Abstract
Purpose
Osteosarcoma (OS) is one of the most prevalent primary malignant bone tumors in adolescent. HOTAIR is highly expressed and associated with the epigenetic modifications, especially DNA methylation, in cancer. However, the regulation mechanism between HOTAIR and DNA methylation and the biological effects of them in the pathogenesis of osteosarcoma remains elusive.
Method
Through RNA-sequencing and computational analysis, followed by a variety of experimental validations, we report a novel interplay between HOTAIR, miR-126, and DNA methylation in OS.
Results
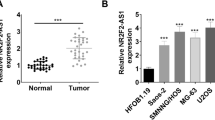
We found that HOTAIR is highly expressed in OS cells and the knockdown of HOTAIR leads to the down-regulation of DNMT1, as well as the decrease of global DNA methylation level. RNA-sequencing analysis of HOTAIR-regulated gene shows that CDKN2A is significantly repressed by HOTAIR. A series of experiments show that HOTAIR represses the expression of CDKN2A through inhibiting the promoter activity of CDKN2A by DNA hypermethylation. Further evidence shows that HOTAIR activates the expression of DNMT1 through repressing miR-126, which is the negative regulator of DNMT1. Functionally, HOTAIR depletion increases the sensibility of OS cells to DNMT1 inhibitor through regulating the viability and apoptosis of OS cells via HOTAIR-miR126-DNMT1-CDKN2A axis.
Conclusion
These results not only enrich our understanding of the regulation relationship between non-coding RNA, DNA methylation, and gene expression, however, also provide a novel direction in developing more sophisticated therapeutic strategies for OS patients.







Similar content being viewed by others
Abbreviations
- OS:
-
Osteosarcoma
- MDS:
-
Myelodysplastic syndrome
- DAC:
-
Decitabine
- RT:
-
Reverse transcription
- ChIP:
-
Chromatin immunoprecipitation
- qPCR:
-
Quantitative PCR
- MedIP:
-
Methylated DNA immunoprecipitation
References
Akhavan-Niaki H, Samadani AA (2013) DNA methylation and cancer development: molecular mechanism. Cell Biochem Biophys 67:501–513. doi:10.1007/s12013-013-9555-2
Akiyama T, Dass CR, Choong PF (2008) Novel therapeutic strategy for osteosarcoma targeting osteoclast differentiation, bone-resorbing activity, and apoptosis pathway. Mol Cancer Ther 7:3461–3469. doi:10.1158/1535-7163.MCT-08-0530
Alhejaily A, Day AG, Feilotter HE, Baetz T, Lebrun DP (2014) Inactivation of the CDKN2A tumor-suppressor gene by deletion or methylation is common at diagnosis in follicular lymphoma and associated with poor clinical outcome. Clin Cancer Res 20:1676–1686. doi:10.1158/1078-0432.CCR-13-2175
Battistelli C et al (2017) The Snail repressor recruits EZH2 to specific genomic sites through the enrollment of the lncRNA HOTAIR in epithelial-to-mesenchymal transition. Oncogene 36:942–955. doi:10.1038/onc.2016.260
Braconi C et al (2011) microRNA-29 can regulate expression of the long non-coding RNA gene MEG3 in hepatocellular cancer. Oncogene 30:4750–4756. doi:10.1038/onc.2011.193
Broadhead ML, Clark JC, Myers DE, Dass CR, Choong PF (2011) The molecular pathogenesis of osteosarcoma: a review. Sarcoma 2011:959248. doi:10.1155/2011/959248
Cai B, Song XQ, Cai JP, Zhang S (2014) HOTAIR: a cancer-related long non-coding RNA. Neoplasma 61:379–391
Calin GA et al (2007) Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell 12:215–229. doi:10.1016/j.ccr.2007.07.027
Chu H et al (2017) The HOTAIR, PRNCR1 and POLR2E polymorphisms are associated with cancer risk: a meta-analysis. Oncotarget. doi:10.18632/oncotarget.14920
Deng J, Yang M, Jiang R, An N, Wang X, Liu B (2017) Long non-coding RNA HOTAIR regulates the proliferation, self-renewal capacity, tumor formation and migration of the cancer stem-like cell (CSC) subpopulation enriched from breast cancer cells. PLoS One 12:e0170860. doi:10.1371/journal.pone.0170860
Fang S et al (2016) Long noncoding RNA-HOTAIR affects chemoresistance by regulating HOXA1 methylation in small cell lung cancer cells. Lab Invest 96:60–68. doi:10.1038/labinvest.2015.123
Feng J, Meyer CA, Wang Q, Liu JS, Shirley Liu X, Zhang Y (2012) GFOLD: a generalized fold change for ranking differentially expressed genes from RNA-seq data. Bioinformatics 28:2782–2788. doi:10.1093/bioinformatics/bts515
Gokul G, Khosla S (2013) DNA methylation and cancer. Subcell Biochem 61:597–625. doi:10.1007/978-94-007-4525-4_26
Gupta RA et al (2010) Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464:1071–1076. doi:10.1038/nature08975
Herman JG, Baylin SB (2003) Gene silencing in cancer in association with promoter hypermethylation. N Engl J Med 349:2042–2054. doi:10.1056/NEJMra023075
Jung Y et al (2007) Potential advantages of DNA methyltransferase 1 (DNMT1)-targeted inhibition for cancer therapy. J Mol Med (Berl) 85:1137–1148. doi:10.1007/s00109-007-0216-z
Kallen AN et al (2013) The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol Cell 52:101–112. doi:10.1016/j.molcel.2013.08.027
Kim K et al (2013) HOTAIR is a negative prognostic factor and exhibits pro-oncogenic activity in pancreatic cancer. Oncogene 32:1616–1625. doi:10.1038/onc.2012.193
Kogo R et al (2011) Long noncoding RNA HOTAIR regulates polycomb-dependent chromatin modification and is associated with poor prognosis in colorectal cancers. Cancer Res 71:6320–6326. doi:10.1158/0008-5472.CAN-11-1021
Kohonen-Corish MR et al (2014) KRAS mutations and CDKN2A promoter methylation show an interactive adverse effect on survival and predict recurrence of rectal cancer. Int J Cancer 134:2820–2828. doi:10.1002/ijc.28619
Lamoureux F et al (2007) Therapeutic relevance of osteoprotegerin gene therapy in osteosarcoma: blockade of the vicious cycle between tumor cell proliferation and bone resorption. Cancer Res 67:7308–7318. doi:10.1158/0008-5472.CAN-06-4130
Lau WM, Ho TH, Hui KM (2007) p16INK4A-silencing augments DNA damage-induced apoptosis in cervical cancer cells. Oncogene 26:6050–6060. doi:10.1038/sj.onc.1210405
Lee SJ et al (2005) DNMT3B polymorphisms and risk of primary lung cancer. Carcinogenesis 26:403–409. doi:10.1093/carcin/bgh307
Ley TJ et al (2010) DNMT3A mutations in acute myeloid leukemia. N Engl J Med 363:2424–2433. doi:10.1056/NEJMoa1005143
Li Y et al (2014) Enhancement of radiosensitivity by 5-Aza-CdR through activation of G2/M checkpoint response and apoptosis in osteosarcoma cells. Tumour Biol 35:4831–4839. doi:10.1007/s13277-014-1634-5
Li F, Cao L, Hang D, Wang F, Wang Q (2015) Long non-coding RNA HOTTIP is up-regulated and associated with poor prognosis in patients with osteosarcoma Int J. Clin Exp Pathol 8:11414–11420
Liang WC et al (2015) The lncRNA H19 promotes epithelial to mesenchymal transition by functioning as miRNA sponges in colorectal cancer. Oncotarget 6:22513–22525. doi:10.18632/oncotarget.4154
Lin J et al (2011) Recurrent DNMT3A R882 mutations in Chinese patients with acute myeloid leukemia and myelodysplastic syndrome. PLoS One 6:e26906. doi:10.1371/journal.pone.0026906
Liu R et al (2015) DNMT1-microRNA126 epigenetic circuit contributes to esophageal squamous cell carcinoma growth via ADAM9-EGFR-AKT signaling. Clin Cancer Res 21:854–863. doi:10.1158/1078-0432.CCR-14-1740
Margueron R, Reinberg D (2011) The Polycomb complex PRC2 and its mark in life. Nature 469:343–349. doi:10.1038/nature09784
Otto T, Sicinski P (2017) Cell cycle proteins as promising targets in cancer therapy. Nat Rev Cancer 17:93–115. doi:10.1038/nrc.2016.138
Pasini D et al (2010) JARID2 regulates binding of the Polycomb repressive complex 2 to target genes in ES cells. Nature 464:306–310. doi:10.1038/nature08788
Savage SA et al (2007) Analysis of genes critical for growth regulation identifies Insulin-like Growth Factor 2 Receptor variations with possible functional significance as risk factors for osteosarcoma. Cancer Epidemiol Biomarkers Prev 16:1667–1674. doi:10.1158/1055-9965.EPI-07-0214
Shen H, Wang L, Spitz MR, Hong WK, Mao L, Wei Q (2002) A novel polymorphism in human cytosine DNA-methyltransferase-3B promoter is associated with an increased risk of lung cancer. Cancer Res 62:4992–4995
Suzuki M, Sunaga N, Shames DS, Toyooka S, Gazdar AF, Minna JD (2004) RNA interference-mediated knockdown of DNA methyltransferase 1 leads to promoter demethylation and gene re-expression in human lung and breast cancer cells. Cancer Res 64:3137–3143
Teschendorff AE et al (2015) HOTAIR and its surrogate DNA methylation signature indicate carboplatin resistance in ovarian cancer. Genome Med 7:108. doi:10.1186/s13073-015-0233-4
Ting AH, Jair KW, Suzuki H, Yen RW, Baylin SB, Schuebel KE (2004) CpG island hypermethylation is maintained in human colorectal cancer cells after RNAi-mediated depletion of DNMT1. Nat Genet 36:582–584. doi:10.1038/ng1365
Turcan S et al (2013) Efficient induction of differentiation and growth inhibition in IDH1 mutant glioma cells by the DNMT Inhibitor. Decitabine Oncotarget 4:1729–1736. doi:10.18632/oncotarget.1412
Villa R et al (2007) Role of the polycomb repressive complex 2 in acute promyelocytic leukemia. Cancer Cell 11:513–525. doi:10.1016/j.ccr.2007.04.009
Volanakis EJ, Boothby MR, Sherr CJ (2013) Epigenetic regulation of the Ink4a-Arf (Cdkn2a) tumor suppressor locus in the initiation and progression of Notch1-driven T cell acute lymphoblastic leukemia. Exp Hematol 41:377–386. doi:10.1016/j.exphem.2012.11.006
Wu Y et al (2015) Long non-coding RNA HOTAIR promotes tumor cell invasion and metastasis by recruiting EZH2 and repressing E-cadherin in oral squamous cell carcinoma. Int J Oncol 46:2586–2594. doi:10.3892/ijo.2015.2976
Xia M, Yao L, Zhang Q, Wang F, Mei H, Guo X, Huang W (2017) Long noncoding RNA HOTAIR promotes metastasis of renal cell carcinoma by up-regulating histone H3K27 demethylase JMJD3. Oncotarget 8:19795–19802. doi:10.18632/oncotarget.15047
Xu ZY et al (2013) Knockdown of long non-coding RNA HOTAIR suppresses tumor invasion and reverses epithelial-mesenchymal transition in gastric cancer. Int J Biol Sci 9:587–597. doi:10.7150/ijbs.6339
Yan J, Dang Y, Liu S, Zhang Y, Zhang G (2016) LncRNA HOTAIR promotes cisplatin resistance in gastric cancer by targeting miR-126 to activate the PI3K/AKT/MRP1 genes. Tumour Biol. doi:10.1007/s13277-016-5448-5
Yang Z, Zhou L, Wu LM, Lai MC, Xie HY, Zhang F, Zheng SS (2011) Overexpression of long non-coding RNA HOTAIR predicts tumor recurrence in hepatocellular carcinoma patients following liver transplantation. Ann Surg Oncol 18:1243–1250. doi:10.1245/s10434-011-1581-y
Yao C et al (2013) Perifosine induces cell apoptosis in human osteosarcoma cells: new implication for osteosarcoma therapy? Cell Biochem Biophys 65:217–227. doi:10.1007/s12013-012-9423-5
Zhang J, Xiao X, Liu J (2015) The role of circulating miRNAs in multiple myeloma. Sci China Life Sci 58:1262–1269. doi:10.1007/s11427-015-4969-2
Zhang YY, Huang SH, Zhou HR, Chen CJ, Tian LH, Shen JZ (2016) Role of HOTAIR in the diagnosis and prognosis of acute leukemia. Oncol Rep 36:3113–3122. doi:10.3892/or.2016.5147
Zhao S et al (2011) MicroRNA-126 regulates DNA methylation in CD4+ T cells and contributes to systemic lupus erythematosus by targeting DNA methyltransferase 1. Arthritis Rheum 63:1376–1386. doi:10.1002/art.30196
Author information
Authors and Affiliations
Contributions
FS and XL designed the study; XL, HL, GF, MH, YS, and KX performed experiments; YL and HL performed data analysis; XL and FS wrote the manuscript; FS, XL, and HL revised the manuscript; and all authors approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Xingang Li and Hongming Lu are co-first authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, X., Lu, H., Fan, G. et al. A novel interplay between HOTAIR and DNA methylation in osteosarcoma cells indicates a new therapeutic strategy. J Cancer Res Clin Oncol 143, 2189–2200 (2017). https://doi.org/10.1007/s00432-017-2478-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-017-2478-3