Abstract
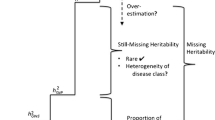
About 10 years ago, after the first large-scale genome-wide association studies (GWAS) were conducted to find genes associated with common complex diseases, investigators were surprised to find that the amount of heritability explained by the significant hits was very low for almost all the studied traits. Indeed, when compared to heritability estimates expected from the observed trait concordance within families, the heritability explained by the associated variants was always much smaller, more than ten times smaller for some traits. There was thus a problem of “missing heritability” and different hypotheses were proposed to help find this “missing heritability”. These hypotheses involved novel research strategies in which different groups engaged including among others increasing sample sizes of GWAS or looking for rare variants and structural variations that were not captured by the SNP-chips used in GWAS. How successful have these efforts been in finding the “missing heritability”? Could it be that the problem of “missing heritability” was ill-defined? These are the questions that will be addressed in this paper by taking some different examples of complex traits.
Similar content being viewed by others
References
Altmuller J, Palmer LJ, Fischer G, Scherb H, Wjst M (2001) Genomewide scans of complex human diseases: true linkage is hard to find. Am J Hum Genet 69:936–950. https://doi.org/10.1086/324069
Aschard H (2016) A perspective on interaction effects in genetic association studies. Genet Epidemiol 40:678–688. https://doi.org/10.1002/gepi.21989
Aschard H, Chen J, Cornelis MC, Chibnik LB, Karlson EW, Kraft P (2012) Inclusion of gene-gene and gene-environment interactions unlikely to dramatically improve risk prediction for complex diseases. Am J Hum Genet 90:962–972. https://doi.org/10.1016/j.ajhg.2012.04.017
Auer PL, Lettre G (2015) Rare variant association studies: considerations, challenges and opportunities. Genome Med 7:16. https://doi.org/10.1186/s13073-015-0138-2
Bellenguez C, Charbonnier C, Grenier-Boley B, Quenez O, Le Guennec K, Nicolas G, Chauhan G, Wallon D, Rousseau S, Richard AC, Boland A, Bourque G, Munter HM, Olaso R, Meyer V, Rollin-Sillaire A, Pasquier F, Letenneur L, Redon R, Dartigues JF, Tzourio C, Frebourg T, Lathrop M, Deleuze JF, Hannequin D, Genin E, Amouyel P, Debette S, Lambert JC, Campion D, Collaborators CM (2017) Contribution to Alzheimer’s disease risk of rare variants in TREM2, SORL1, and ABCA7 in 1779 cases and 1273 controls. Neurobiol Aging 59:220e1–220e9. https://doi.org/10.1016/j.neurobiolaging.2017.07.001
Benchek PH, Morris NJ (2013) How meaningful are heritability estimates of liability? Hum Genet 132:1351–1360. https://doi.org/10.1007/s00439-013-1334-z
Bourrat P, Lu Q, Jablonka E (2017) Why the missing heritability might not be in the DNA. Bioessays. https://doi.org/10.1002/bies.201700067
Boyle EA, Li YI, Pritchard JK (2017) An expanded view of complex traits: from polygenic to omnigenic. Cell 169:1177–1186. https://doi.org/10.1016/j.cell.2017.05.038
Browning SR, Browning BL (2011) Population structure can inflate SNP-based heritability estimates. Am J Hum Genet 89:191–193. https://doi.org/10.1016/j.ajhg.2011.05.025(author reply 193-5)
Bulik-Sullivan BK, Loh PR, Finucane HK, Ripke S, Yang J, Schizophrenia Working Group of the Psychiatric Genomics C, Patterson N, Daly MJ, Price AL, Neale BM (2015) LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet 47:291–295. https://doi.org/10.1038/ng.3211
Buniello A, MacArthur JAL, Cerezo M, Harris LW, Hayhurst J, Malangone C, McMahon A, Morales J, Mountjoy E, Sollis E, Suveges D, Vrousgou O, Whetzel PL, Amode R, Guillen JA, Riat HS, Trevanion SJ, Hall P, Junkins H, Flicek P, Burdett T, Hindorff LA, Cunningham F, Parkinson H (2019) The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res 47:D1005–D1012. https://doi.org/10.1093/nar/gky1120
Carlborg O, Haley CS (2004) Epistasis: too often neglected in complex trait studies? Nat Rev Genet 5:618–625. https://doi.org/10.1038/nrg1407
Colella S, Yau C, Taylor JM, Mirza G, Butler H, Clouston P, Bassett AS, Seller A, Holmes CC, Ragoussis J (2007) QuantiSNP: an Objective Bayes Hidden-Markov Model to detect and accurately map copy number variation using SNP genotyping data. Nucleic Acids Res 35:2013–2025. https://doi.org/10.1093/nar/gkm076
Cordell HJ (2002) Epistasis: what it means, what it doesn’t mean, and statistical methods to detect it in humans. Hum Mol Genet 11:2463–2468
Cuccaro D, De Marco EV, Cittadella R, Cavallaro S (2017) Copy number variants in Alzheimer’s disease. J Alzheimers Dis 55:37–52. https://doi.org/10.3233/JAD-160469
Dandine-Roulland C, Bellenguez C, Debette S, Amouyel P, Genin E, Perdry H (2016) Accuracy of heritability estimations in presence of hidden population stratification. Sci Rep 6:26471. https://doi.org/10.1038/srep26471
de Los Campos G, Sorensen D, Gianola D (2015) Genomic heritability: what is it? PLoS Genet 11:e1005048. https://doi.org/10.1371/journal.pgen.1005048
Dempfle A, Scherag A, Hein R, Beckmann L, Chang-Claude J, Schafer H (2008) Gene-environment interactions for complex traits: definitions, methodological requirements and challenges. Eur J Hum Genet 16:1164–1172. https://doi.org/10.1038/ejhg.2008.106
Dewan A, Liu M, Hartman S, Zhang SS, Liu DT, Zhao C, Tam PO, Chan WM, Lam DS, Snyder M, Barnstable C, Pang CP, Hoh J (2006) HTRA1 promoter polymorphism in wet age-related macular degeneration. Science 314:989–992. https://doi.org/10.1126/science.1133807
Evans LM, Tahmasbi R, Vrieze SI, Abecasis GR, Das S, Gazal S, Bjelland DW, de Candia TR, Haplotype Reference C, Goddard ME, Neale BM, Yang J, Visscher PM, Keller MC (2018) Comparison of methods that use whole genome data to estimate the heritability and genetic architecture of complex traits. Nat Genet 50:737–745. https://doi.org/10.1038/s41588-018-0108-x
Falconer DS (1965) The inheritance of liability to certain diseases, estimated from the incidence among relatives. Ann Hum Genet 29:51–76
Feldman MW, Ramachandran S (2018) Missing compared to what? Revisiting heritability, genes and culture. Philos Trans R Soc Lond B Biol Sci. https://doi.org/10.1098/rstb.2017.0064
Fisher RA (1918) The correlation between relatives on the supposition of Mendelian inheritance. Trans R Soc Edinb Earth Sci 52:399–433
Fuchsberger C, Flannick J, Teslovich TM, Mahajan A, Agarwala V, Gaulton KJ, Ma C, Fontanillas P, Moutsianas L, McCarthy DJ, Rivas MA, Perry JRB, Sim X, Blackwell TW, Robertson NR, Rayner NW, Cingolani P, Locke AE, Tajes JF, Highland HM, Dupuis J, Chines PS, Lindgren CM, Hartl C, Jackson AU, Chen H, Huyghe JR, van de Bunt M, Pearson RD, Kumar A, Muller-Nurasyid M, Grarup N, Stringham HM, Gamazon ER, Lee J, Chen Y, Scott RA, Below JE, Chen P, Huang J, Go MJ, Stitzel ML, Pasko D, Parker SCJ, Varga TV, Green T, Beer NL, Day-Williams AG, Ferreira T, Fingerlin T, Horikoshi M, Hu C, Huh I, Ikram MK, Kim BJ, Kim Y, Kim YJ, Kwon MS, Lee J, Lee S, Lin KH, Maxwell TJ, Nagai Y, Wang X, Welch RP, Yoon J, Zhang W, Barzilai N, Voight BF, Han BG, Jenkinson CP, Kuulasmaa T, Kuusisto J, Manning A, Ng MCY, Palmer ND, Balkau B, Stancakova A, Abboud HE, Boeing H, Giedraitis V, Prabhakaran D, Gottesman O, Scott J, Carey J, Kwan P, Grant G, Smith JD, Neale BM, Purcell S, Butterworth AS, Howson JMM, Lee HM, Lu Y, Kwak SH, Zhao W, Danesh J, Lam VKL, Park KS, Saleheen D et al (2016) The genetic architecture of type 2 diabetes. Nature 536:41–47. https://doi.org/10.1038/nature18642
Gamazon ER, Cox NJ, Davis LK (2014) Structural architecture of SNP effects on complex traits. Am J Hum Genet 95:477–489. https://doi.org/10.1016/j.ajhg.2014.09.009
Gatz M, Reynolds CA, Fratiglioni L, Johansson B, Mortimer JA, Berg S, Fiske A, Pedersen NL (2006) Role of genes and environments for explaining Alzheimer disease. Arch Gen Psychiatry 63:168–174. https://doi.org/10.1001/archpsyc.63.2.168
Gazal S, Loh PR, Finucane HK, Ganna A, Schoech A, Sunyaev S, Price AL (2018) Functional architecture of low-frequency variants highlights strength of negative selection across coding and non-coding annotations. Nat Genet 50:1600–1607. https://doi.org/10.1038/s41588-018-0231-8
Genin E, Clerget-Darpoux F (2015a) The missing heritability paradigm: a dramatic resurgence of the GIGO syndrome in genetics. Hum Hered 79:1–4. https://doi.org/10.1159/000370327
Genin E, Clerget-Darpoux F (2015b) Revisiting the polygenic additive liability model through the example of diabetes mellitus. Hum Hered 80:171–177. https://doi.org/10.1159/000447683
Golan D, Lander ES, Rosset S (2014) Measuring missing heritability: inferring the contribution of common variants. Proc Natl Acad Sci USA 111:E5272–E5281. https://doi.org/10.1073/pnas.1419064111
Gudbjartsson DF, Walters GB, Thorleifsson G, Stefansson H, Halldorsson BV, Zusmanovich P, Sulem P, Thorlacius S, Gylfason A, Steinberg S, Helgadottir A, Ingason A, Steinthorsdottir V, Olafsdottir EJ, Olafsdottir GH, Jonsson T, Borch-Johnsen K, Hansen T, Andersen G, Jorgensen T, Pedersen O, Aben KK, Witjes JA, Swinkels DW, den Heijer M, Franke B, Verbeek AL, Becker DM, Yanek LR, Becker LC, Tryggvadottir L, Rafnar T, Gulcher J, Kiemeney LA, Kong A, Thorsteinsdottir U, Stefansson K (2008) Many sequence variants affecting diversity of adult human height. Nat Genet 40:609–615. https://doi.org/10.1038/ng.122
Guo SW, Lange K (2000) Genetic mapping of complex traits: promises, problems, and prospects. Theor Popul Biol 57:1–11. https://doi.org/10.1006/tpbi.2000.1449
Haley CS, Visscher PM (1998) Strategies to utilize marker-quantitative trait loci associations. J Dairy Sci 81(Suppl 2):85–97
Hindorff LA, Sethupathy P, Junkins HA, Ramos EM, Mehta JP, Collins FS, Manolio TA (2009) Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci USA 106:9362–9367. https://doi.org/10.1073/pnas.0903103106
International HapMap C (2005) A haplotype map of the human genome. Nature 437:1299–1320. https://doi.org/10.1038/nature04226
International HapMap C (2002) https://www.genome.gov/10005336/2002-release-genetic-variation-mapping-launch/. Accessed 10 Nov 2018
Jun G, Manning A, Almeida M, Zawistowski M, Wood AR, Teslovich TM, Fuchsberger C, Feng S, Cingolani P, Gaulton KJ, Dyer T, Blackwell TW, Chen H, Chines PS, Choi S, Churchhouse C, Fontanillas P, King R, Lee S, Lincoln SE, Trubetskoy V, DePristo M, Fingerlin T, Grossman R, Grundstad J, Heath A, Kim J, Kim YJ, Laramie J, Lee J, Li H, Liu X, Livne O, Locke AE, Maller J, Mazur A, Morris AP, Pollin TI, Ragona D, Reich D, Rivas MA, Scott LJ, Sim X, Tearle RG, Teo YY, Williams AL, Zollner S, Curran JE, Peralta J, Akolkar B, Bell GI, Burtt NP, Cox NJ, Florez JC, Hanis CL, McKeon C, Mohlke KL, Seielstad M, Wilson JG, Atzmon G, Below JE, Dupuis J, Nicolae DL, Lehman D, Park T, Won S, Sladek R, Altshuler D, McCarthy MI, Duggirala R, Boehnke M, Frayling TM, Abecasis GR, Blangero J (2018) Evaluating the contribution of rare variants to type 2 diabetes and related traits using pedigrees. Proc Natl Acad Sci USA 115:379–384. https://doi.org/10.1073/pnas.1705859115
Kanoungi G, Nothnagel M (2018) Pathway-induced allelic spectra of diseases in the presence of strong genetic effects. Hum Genet 137:215–230. https://doi.org/10.1007/s00439-018-1872-5
Koch L (2014) Epigenetics: an epigenetic twist on the missing heritability of complex traits. Nat Rev Genet 15:218. https://doi.org/10.1038/nrg3698
Kryukov GV, Pennacchio LA, Sunyaev SR (2007) Most rare missense alleles are deleterious in humans: implications for complex disease and association studies. Am J Hum Genet 80:727–739. https://doi.org/10.1086/513473
Lambert JC, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, DeStafano AL, Bis JC, Beecham GW, Grenier-Boley B, Russo G, Thorton-Wells TA, Jones N, Smith AV, Chouraki V, Thomas C, Ikram MA, Zelenika D, Vardarajan BN, Kamatani Y, Lin CF, Gerrish A, Schmidt H, Kunkle B, Dunstan ML, Ruiz A, Bihoreau MT, Choi SH, Reitz C, Pasquier F, Cruchaga C, Craig D, Amin N, Berr C, Lopez OL, De Jager PL, Deramecourt V, Johnston JA, Evans D, Lovestone S, Letenneur L, Moron FJ, Rubinsztein DC, Eiriksdottir G, Sleegers K, Goate AM, Fievet N, Huentelman MW, Gill M, Brown K, Kamboh MI, Keller L, Barberger-Gateau P, McGuiness B, Larson EB, Green R, Myers AJ, Dufouil C, Todd S, Wallon D, Love S, Rogaeva E, Gallacher J, St George-Hyslop P, Clarimon J, Lleo A, Bayer A, Tsuang DW, Yu L, Tsolaki M, Bossu P, Spalletta G, Proitsi P, Collinge J, Sorbi S, Sanchez-Garcia F, Fox NC, Hardy J, Deniz Naranjo MC, Bosco P, Clarke R, Brayne C, Galimberti D, Mancuso M, Matthews F, European Alzheimer’s Disease I, Genetic, Environmental Risk in Alzheimer’s D, Alzheimer’s Disease Genetic C, Cohorts for H, Aging Research in Genomic E, Moebus S, Mecocci P, Del Zompo M, Maier W, Hampel H, Pilotto A, Bullido M, Panza F, Caffarra P et al (2013) Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet 45:1452–1458. https://doi.org/10.1038/ng.2802
Lee JJ, Chow CC (2014) Conditions for the validity of SNP-based heritability estimation. Hum Genet 133:1011–1022. https://doi.org/10.1007/s00439-014-1441-5
Lee SH, Wray NR, Goddard ME, Visscher PM (2011) Estimating missing heritability for disease from genome-wide association studies. Am J Hum Genet 88:294–305. https://doi.org/10.1016/j.ajhg.2011.02.002
Lettre G, Jackson AU, Gieger C, Schumacher FR, Berndt SI, Sanna S, Eyheramendy S, Voight BF, Butler JL, Guiducci C, Illig T, Hackett R, Heid IM, Jacobs KB, Lyssenko V, Uda M, Diabetes Genetics I, Fusion Kora, Prostate LC, Ovarian Cancer Screening T, Nurses’ Health S, SardiNia Boehnke M, Chanock SJ, Groop LC, Hu FB, Isomaa B, Kraft P, Peltonen L, Salomaa V, Schlessinger D, Hunter DJ, Hayes RB, Abecasis GR, Wichmann HE, Mohlke KL, Hirschhorn JN (2008) Identification of ten loci associated with height highlights new biological pathways in human growth. Nat Genet 40:584–591. https://doi.org/10.1038/ng.125
Lewontin RC (1974) Annotation: the analysis of variance and the analysis of causes. Am J Hum Genet 26:400–411
Lush JL (1937) Animal Breeding Plans. Ames, IA
Lynch M, Walsh B (1998) Genetic and the analysis of quantitative traits. Sinauer, Sunderland
Mace A, Tuke MA, Deelen P, Kristiansson K, Mattsson H, Noukas M, Sapkota Y, Schick U, Porcu E, Rueger S, McDaid AF, Porteous D, Winkler TW, Salvi E, Shrine N, Liu X, Ang WQ, Zhang W, Feitosa MF, Venturini C, van der Most PJ, Rosengren A, Wood AR, Beaumont RN, Jones SE, Ruth KS, Yaghootkar H, Tyrrell J, Havulinna AS, Boers H, Magi R, Kriebel J, Muller-Nurasyid M, Perola M, Nieminen M, Lokki ML, Kahonen M, Viikari JS, Geller F, Lahti J, Palotie A, Koponen P, Lundqvist A, Rissanen H, Bottinger EP, Afaq S, Wojczynski MK, Lenzini P, Nolte IM, Sparso T, Schupf N, Christensen K, Perls TT, Newman AB, Werge T, Snieder H, Spector TD, Chambers JC, Koskinen S, Melbye M, Raitakari OT, Lehtimaki T, Tobin MD, Wain LV, Sinisalo J, Peters A, Meitinger T, Martin NG, Wray NR, Montgomery GW, Medland SE, Swertz MA, Vartiainen E, Borodulin K, Mannisto S, Murray A, Bochud M, Jacquemont S, Rivadeneira F, Hansen TF, Oldehinkel AJ, Mangino M, Province MA, Deloukas P, Kooner JS, Freathy RM, Pennell C, Feenstra B, Strachan DP, Lettre G, Hirschhorn J, Cusi D, Heid IM, Hayward C, Mannik K, Beckmann JS, Loos RJF, Nyholt DR, Metspalu A, Eriksson JG et al (2017) CNV-association meta-analysis in 191,161 European adults reveals new loci associated with anthropometric traits. Nat Commun 8:744. https://doi.org/10.1038/s41467-017-00556-x
Mackay TF, Moore JH (2014) Why epistasis is important for tackling complex human disease genetics. Genome Med 6:124. https://doi.org/10.1186/gm561
Maher B (2008) Personal genomes: the case of the missing heritability. Nature 456:18–21. https://doi.org/10.1038/456018a
Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ, McCarthy MI, Ramos EM, Cardon LR, Chakravarti A, Cho JH, Guttmacher AE, Kong A, Kruglyak L, Mardis E, Rotimi CN, Slatkin M, Valle D, Whittemore AS, Boehnke M, Clark AG, Eichler EE, Gibson G, Haines JL, Mackay TF, McCarroll SA, Visscher PM (2009) Finding the missing heritability of complex diseases. Nature 461:747–753. https://doi.org/10.1038/nature08494
Marenne G, Chanock SJ, Malats N, Genin E (2013) Advantage of using allele-specific copy numbers when testing for association in regions with common copy number variants. PLoS One 8:e75350. https://doi.org/10.1371/journal.pone.0075350
Marouli E, Graff M, Medina-Gomez C, Lo KS, Wood AR, Kjaer TR, Fine RS, Lu Y, Schurmann C, Highland HM, Rueger S, Thorleifsson G, Justice AE, Lamparter D, Stirrups KE, Turcot V, Young KL, Winkler TW, Esko T, Karaderi T, Locke AE, Masca NG, Ng MC, Mudgal P, Rivas MA, Vedantam S, Mahajan A, Guo X, Abecasis G, Aben KK, Adair LS, Alam DS, Albrecht E, Allin KH, Allison M, Amouyel P, Appel EV, Arveiler D, Asselbergs FW, Auer PL, Balkau B, Banas B, Bang LE, Benn M, Bergmann S, Bielak LF, Bluher M, Boeing H, Boerwinkle E, Boger CA, Bonnycastle LL, Bork-Jensen J, Bots ML, Bottinger EP, Bowden DW, Brandslund I, Breen G, Brilliant MH, Broer L, Burt AA, Butterworth AS, Carey DJ, Caulfield MJ, Chambers JC, Chasman DI, Chen YI, Chowdhury R, Christensen C, Chu AY, Cocca M, Collins FS, Cook JP, Corley J, Galbany JC, Cox AJ, Cuellar-Partida G, Danesh J, Davies G, de Bakker PI, de Borst GJ, de Denus S, de Groot MC, de Mutsert R, Deary IJ, Dedoussis G, Demerath EW, den Hollander AI, Dennis JG, Di Angelantonio E, Drenos F, Du M, Dunning AM, Easton DF, Ebeling T, Edwards TL, Ellinor PT, Elliott P, Evangelou E, Farmaki AE, Faul JD et al (2017) Rare and low-frequency coding variants alter human adult height. Nature 542:186–190. https://doi.org/10.1038/nature21039
McCarroll SA (2008) Extending genome-wide association studies to copy-number variation. Hum Mol Genet 17:R135–R142. https://doi.org/10.1093/hmg/ddn282
Moore DS, Shenk D (2017) The heritability fallacy. Wiley Interdiscip Rev Cogn Sci. https://doi.org/10.1002/wcs.1400
Nelson RM, Pettersson ME, Carlborg O (2013) A century after Fisher: time for a new paradigm in quantitative genetics. Trends Genet 29:669–676. https://doi.org/10.1016/j.tig.2013.09.006
Nicolas G, Charbonnier C, Campion D (2016) From common to rare variants: the genetic component of Alzheimer disease. Hum Hered 81:129–141. https://doi.org/10.1159/000452256
Ozaki K, Ohnishi Y, Iida A, Sekine A, Yamada R, Tsunoda T, Sato H, Sato H, Hori M, Nakamura Y, Tanaka T (2002) Functional SNPs in the lymphotoxin-alpha gene that are associated with susceptibility to myocardial infarction. Nat Genet 32:650–654. https://doi.org/10.1038/ng1047
Reich DE, Lander ES (2001) On the allelic spectrum of human disease. Trends Genet 17:502–510
Ridge PG, Mukherjee S, Crane PK, Kauwe JS, Alzheimer’s Disease Genetics C (2013) Alzheimer’s disease: analyzing the missing heritability. PLoS One 8:e79771. https://doi.org/10.1371/journal.pone.0079771
Risch N, Merikangas K (1996) The future of genetic studies of complex human diseases. Science 273:1516–1517
Ritland K (1996) A marker-based method for inferences about quantitative inheritance in natural populations. Evolution 50:1062–1073. https://doi.org/10.1111/j.1558-5646.1996.tb02347.x
Rovelet-Lecrux A, Hannequin D, Raux G, Le Meur N, Laquerriere A, Vital A, Dumanchin C, Feuillette S, Brice A, Vercelletto M, Dubas F, Frebourg T, Campion D (2006) APP locus duplication causes autosomal dominant early-onset Alzheimer disease with cerebral amyloid angiopathy. Nat Genet 38:24–26. https://doi.org/10.1038/ng1718
Rovelet-Lecrux A, Legallic S, Wallon D, Flaman JM, Martinaud O, Bombois S, Rollin-Sillaire A, Michon A, Le Ber I, Pariente J, Puel M, Paquet C, Croisile B, Thomas-Anterion C, Vercelletto M, Levy R, Frebourg T, Hannequin D, Campion D, Investigators of the Gp (2012) A genome-wide study reveals rare CNVs exclusive to extreme phenotypes of Alzheimer disease. Eur J Hum Genet 20:613–617. https://doi.org/10.1038/ejhg.2011.225
Saint Pierre A, Genin E (2014) How important are rare variants in common disease? Brief Funct Genom 13:353–361. https://doi.org/10.1093/bfgp/elu025
Sandoval-Motta S, Aldana M, Martinez-Romero E, Frank A (2017) the human microbiome and the missing heritability problem. Front Genet 8:80. https://doi.org/10.3389/fgene.2017.00080
Schork NJ, Murray SS, Frazer KA, Topol EJ (2009) Common vs. rare allele hypotheses for complex diseases. Curr Opin Genet Dev 19:212–219. https://doi.org/10.1016/j.gde.2009.04.010
Shen L, Jia J (2016) An overview of genome-wide association studies in Alzheimer’s disease. Neurosci Bull 32:183–190. https://doi.org/10.1007/s12264-016-0011-3
Speed D, Hemani G, Johnson MR, Balding DJ (2012) Improved heritability estimation from genome-wide SNPs. Am J Hum Genet 91:1011–1021. https://doi.org/10.1016/j.ajhg.2012.10.010
Speed D, Cai N, Consortium U, Johnson MR, Nejentsev S, Balding DJ (2017) Reevaluation of SNP heritability in complex human traits. Nat Genet 49:986–992. https://doi.org/10.1038/ng.3865
Tenesa A, Haley CS (2013) The heritability of human disease: estimation, uses and abuses. Nat Rev Genet 14:139–149. https://doi.org/10.1038/nrg3377
Thomson G (2001) Significance levels in genome scans. Adv Genet 42:475–486
Trerotola M, Relli V, Simeone P, Alberti S (2015) Epigenetic inheritance and the missing heritability. Hum Genom 9:17. https://doi.org/10.1186/s40246-015-0041-3
Visscher PM, Wray NR (2015) Concepts and misconceptions about the polygenic additive model applied to disease. Hum Hered 80:165–170. https://doi.org/10.1159/000446931
Visscher PM, Hill WG, Wray NR (2008) Heritability in the genomics era–concepts and misconceptions. Nat Rev Genet 9:255–266. https://doi.org/10.1038/nrg2322
Wang K, Li M, Hadley D, Liu R, Glessner J, Grant SF, Hakonarson H, Bucan M (2007) PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res 17:1665–1674. https://doi.org/10.1101/gr.6861907
Weedon MN, Lango H, Lindgren CM, Wallace C, Evans DM, Mangino M, Freathy RM, Perry JR, Stevens S, Hall AS, Samani NJ, Shields B, Prokopenko I, Farrall M, Dominiczak A, Diabetes Genetics I, Wellcome Trust Case Control C, Johnson T, Bergmann S, Beckmann JS, Vollenweider P, Waterworth DM, Mooser V, Palmer CN, Morris AD, Ouwehand WH, Cambridge GEMC, Zhao JH, Li S, Loos RJ, Barroso I, Deloukas P, Sandhu MS, Wheeler E, Soranzo N, Inouye M, Wareham NJ, Caulfield M, Munroe PB, Hattersley AT, McCarthy MI, Frayling TM (2008) Genome-wide association analysis identifies 20 loci that influence adult height. Nat Genet 40:575–583. https://doi.org/10.1038/ng.121
Wellcome Trust Case Control C, Craddock N, Hurles ME, Cardin N, Pearson RD, Plagnol V, Robson S, Vukcevic D, Barnes C, Conrad DF, Giannoulatou E, Holmes C, Marchini JL, Stirrups K, Tobin MD, Wain LV, Yau C, Aerts J, Ahmad T, Andrews TD, Arbury H, Attwood A, Auton A, Ball SG, Balmforth AJ, Barrett JC, Barroso I, Barton A, Bennett AJ, Bhaskar S, Blaszczyk K, Bowes J, Brand OJ, Braund PS, Bredin F, Breen G, Brown MJ, Bruce IN, Bull J, Burren OS, Burton J, Byrnes J, Caesar S, Clee CM, Coffey AJ, Connell JM, Cooper JD, Dominiczak AF, Downes K, Drummond HE, Dudakia D, Dunham A, Ebbs B, Eccles D, Edkins S, Edwards C, Elliot A, Emery P, Evans DM, Evans G, Eyre S, Farmer A, Ferrier IN, Feuk L, Fitzgerald T, Flynn E, Forbes A, Forty L, Franklyn JA, Freathy RM, Gibbs P, Gilbert P, Gokumen O, Gordon-Smith K, Gray E, Green E, Groves CJ, Grozeva D, Gwilliam R, Hall A, Hammond N, Hardy M, Harrison P, Hassanali N, Hebaishi H, Hines S, Hinks A, Hitman GA, Hocking L, Howard E, Howard P, Howson JM, Hughes D, Hunt S, Isaacs JD, Jain M, Jewell DP, Johnson T, Jolley JD, Jones IR et al (2010) Genome-wide association study of CNVs in 16,000 cases of eight common diseases and 3,000 shared controls. Nature 464:713–720. https://doi.org/10.1038/nature08979
Wray NR, Wijmenga C, Sullivan PF, Yang J, Visscher PM (2018) Common disease is more complex than implied by the core gene omnigenic model. Cell 173:1573–1580. https://doi.org/10.1016/j.cell.2018.05.051
Wright S (1920) The relative importance of heredity and environment in determining the piebald pattern of guinea-pigs. Proc Natl Acad Sci USA 6:320–332
Xue A, Wu Y, Zhu Z, Zhang F, Kemper KE, Zheng Z, Yengo L, Lloyd-Jones LR, Sidorenko J, Wu Y, e QC, McRae AF, Visscher PM, Zeng J, Yang J (2018) Genome-wide association analyses identify 143 risk variants and putative regulatory mechanisms for type 2 diabetes. Nat Commun 9:2941. https://doi.org/10.1038/s41467-018-04951-w
Yang J, Benyamin B, McEvoy BP, Gordon S, Henders AK, Nyholt DR, Madden PA, Heath AC, Martin NG, Montgomery GW, Goddard ME, Visscher PM (2010) Common SNPs explain a large proportion of the heritability for human height. Nat Genet 42:565–569. https://doi.org/10.1038/ng.608
Yang J, Lee SH, Goddard ME, Visscher PM (2011) GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet 88:76–82. https://doi.org/10.1016/j.ajhg.2010.11.011
Yengo L, Sidorenko J, Kemper KE, Zheng Z, Wood AR, Weedon MN, Frayling TM, Hirschhorn J, Yang J, Visscher PM (2018) Meta-analysis of genome-wide association studies for height and body mass index in ~ 700,000 individuals of European ancestry. bioRxiv. https://doi.org/10.1101/274654
Young AI, Frigge ML, Gudbjartsson DF, Thorleifsson G, Bjornsdottir G, Sulem P, Masson G, Thorsteinsdottir U, Stefansson K, Kong A (2018) Relatedness disequilibrium regression estimates heritability without environmental bias. Nat Genet 50:1304–1310. https://doi.org/10.1038/s41588-018-0178-9
Zeng J, de Vlaming R, Wu Y, Robinson MR, Lloyd-Jones LR, Yengo L, Yap CX, Xue A, Sidorenko J, McRae AF, Powell JE, Montgomery GW, Metspalu A, Esko T, Gibson G, Wray NR, Visscher PM, Yang J (2018) Signatures of negative selection in the genetic architecture of human complex traits. Nat Genet 50:746–753. https://doi.org/10.1038/s41588-018-0101-4
Zhu Z, Bakshi A, Vinkhuyzen AA, Hemani G, Lee SH, Nolte IM, van Vliet-Ostaptchouk JV, Snieder H, LifeLines Cohort S, Esko T, Milani L, Magi R, Metspalu A, Hill WG, Weir BS, Goddard ME, Visscher PM, Yang J (2015) Dominance genetic variation contributes little to the missing heritability for human complex traits. Am J Hum Genet 96:377–385. https://doi.org/10.1016/j.ajhg.2015.01.001
Zuk O, Hechter E, Sunyaev SR, Lander ES (2012) The mystery of missing heritability: genetic interactions create phantom heritability. Proc Natl Acad Sci USA 109:1193–1198. https://doi.org/10.1073/pnas.1119675109
Zuk O, Schaffner SF, Samocha K, Do R, Hechter E, Kathiresan S, Daly MJ, Neale BM, Sunyaev SR, Lander ES (2014) Searching for missing heritability: designing rare variant association studies. Proc Natl Acad Sci USA 111:E455–E464. https://doi.org/10.1073/pnas.1322563111
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Génin, E. Missing heritability of complex diseases: case solved?. Hum Genet 139, 103–113 (2020). https://doi.org/10.1007/s00439-019-02034-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-019-02034-4