Abstract
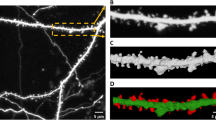
Dendritic spines are postsynaptic structures the morphology of which correlates with the strength of synaptic efficacy. Measurements of spine density and spine morphology are achievable using recent imaging and bioinformatics tools. The three-dimensional automated analysis requires optimization of image acquisition and treatment. Here, we studied the critical steps for optimal confocal microscopy imaging of dendritic spines. We characterize the deconvolution process and show that it improves spine morphology analysis. With this method, images of dendritic spines from medium spiny neurons are automatically detected by the software Neuronstudio, which retrieves spine density as well as spine diameter and volume. This approach is illustrated with three-dimensional analysis of dendritic spines in a mouse model of Huntington’s disease: the transgenic R6/2 mice. In symptomatic mutant mice, we confirm the decrease in spine density, and the method brings further information and show a decrease in spine volume and dendrite diameter. Moreover, we show a significant decrease in spine density at presymptomatic age which so far has gone unnoticed.





Similar content being viewed by others
References
Arellano JI, Benavides-Piccione R, Defelipe J, Yuste R (2007) Ultrastructure of dendritic spines: correlation between synaptic and spine morphologies. Front Neurosci 1(1):131–143
Boutet de Monvel J, Le Calvez S, Ulfendahl M (2001) Image restoration for confocal microscopy: improving the limits of deconvolution, with application to the visualization of the mammalian hearing organ. Biophys J 80(5):2455–2470
Cheng J, Zhou X, Miller E, Witt RM, Zhu J, Sabatini BL, Wong ST (2007) A novel computational approach for automatic dendrite spines detection in two-photon laser scan microscopy. J Neurosci Methods 165(1):122–134
Conchello JA, Lichtman JW (2005) Optical sectioning microscopy. Nat Methods 2(12):920–931
Crook ZR, Housman D (2011) Huntington’s disease: can mice lead the way to treatment? Neuron 69(3):423–435
Diaspro A, Federici F, Robello M (2002) Influence of refractive-index mismatch in high-resolution three-dimensional confocal microscopy. Appl Opt 41(4):685–690
Donohue DE, Ascoli GA (2011) Automated reconstruction of neuronal morphology: an overview. Brain Res Rev 67(1–2):94–102
Dumitriu D, Hao J, Hara Y, Kaufmann J, Janssen WG, Lou W, Rapp PR, Morrison JH (2010) Selective changes in thin spine density and morphology in monkey prefrontal cortex correlate with aging-related cognitive impairment. J Neurosci 30(22):7507–7515
Dusch E, Dorval T, Vincent N, Wachsmuth M, Genovesio A (2007) Three-dimensional point spread function model for line-scanning confocal microscope with high-aperture objective. J Microsc 228(Pt 2):132–138
Fiala JC, Spacek J, Harris KM (2002) Dendritic spine pathology: cause or consequence of neurological disorders? Brain Res Brain Res Rev 39(1):29–54
Gan WB, Grutzendler J, Wong WT, Wong RO, Lichtman JW (2000) Multicolor “DiOlistic” labeling of the nervous system using lipophilic dye combinations. Neuron 27(2):219–225
García-López P, García-Marín V, Freire M (2007) The discovery of dendritic spines by Cajal in 1888 and its relevance in the present neuroscience. Prog Neurobiol 83(2):110–130
Gibson Lanni (1991) Experimental test of an analytic model of aberration in an oil-immersion objective lens used in 3D light microscopy. J Opt Soc Am A 9(1):154–166
Gillette TA, Brown KM, Svoboda K, Liu Y, Ascoli GA (2011) DIADEMchallenge.org: A compendium of resources fostering the continuous development of automated neuronal reconstruction. Neuroinformatics 9(2–3):303–304
Grossman AW, Aldridge GM, Lee KJ, Zeman MK, Jun CS, Azam HS, Arii T, Imoto K, Greenough WT, Rhyu IJ (2010) Developmental characteristics of dendritic spines in the dentate gyrus of Fmr1 knockout mice. Brain Res 1355:221–227
Grutzendler J, Tsai J, Gan WB (2003) Rapid labeling of neuronal populations by ballistic delivery of fluorescent dyes. Methods 30(1):79–85
Harris KM, Stevens JK (1989) Dendritic spines of CA 1 pyramidal cells in the rat hippocampus: serial electron microscopy with reference to their biophysical characteristics. J Neurosci 9(8):2982–2997
Hell S, Reiner G, Cremer C, Stelzer HK (1993) Aberrations in confocal fluorescence microscopy induced by mismatches in refractive index. J Microsc 169(3):391–405
Holtmaat A, Svoboda K (2009) Experience-dependent structural synaptic plasticity in the mammalian brain. Nat Rev Neurosci 10(9):647–658
Hotulainen P, Hoogenraad CC (2010) Actin in dendritic spines: connecting dynamics to function. J Cell Biol 189(4):619–629
Inoue S (2006) Foundations of confocal scanned imaging in light microscopy. In: Pawley JB (ed) Handbook of biological confocal microscopy, 3rd edn. Springer, New York, pp 1–16
Kim BG, Dai HN, McAtee M, Vicini S, Bregman BS (2007) Labeling of dendritic spines with the carbocyanine dye DiI for confocal microscopic imaging in lightly fixed cortical slices. J Neurosci Methods 162(1–2):237–243
Klapstein GJ, Fisher RS, Zanjani H, Cepeda C, Jokel ES, Chesselet MF, Levine MS (2001) Electrophysiological and morphological changes in striatal spiny neurons in R6/2 Huntington’s disease transgenic mice. J Neurophysiol 86(6):2667–2677
Koh IY, Lindquist WB, Zito K, Nimchinsky EA, Svoboda K (2002) An image analysis algorithm for dendritic spines. Neural Comput 14(6):1283–1310
Lai X, Lin Z, Ward ES, Ober RJ (2005) Noise suppression of point spread functions and its influence on deconvolution of three-dimensional fluorescence microscopy image sets. J Microsc 217(Pt 1):93–108
Li JY, Popovic N, Brundin P (2005) The use of the R6 transgenic mouse models of Huntington’s disease in attempts to develop novel therapeutic strategies. NeuroRx 2(3):447–464
Luebke JI, Weaver CM, Rocher AB, Rodriguez A, Crimins JL, Dickstein DL, Wearne SL, Hof PR (2010) Dendritic vulnerability in neurodegenerative disease: insights from analyses of cortical pyramidal neurons in transgenic mouse models. Br Struct Funct 214:181–199
Mangiarini L, Sathasivam K, Seller M, Cozens B, Harper A, Hetherington C, Lawton M, Trottier Y, Lehrach H, Davies SW, Bates GP (1996) Exon 1 of the HD gene with an expanded CAG repeat is sufficient to cause a progressive neurological phenotype in transgenic mice. Cell 87(3):493–506
McNally JG, Karpova T, Cooper J, Conchello JA (1999) Three-dimensional imaging by deconvolution microscopy. Methods 19(3):373–385
Meijering E (2010) Neuron tracing in perspectives. Cytometry 77A:693–704
North AJ (2006) Seeing is believing? A beginner’s guide to practical pitfalls in image acquisition. J Cell Biol 172(1):9–18
Pawley J (2000) The 39 steps: a cautionary tale of quantitative 3-D fluorescence microscopy. Biotechniques 28(5):884–886
Pawley J (2006) Points, pixels, and gray levels: digitizing image data. In: Pawley JB (ed) Handbook of biological confocal microscopy, 3rd edn. Springer, New York, pp 59–79
Peng H (2008) Bioimage informatics: a new area of engineering biology. Bioinformatics 24(17):1827–1836
Penzes P, Cahill ME, Jones KA, Van Leeuwen JE, Woolfrey KM (2011) Dendritic spine pathology in neuropsychiatric disorders. Nat Neurosci 14(3):285–293
Peters A, Kaiserman-Abramof IR (1970) The small pyramidal neuron of the rat cerebral cortex The perikaryon, dendrites and spines. Am J Anat 127(4):321–355
Preza C, Miller MI, Thomas LJ, McNally JG (1992) Regularized linear method for reconstruction of three-dimensional microscopic objects from optical sections. J Opt Soc Am A 9(2):219–228
Rodriguez A, Ehlenberger DB, Hof PR, Wearne SL (2006) Rayburst sampling, an algorithm for automated three-dimensional shape analysis from laser scanning microscopy images. Nat Protoc 1(4):2152–2161
Rodriguez A, Ehlenberger DB, Dickstein DL, Hof PR, Wearne SL (2008) Automated three-dimensional detection and shape classification of dendritic spines from fluorescence microscopy images. Plos One 3(4):e1997
Ronneberger O, Baddeley D, Scheipl F, Verveer PJ, Burkhardt H, Cremer C, Fahrmeir L, Cremer T, Joffe B (2008) Spatial quantitative analysis of fluorescently labeled nuclear structures: problems methods, pitfalls. Chromosome Res 16(3):523–562
Rostaing P, Real E, Siksou L, Lechaire JP, Boudier T, Boeckers TM, Gertler F, Gundelfinger ED, Triller A, Marty S (2006) Analysis of synaptic ultrastructure without fixative using high-pressure freezing and tomography. Eur J Neurosci 24(12):3463–3474
Roze E, Bonnet C, Betuing S, Caboche J (2010) Huntington’s disease. Adv Exp Med Biol 685:45–63
Russo SJ, Dietz DM, Dumitriu D, Morrison JH, Malenka RC, Nestler EJ (2010) The addicted synapse: mechanisms of synaptic and structural plasticity in nucleus accumbens. Trends Neurosci 33(6):267–276
Sarder P, Nehorai A (2006) Deconvolution methods for 3-D fluorescence microscopy images. Signal Process. Mag IEEE 23:32–45
Schrader M, Hell SW, van der Voort HTM (1996) Potential of confocal microscopes to resolve in the 50–100 nm range. Appl Phys Lett 69(24):3644–3646
Scorcioni R, Polavaram S, Ascoli GA (2008) L-Measure: a web-accessible tool for the analysis, comparison and search of digital reconstructions of neuronal morphologies. Nat Protoc 3(5):866–876
Seabold GK, Daunais JB, Rau A, Grant KA, Alvarez VA (2010) DiOLISTIC labeling of neurons from rodent and non-human primate brain slices. J Vis Exp (41). pii: 2081
Shaw P (1994) Deconvolution in 3-D optical microscopy. Histochem J 26(9):687–694
Shaw P, Rawlins D (1991) The point-spread function of a confocal microscope: its measurement and use in deconvolution of 3-D data. J Microsc 163(2):151–165
Shen H, Sesack SR, Toda S, Kalivas PW (2008) Automated quantification of dendritic spine density and spine head diameter in medium spiny neurons of the nucleus accumbens. Brain Struct Funct 213(1–2):149–157
Shen HW, Toda S, Moussawi K, Bouknight A, Zahm DS, Kalivas PW (2009) Altered dendritic spine plasticity in cocaine-withdrawn rats. J Neurosci 29(9):2876–2884
Sheppard CJR, Gan X, Gu M, Roy M (2006) Signal to noise ratio in confocal microscopy. In: Pawley JB (ed) Handbook of biological confocal microscopy, 3rd edn. Springer, New York, pp 442–451
Sibarita JB (2005) Deconvolution microscopy. Adv Biochem Eng Biotechnol 95:201–243
Swedlow JR, Eliceiri KW (2009) Open source bioimage informatics for cell biology. Trends Cell Biol 19(11):656–660
Tao-Cheng JH, Gallant PE, Brightman MW, Dosemeci A, Reese TS (2007) Structural changes at synapses after delayed perfusion fixation in different regions of the mouse brain. J Comp Neurol 501(5):731–740
van der Voort H, Strasters K (1995) Restoration of confocal images for quantitative image analysis. J Microsc 178(2):165–181
Vecellio M, Schwaller B, Meyer M, Hunziker W, Celio MR (2000) Alterations in Purkinje cell spines of calbindin D-28 k and parvalbumin knock-out mice. Eur J Neurosci 12(3):945–954
Wallace W, Bear MF (2004) A morphological correlate of synaptic scaling in visual cortex. J Neurosci 24(31):6928–6938
Wallace W, Schaefer LH (1080) Swedlow JR (2001) A working person’s guide to deconvolution in light microscopy. Biotechniques 31(5):1076–1078 1082 passim
Wearne SL, Rodriguez A, Ehlenberger DB, Rocher AB, Henderson SC, Hof PR (2005) New techniques for imaging, digitization and analysis of three-dimensional neural morphology on multiple scales. Neuroscience 136:661–680
Wimmer VC, Möller A (2010) High-resolution confocal imaging in tissue. Methods Mol Biol 611:183–191
Wolf DE (2007) The optics of microscope image formation. Methods Cell Biol 81:11–42
Wu CC, Reilly JF, Young WG, Morrison JH, Bloom FE (2004) High-throughput morphometric analysis of individual neurons. Cereb Cortex 14(5):543–554
Xu X, Wong ST (2006) Optical microscopic image processing of dendritic spines morphology. Signal Process Mag IEEE 23(4):132–135
Yoo H, Song I, Gweon DG (2006) Measurement and restoration of the point spread function of fluorescence confocal microscopy. J Microsc 221(Pt 3):172–176
Yuan X, Trachtenberg JT, Potter SM, Roysam B (2009) MDL constrained 3-D grayscale skeletonization algorithm for automated extraction of dendrites and spines from fluorescence confocal images. Neuroinformatics 7(4):213–232
Yuste R, Bonhoeffer T (2001) Morphological changes in dendritic spines associated with long-term synaptic plasticity. Annu Rev Neurosci 24:1071–1089
Zhang Y, Zhou X, Witt RM, Sabatini BL, Adjeroh D, Wong ST (2007) Dendritic spine detection using curvilinear structure detector and LDA classifier. Neuroimage 36(2):346–360
Zhang Y, Chen K, Baron M, Teylan MA, Kim Y, Song Z, Greengard P, Wong ST (2010) A neurocomputational method for fully automated 3D dendritic spine detection and segmentation of medium-sized spiny neurons. Neuroimage 50(4):1472–1484
Acknowledgments
This work was supported by Centre National pour la Recherche Scientifique (CNRS), University Pierre and Marie Curie (UPMC), the hereditary disease foundation (HDF) and l’ “Agence Nationale pour la Recherche” (ANR-08-BLAN). We wish to thank Dr. Virginie Georget, Dr; Susanne Bolte and Richard Schwartzmann from the Cellular Imaging facility of the IFR 83 (Institut Fédératif de Recherche). We thank Dr. Anina Moritz, Dr. Sophie Scotto and Emma Cahill for critical reading and helpful comments on the manuscript. All authors report no biomedical financial interests or potential conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Heck, N., Betuing, S., Vanhoutte, P. et al. A deconvolution method to improve automated 3D-analysis of dendritic spines: application to a mouse model of Huntington’s disease. Brain Struct Funct 217, 421–434 (2012). https://doi.org/10.1007/s00429-011-0340-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00429-011-0340-y