust exploration capabilities in comparison with CHIO.
Compared with PSO, GWO, and WOA, COVIDOA is superior according to most of the test problems' best cost, average cost, and STD values. It has a higher convergence speed as it reaches the global minimum after the first few iterations, as in functions (F3, F8, F7, F15, and F16).
The curves in Figs. 11 and 12 represent the relationship between the iterations and the corresponding best cost for the classical test functions. The obtained results using the selected test problems are divided into two groups and displayed in Figs. 11 and 12. Figure 11 represents the test problems for which the COVIDOA algorithm outperforms the other algorithms. In contrast, Fig. 12 shows the results of test problems in which the COVIDOA algorithm has a performance very close to the others.
Additionally, to prove the results' statistical significance, the test results of the 20 classical benchmark functions are compared using Wilcoxon rank-sum test at the 5% significance level [18]. A null hypothesis is a type of hypothesis used in statistics that assumes no significant difference between the two methods' average values. A small p-value (typically ≤ 0.05) indicates strong evidence against the null hypothesis [70].
Table 5 introduces the p values computed by Wilcoxon rank-sum test that compares the COIDOA with eight well-known metaheuristic algorithms for the 20 classical benchmark functions. We observed from Table 5 that all p values are less than a 5% significance level for all comparative algorithms, strong evidence against the null hypothesis. Therefore, we conclude that the COVIDOA is better than all other comparative algorithms.
CEC benchmark functions, COVIDOA, and state-of-the-art algorithms search for the optimum cost for 250 iterations with 1000 solutions in each generation. The results of the best cost, average cost, and STD values are discussed in Table 6, and the convergence curves are shown in Fig. 13. COVIDOA is superior to the other algorithms in CEC01, CEC06, and CEC01. The CEC03 problem reaches the minimum best cost and the second minimum average cost ad STD value. In the case of CEC07, however, it is not the best; it achieves excellent results compared to GA, FPA, GWO, WOA, SOA, and CHIO algorithms.
All test results for the CEC benchmark functions were compared using the Wilcoxon rank-sum test to prove their statistical significance. Table 7 shows the p values computed by Wilcoxon rank-sum test that compares the COIDOA with other well-known algorithms for CEC benchmark functions. It is evident from Table 7 that all p values are less than 5% which proves the statistical significance of COVIDOA.
To test the impact of changing parameter values on the performance of OVIDOA, we used nine different scenarios by changing the values of the parameters MR (Mutation Rate) and numOfProtiens. We utilized the values of 0.1, 0.01, ad 0.001 for MR, 2, 4, and 6 for numOfProtiens which produces nine scenarios, as shown in Table 8. The results of each scenario on the selected five IEEE CEC benchmark problems are presented in Table 9. We noticed that scenario 1 (MR = 0.1 and numOfProtiens = 2) has better results, followed by scenario 4. The common between these two scenarios is MR = 0.1 which represents a higher mutation rate. This comparison shows that higher MR values are better for improving the performance of the proposed algorithm.
For testing COVIDOA on CEC real-world problems, we obtain our results over 500 iterations. The proposed and state-of-the-art algorithms were run 25 independent times as suggested by IEEE-CEC 2011 Competition [16]. Table 10 and Fig. 14 show the results of the selected CEC real-world problems. The proposed algorithm achieves the optimum best cost, average cost, and STD values for all five selected problems.
Although the general steps of COVIDOA and other evolutionary algorithms, such as GA and DE, are very similar, COVIDOA is superior to them, as shown in Tables 1, 2, 3, 4, 5, 6, 7, 8, 9 and 10. This progress is caused by the additional step proposed in the replication phase of COVIDOA, the frameshifting technique. Adding frameshifting technique in the replication process helps COVIDOA update solutions in each generation, helping to reach global optimum rapidly.
3.3 Explorations and exploitation capabilities of COVIDOA
It is essential to test the efficiency of the proposed algorithm. In other words, it is necessary to test its exploration and exploitation capabilities. In exploration, the algorithm searches for new solutions in new regions, while exploitation means using existing solutions and improving their fitness [21]. Mutation and crossover steps in COVIDOA are used to create new solutions, so they are methods to explore the problem space. On the other hand, selecting an existing parent virus and applying the frameshifting technique to produce new children represents the exploitation of the current solution features. Unimodal test functions F1, F7, F10, and F13 can evaluate the exploitation feature because they have only one global optimum solution. Multimodal functions F2, F3, F4, and F5 can help assess the exploration capability of COVIDOA because they have many optimum solutions.
3.4 Convergence of COVIDOA
In Table 4, the convergence speed of COVIDOA and the other algorithms for the classical benchmark functions are classified into three groups: Fast, Moderate, and Low, where algorithms that reach the minimum cost in the first 100 iterations are classified as fast convergence algorithms, those that get the minimum cost from iteration 100 to 300 are moderate convergence algorithms, and the others classified as slow convergence algorithms. As shown in Table 5 and Fig. 13, the proposed algorithm has fast convergence in the majority (16 from 20) of the test problems and moderate in the others. In contrast, other state-of-the-art algorithms may slow the test problems' convergence.
Overall results reveal that COVIDOA reaches the minimum best cost, average cost, and standard deviation in most test problems. It also has high exploration and exploitation capabilities and a high convergence speed during iterations.
4 Conclusion
A novel evolutionary optimization algorithm (COVIDOA) inspired by the replication lifecycle of SARS-CoV-2 is presented. The proposed COVIDOA was tested by solving 20 classical benchmark problems, five CEC benchmark test functions, and five CEC 2011 real-world problems. The proposed COVIDOA is compared with the state-of-the-art nature-inspired optimization algorithms in terms of best cost, average cost, standard deviation, and convergence speed. The proposed algorithm is implemented using MATLAB R2016a software, and the source code of the state-of-the-art algorithms and the benchmark problems are downloaded from the mathworks.com website. The experimental results proved that the proposed algorithm outperforms the state-of-the-art optimization algorithms in most test problems and has very close results to other algorithms in the rest of the test problems. COVIDOA has high exploitation and exploration capabilities and convergence speed compared to other metaheuristics.
Future work may include the implementation of COVIDOA in solving large-scale problems in different fields.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Ab Wahab MN, Nefti-Meziani S, Atyabi A (2015) A comprehensive review of swarm optimization algorithms. PLoS ONE 10(5):e0122827
Abdullah JM, Ahmed T (2019) Fitness-dependent optimizer: inspired by the bee swarming reproductive process. IEEE Access 7:43473–43486
Alatas B, Can U (2015) Physics based metaheuristic optimization algorithms for global optimization
Al-Betar MA, Alyasseri ZAA, Awadallah MA et al (2021) Coronavirus herd immunity optimizer (CHIO). Neural Comput Appl 33:5011–5042. https://doi.org/10.1007/s00521-020-05296-6
Andre J, Siarry P, Dognon T (2001) An improvement of the standard genetic algorithm fighting premature convergence in continuous optimization. Adv Eng Softw 32(1):49–60
Askarzadeh A (2014) Bird mating optimizer: an optimization algorithm inspired by bird mating strategies. Commun Nonlinear Sci Numer Simul 19(4):1213–1228
Baluja S (1994) Population-based incremental learning. a method for integrating genetic search based function optimization and competitive learning. Carnegie-Mellon Univ Pittsburgh Pa Dept Of Computer Science
Bar-On YM et al (2020) SARS-CoV-2 (COVID-19) by the numbers. Elife 9:e57309. https://doi.org/10.7554/eLife.57309
Bergmann CC, Silverman RH (2020) COVID-19: Coronavirus replication, pathogenesis, and therapeutic strategies. Cleve Clin J Med 87(6):321–327. https://doi.org/10.3949/ccjm.87a.20047
Birbil Şİ, Fang SC (2003) An electromagnetism-like mechanism for global optimization. J Global Optim 25(3):263–282
Brameier MF, Banzhaf W (2007) Linear genetic programming. Springer
Brian DA, Baric RS (2005) Coronavirus genome structure and replication. Curr Top Microbiol Immunol 287:1–30. https://doi.org/10.1007/3-540-26765-4_1
Brooks SP, Morgan BJ (1995) Optimization using simulated annealing. J R Stat Soc Ser D (Stat) 44(2):241–257
Cobb M (2015) Who discovered messenger RNA? Curr Biol 25(13):526–532
Darwish A (2018) Bio-inspired computing: algorithms review, deep analysis, and the scope of applications. Fut Comput Inform J 3(2):231–246
Das S, Suganthan PN (2010) Problem definitions and evaluation criteria for cec 2011 competition on testing evolutionary algorithms on real world optimization problems. Jadavpur University, Nanyang Technological University, Kolkata, pp 341–359
Das S, Biswas A, Dasgupta S, Abraham A (2009) Bacterial foraging optimization algorithm: theoretical foundations, analysis, and applications. Found Comput Intell 3:23–55
Derrac J, García S, Molina D, Herrera F (2011) A practical tutorial on the use of nonparametric statistical tests as a methodology for comparing evolutionary and swarm intelligence algorithms. Swarm Evol Comput 1(1):3–18
Dhiman G, Kumar V (2019) Seagull optimization algorithm: theory and its applications for large-scale industrial engineering problems. Knowl-Based Syst 165:169–196
Domingo E, Escarmís C, Sevilla N, Moya A, Elena SF, Quer J, Holland JJ (1996) Basic concepts in RNA virus evolution. FASEB J 10(8):859–864
Epitropakis M, Plagianakos V, Vrahatis M (2008) Balancing the exploration and exploitation capabilities of the Differential Evolution Algorithm, pp 2686–2693
Eskandar H, Sadollah A, Bahreininejad A, Hamdi M (2014) Water cycle algorithm–a novel metaheuristic optimization method for solving constrained engineering optimization problems. Comput Struct 110:151–166
Fogel DB (1997) The advantages of evolutionary computation. BCEC, pp 1–11
Formato RA (2007) Central force optimization: a new metaheuristic with applications in applied electromagnetics. Prog Electromagn Res 77:425–491
Gandomi AH, Alavi AH (2015) Krill herd: a new bio-inspired optimization algorithm. Commun Nonlinear Sci Numer Simul 17(12):4831–4845
Holland JH (1992) Genetic algorithms. Sci Am 267:66–72
Hosseini E, Ghafoor KZ, Sadiq AS, Guizani M, Emrouznejad A (2020) Covid-19 optimizer algorithm, modeling, and controlling of coronavirus distribution process. IEEE J Biomed Health Inform 24(10):2765–2775
Hsiao YT, Chuang CL, Jiang JA, Chien CC (2005) A novel optimization algorithm: space gravitational optimization. In: 2005 IEEE international conference on systems, man and cybernetics, vol 3, pp 2323–2328
https://commons.wikimedia.org/wiki/File:3D_medical_animation_corona_virus.jpg
https://www.mathworks.com/matlabcentral/fileexchange/72123-cec-06-2019-matlabimplementation
https://www.microscope.com/coronavirus-under-an-electron-microscope/
Hussain K, Salleh MNM, Cheng S, Shi Y (2019) Metaheuristic research: a comprehensive survey. Artif Intell Rev 52(4):2191–2233
Ivanov IP, Atkins JF (2015) Ribosomal frameshifting in decoding antizyme mRNAs from yeast and protists to humans: close to 300 cases reveal remarkable diversity despite underlying conservation. Nucleic Acids Res 35(6):1842–1858
Jacks T, Madhani HD, Masiarz FR, Varmus HE (1988) Signals for ribosomal frameshifting in the Rous sarcoma virus gag-pol region. Cell 55(3):447–458
James JQ, Li VO (2015) A social spider algorithm for global optimization. Appl Soft Comput 30:614–627
Jamil M, Yang X-S (2013) A literature survey of benchmark functions for global optimization problems. Int J Math Model Numer Optim 4(2):150–194
Kang S, Peng W, Zhu Y, Lu S, Zhou M, Lin W, Deng M (2019) Recent Progress in understanding 2019 novel coronavirus associated with human respiratory disease: detection, mechanism and treatment. Int J Antimicrob Agents 55(5):2020
Karaboga D (2005) An idea based on honey bee swarm for numerical optimization. S2CID 8215393, vol 200, pp 1–10
Kaveh A, Talatahari S (2010) A novel heuristic optimization method: charged system search. Acta Mech 213(3):267–289
Kelly JA, Olson AN, Neupane K, Munshi S, San Emeterio J, Pollack L, Dinman JD (2020) Structural and functional conservation of the programmed− 1 ribosomal frameshift signal of SARS coronavirus 2 (SARS-CoV-2). J Biol Chem 295(31):10741–10748
Kennedy J, Eberhart R (1995) Particle swarm optimization. In: Proceedings of ICNN'95-international conference on neural networks, vol 4, pp 1942–1948
Khan MI, Khan ZA, Baig MH, Ahmad I, Farouk AE, Song YG, Dong JJ (2020) Comparative genome analysis of novel coronavirus (SARS-CoV-2) from different geographical locations and the effect of mutations on major target proteins, An in-silico insight. PLoS ONE 15(9):e0238344
Lipowski A, Lipowska D (2012) Roulette-wheel selection via stochastic acceptance. Physica A 391(6):2193–2196
Martínez-Álvarez F, Asencio-Cortés G, Torres JF, Gutiérrez-Avilés D, Melgar-García L, Pérez-Chacón R, Rubio-Escudero C, Santos JC, Lora AT (2020) Coronavirus optimization algorithm: a bioinspired metaheuristic based on the COVID-19 propagation model. Big Data
Meng X, Liu Y, Gao X, Zhang H (2014) A new bio-inspired algorithm: chicken swarm optimization. In: International conference in swarm intelligence, pp 86–94
Milewska A, Kula-Pacurar A, Wadas J, Suder A, Szczepanski A, Dabrowska A, Rajfur Z (2020) Replication of SARS-CoV-2 in human respiratory epithelium. J Virol 94(15)
Mirjalili S, Lewis A (2016) The whale optimization algorithm. Adv Eng Softw 95:51–67
Mirjalili S, Mirjalili SM, Lewis A (2014) Grey wolf optimizer. Adv Eng Softw 69:46–61
Moghal A, Mohler K, Ibba M (2014) Mistranslation of the genetic code. FEBS Lett 588(23)
Moscato P, Cotta C, Mendes A (2004) Memetic algorithms. New Optim Techn Eng, pp 53–85
Napthine S, Ling R, Finch LK, Jones JD, Bell S, Brierley I, Firth AE (2017) Protein-directed ribosomal frameshifting temporally regulates gene expression. Nat Commun. https://doi.org/10.1038/ncomms15582
Nordin P, Keller RE, Francone FD (1998) Genetic programming: an introduction: on the automatic evolution of computer programs and its applications. Morgan Kaufmann Publishers Inc., Burlington
O’Neill M, Ryan C (2001) Grammatical evolution. IEEE Trans Evol Comput 5(4):349–358
Pál KF (2006) Hysteretic optimization, faster and simpler. Physica A 360(2):525–533
Pascual MR (2020) Coronavirus SARS-CoV-2: Analysis of subgenomic mRNA transcription, 3CLpro and PL2pro protease cleavage sites and protein synthesis. Preprint https://doi.org/10.48550/arxiv.2004.00746
Rapley R, Whitehouse D (Eds) (2015) Molecular biology and biotechnology. Royal Society of Chemistry.
Rashedi E, Nezamabadi-Pour H, Saryazdi S (2009) GSA: a gravitational search algorithm. Inf Sci 179(13):2232–2248
Rashedi E, Nezamabadi-Pour H, Saryazdi S (2010) BGSA: binary gravitational search algorithm. Natural Comput 9(3):727–745
Rechenberg I (1989) Evolution strategy: Nature's way of optimization. In: Optimization: methods and applications, possibilities and limitations, pp 106–126
Rocca P, Oliveri G, Massa A (2011) Differential evolution as applied to electromagnetics. IEEE Antennas Propag Mag 53(1):38–49
Sacco WF, Oliveira CREA (2005) A new stochastic optimization algorithm based on a particle collision metaheuristic. In: Proceedings of 6th WCSMO
Schoeman D, Fielding BC (2019) Coronavirus envelope protein. Curr Knowl 16(1):1–22
Shah-Hosseini H (2009) The intelligent water drops algorithm: a nature-inspired swarm-based optimization algorithm. Int J Bio-inspired Comput 1(1–2):71–79
Shang J, Ye G, Shi K, Wan Y, Luo C, Aihara H, Li F (2020) Structural basis of receptor recognition by SARS-CoV-2. Nature 581(7807):221–224
Sharma J et al (2020) Pharmacological approaches for targeting cystic fibrosis nonsense mutations. Eur J Med Chem 200:112436. https://doi.org/10.1016/j.ejmech.2020.112436
Simon D (2008) Biogeography-based optimization. IEEE Trans Evol Comput 12(6):702–713
Szucs D, Ioannidis J (2017) When null hypothesis significance testing is unsuitable for research: a reassessment. Front Hum Neurosci 11:390
Terpos E et al (2020) Hematological findings and complications of COVID-19. Am J Hematol 95(7):834–847. https://doi.org/10.1002/ajh.25829
Tu YF, Chien CS, Yarmishyn AA, Lin YY, Luo YH, Lin YT, Wang ML (2020) A review of SARS-CoV-2 and the ongoing clinical trials. Int J Mol Sci 21(7)
Wang S, Zhang Y, Liu S, Peng H, Mackey V, Sun L (2020) Coronaviruses and the associated potential therapeutics for the viral infections. J Infect Dis Ther 8(417)
Yamauchi Y, Greber UF (2016) “Principles of virus uncoating,” cues and the snooker ball. Traffic 17(6):569–592
Yang XS (2012) Flower pollination algorithm for global optimization. In: International conference on unconventional computing and natural computation, pp 240–249
Yang XS, Deb S (2009) Cuckoo search via Lévy flights. In: 2009 World Congress on Nature & biologically inspired computing (NaBIC), pp 210–214
Yu X, Gen M (2010) Introduction to evolutionary algorithms. Springer
Zheng YJ (2015) Water wave optimization: a new nature-inspired metaheuristic. Comput Oper Res 55:1–11
Ziebuhr J (2020) The coronavirus replicase: insights into a sophisticated enzyme machinery. The Nidoviruses. Springer, Boston, pp 3–11
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants performed by any authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Khalid, A.M., Hosny, K.M. & Mirjalili, S. COVIDOA: a novel evolutionary optimization algorithm based on coronavirus disease replication lifecycle. Neural Comput & Applic 34, 22465–22492 (2022). https://doi.org/10.1007/s00521-022-07639-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00521-022-07639-x