Abstract
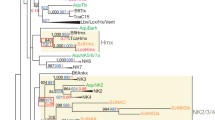
Data on nonbilaterian animals (sponges, cnidarians, and ctenophores) have suggested that Antennapedia (ANTP) class homeobox genes played a crucial role in the early diversification of animal body plans. Estimates of ancestral gene diversity within this important class of developmental regulators have been mostly based on recent analyses of the complete genome of a demosponge species, leading to the proposal that all ANTP families found in nonsponges animals (eumetazoans) derived from an ancestral “proto-NK” six-gene cluster. However, a single sponge species cannot reveal ancestral metazoan traits, in particular because lineage-specific gene duplications or losses are likely to have occurred during the long history of the Porifera. We thus looked for ANTP genes by degenerate polymerase chain reaction search in five species belonging to the Homoscleromorpha, a sponge lineage recently phylogenetically classified outside demosponges and characterized by unique histological features. We identified new genes of the ANTP class called HomoNK. Our phylogenetic analyses placed HomoNK (without significant support) close to the NK6 and NK7 families of cnidarian and bilaterian ANTP genes and did not recover the monophyly of the proposed “proto-NK” cluster. Our expression analyses of the HomoNK gene OlobNK in adult Oscarella lobularis showed that this gene is a strict marker of choanocytes, the most typical sponge cell type characterized by an apical flagellum surrounded by a collar of microvilli. These results are discussed in the light of the predominant neurosensory expression of NK6 and NK7 genes in bilaterians and of the recent proposal that choanocytes could be the sponge homologs of sensory cells.



Similar content being viewed by others
References
Alanentalo T, Chatonnet F, Karlen M, Sulniute R, Ericson J, Andersson E, Ahlgren U (2006) Cloning and analysis of Nkx6.3 during CNS and gastrointestinal development. Gene Expr Patterns 6:162–170
Atschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Aouachaeria A, Geourjon C, Aghajari N, Navratil V, Deléage G, Lethias C, Exposito JY (2006) Insights into early extracellular matrix evolution: Spongin short chain collagen-related proteins are homologous to basement membrane type IV collagens and form a novel family widely distributed in invertebrates. Mol Biol Evol 23:2288–2302
Borchiellini C, Manuel M, Alivon E, Boury-Esnault N, Vacelet J, Le Parco Y (2001) Sponge paraphyly and the origin of Metazoa. J Evol Biol 14:171–179
Borchiellini C, Chombard C, Manuel M, Alivon E, Vacelet J, Boury-Esnault N (2004) Molecular phylogeny of Demospongiae: implications for classification and scenarios of character evolution. Mol Phyl Evol 32:823–837
Boury-Esnault N, Jamieson BGM (1999) Porifera. In: Jamieson PGM (ed) Progress in male gamete ultrastructure and phylogeny. Wiley, Chichester, NY, pp 1–20
Boute N, Exposito JY, Boury-Esnault N, Vacelet J, Noro N, Miyazaki K, Yoshigato K, Garrone R (1996) Type IV collagen in sponges, the missing link in basement membrane ubiquity. Biol Cell 88:37–44
Chourrout D, Delsuc F, Chourrout P, Edvardsen RB, Rentzsch F, Renfer E, Jensen MF, Zhu B, de Jong P, Steele RE, Technau U (2006) Minimal ProtoHox cluster inferred from bilaterians and cnidarian Hox complements. Nature 442:684–687
Dellaporta SL, Xu A, Sagasser S, Jakob W, Moreno MA, Buss LW (2006) Mitochondrial genome of Trichoplax adhaerens supports Placozoa as the basal lower metazoan phylum. Proc Natl Acad Sci U S A 103:8751–8756
de Jong DM, Hislop NR, Hayward DC, Reece-Hoyes JS, Pontynen PC, Ball EE, Miller DJ (2006) Components of both major axial patterning systems of the Bilateria are differentially expressed along the primary axis of a ‘radiate’ animal, the anthozoan cnidarian Acropora millepora. Dev Biol 298:632–643
Derelle R, Manuel M (2007) Ancient connection between NKL genes and the mesoderm? Insights from Tlx expression in a ctenophore. Dev Genes Evol 214:253–261
Dunn CW, Hejnol A, Matus DQ, Pang K, Browne WE, Smith SA, Seaver E, Rouse GW, Obst M, Edgecombe GD, Sorensen MV, Haddock SH, Schmidt-Rhaesa A, Okusu A, Kristensen RM, Wheeler WC, Martindale MQ, Giribet G (2008) Broad phylogenomic sampling improves resolution of the animal tree of life. Nature 452:745–749
Ereskovsky AV, Tokina DB, Bézac C, Boury-Esnault N (2007) Metamorphosis of cinctoblastula larvae (Homoscleromorpha, Porifera). J Morphol 268:518–528
Erpenbeck D, Voigt O, Adamski M, Adamska M, Hooper JN, Worheide G, Degnan BM (2007) Mitochondrial diversity of early-branching metazoa is revealed by the complete mt genome of a haplosclerid demosponge. Mol Biol Evol 24:19–22
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Funayama N, Nakatsukasa M, Hayashi T, Agat K (2005) Isolation of the choanocyte in the fresh water sponge, Ephydatia fluviatilis and its lineage marker, Ef annexin. Dev Growth Differ 47:243–253
Fritzsch B, Beisel KW, Hansen LA (2006) The molecular basis of neurosensory cell formation in ear development: a blueprint for hair cell and sensory neuron regeneration. Bioessays 28:1181–1193
Gaino E, Manconi R, Pronzato R (1995) Organizational plasticity as a successful conservative tactics in sponges. Anim Biol 4:31–43
Garcia-Fernàndez J (2005) The genesis and evolution of homeobox gene clusters. Nat Rev Genet 6:881–892
Gilbert S (2006) Developmental biology, 8th edn. Sinauer, Sunderland, MA
Grimmelikhuijzen CJP, Westfall JA (1995) The nervous systems of cnidarians. In: Breidbach O, Kutsch W (eds) The nervous systems of invertebrates: an evolutionary and comparative approach. Birkhäuser, Basel, Switzerland, pp 7–24
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Hall TA (1999) Bioedit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hill A, Tetrault J, Hill M (2004) Isolation and expression analysis of a poriferan Antp-class Bar-/Bsh-like homeobox gene. Dev Genes Evol 214:515–523
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Huelsenbeck JP, Ronquist F, Nielsen R, Bollback JP (2001) Bayesian inference of phylogeny and its impact on evolutionary biology. Science 294:2310–2314
Hwang JS, Ohyanagi H, Hayakawa S, Osato N, Nishimiya-Fujisawa C, Ikeo K, David CN, Fujisawa T, Gojobori T (2007) The evolutionary emergence of cell type-specific genes inferred from the gene expression analysis of Hydra. Proc Natl Acad Sci U S A 104:14735–14740
Hyman LH (1940) The invertebrates, vol 1. Protozoa through Ctenophora. McGraw-Hill Book, New York and London
Jacobs DK, Nakanishi N, Yuan D, Camara A, Nichols SA, Hartenstein V (2007) Evolution of sensory structures in basal metazoa. Integr Comp Biol 47:712–723
Jager M, Quéinnec E, Houlison E, Manuel M (2006) Expansion of the SOX family predated the emergence of the Bilateria. Mol Phyl Evol 39:468–477
Jenner RA, Wills MA (2007) The choice of model organisms in evo–devo. Nat Rev 8:311–318
Kamm K, Schierwater B (2006) Ancient complexity of the non-hox ANTP gene complement in the Anthozoan Nematostella vectensis. Implications for the evolution of the ANTP superclass. J Exp Zool (Mol Dev Evol) 306B:589–596
King N, Westbrook MJ, Young SL, Kuo A, Abedin M, Chapman J, Fairclough S, Hellsten U, Isogai Y, Letunic I, Marr M, Pincus D, Putnam N, Rokas A, Wright KJ, Zuzow R, Dirks W, Good M, Goodstein D, Lemons D, Li W, Lyons JB, Morris A, Nichols S, Richter DJ, Salamov A, Sequencing JGI, Bork P, Lim WA, Manning G, Miller WT, McGinnis W, Shapiro H, Tjian R, Grigoriev IV, Rokhsar D (2008) The genome of the choanoflagellate Monosiga brevicollis and the origin of metazoans. Nature 451:783–788
Larroux C, Fahey B, Liubicich D, Hinman VF, Gauthier M, Gongora M, Green K, Wörheide G, Leys SP, Degnan BM (2006) Developmental expression of transcription factor genes in a demosponge: insights into the origin of metazoan multicellularity. Evol Dev 8:150–173
Larroux C, Fahey B, Degnan SM, Adamski M, Rokhsar DS, Degnan BM (2007) The NK homeobox gene cluster predates the origin of Hox genes. Curr Biol 17:1–5
Larroux C, Luke GN, Koopman P, Rokhsar DS, Shimeld SM, Degnan BM (2008) Genesis and expansion of metazoan transcription factor gene classes. Mol Biol Evol 25(5):980–996
Leys SP, Meech RW (2006) Physiology of coordination in sponges. Can J Zool 84:288–306
Maldonaldo M (2004) Choanoflagellates, choanocytes, and animal multicellularity. Invert Biol 123:1–22
Manuel M, Le Parco Y (2000) Homeobox gene diversification in the calcareous sponge Sycon raphanus. Mol Phyl Evol 17:97–107
Manuel M, Borchiellini C, Alivon E, Le Parco Y, Vacelet J, Boury-Esnault N (2003) Phylogeny and evolution of calcareous sponges: monophyly of Calcinea and Calcaronea, high level of morphological homoplasy, and the primitive nature of axial symmetry. Syst Biol 52:311–333
Martinez DE, Dirksen M-L, Bode PM, Jamrich M, Steele RE, Bode HR (1997) Budhead, a fork head/HNF-3 homologue, is expressed during axis formation and head specification in Hydra. Dev Biol 192:523–536
Medina M, Collins AG, Silberman JD, Sogin ML (2001) Evaluating hypotheses of basal animal phylogeny using complete sequences of large and small subunit rRNA. Proc Natl Acad Sci U S A 98:9707–9712
Murtha MT, Leckman JF, Ruddle FH (1991) Detection of homeobox genes in development and evolution. Proc Natl Acad Sci U S A 88:10711–10715
Nam J, Nei M (2005) Evolutionary change of the numbers of homeobox genes in bilateral animals. Mol Biol Evol 22:2386–2394
Nielsen (2001) Animal evolution: interrelationships of the living Phyla, 2nd edn. Oxford University Press, Oxford
Nielsen C (2008) Six major steps in animal evolution: are we derived sponge larvae. Evol Dev 10:241–257
Nichols SA (2005) An evaluation of support for order-level monophyly and interrelationships within the class Demospongiae using partial data from the large subunit rDNA and cytochrome oxidase subunit I. Mol Phyl Evol 34:81–96
Peterson KJ, Sperling EA (2007) Poriferan ANTP genes: primitively simple or secondarily reduced? Evol Dev 9:405–408
Putnam NH, Srivastava M, Hellsten U, Dirks B, Chapman J, Salamov A, Terry A, Shapiro H, Lindquist E, Kapitonov VV, Jurka J, Genikhovich G, Grigoriev IV, Lucas SM, Steele RE, Finnerty JR, Technau U, Martindale MQ, Rokhsar DS (2007) Sea anemone genome reveals ancestral eumetazoan gene repertoire and genomic organization. Science 317:86–94
Richelle-Maurer E, Van de Vyver G (1999) Temporal and spatial expression of EmH-3, a homeobox-containing gene isolated from the freshwater sponge Ephydatia muelleri. Mech Ageing Dev 109:203–219
Richelle-Maurer E, Boury-Esnault N, Itskovich VB, Manuel M, Pomponi SA, Van de Vyver G, Borchiellini C (2006) Conservation and phylogeny of a novel class of non-Hox genes of the Antp-superclass in Demospongiae (Porifera). J Mol Evol 63:222–230
Ryan JF, Burton PM, Mazza ME, Kwong GK, Mullikin JC, Finnerty JR (2006) The cnidarian–bialterian ancestor possessed at least 56 homeoboxes: evidence from the starlet sea anemone, Nematostella vectensis. Genome Biol 7:R64
Sperling EA, Pisani D, Peterson KJ (2007) Poriferan paraphyly and its implications for Precambrian palaeobiology. Geol Soc London Spec Publ 286:355–368
Swofford DL (2000) PAUP*. Phylogenetic analysis using parsimony (*and other methods). Version 4. Sinauer, Sunderland, Massachusetts
Takahashi H, Kamiya A, Ishiguro A, Suzuki AC, Saitou N, Toyoda A, Aruga J (2008) Conservation and diversification of msx protein in metazoan evolution. Mol Biol Evol 25:69–82
Telford MJ (2006) Animal phylogeny. Curr Biol 16:981–985
Uhler J, Garbern J, Yang L, Kamholz J, Mellerick DM (2002) Nk6, a novel Drosophila homeobox gene regulated by vnd. Mech Dev 116:105–116
Wang X, Lavrov DV (2007) Mitochondrial genome of the Homoscleromorph Oscarella carmela (Porifera, Demospongiae) reveals unexpected complexity in the common ancestor of sponges and others animals. Mol Biol Evol 24:363–373
Wheland S, Goldman N (2001) A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol Biol Evol 18:691–699
Zhao S, Jiang H, Wang W, Mao B (2007) Cloning and developmental expression of the Xenopus Nkx6 genes. Dev Genes Evol 217:477–483
Acknowledgments
We thank to the diving staff of the COM (Roland Graille, Frederic Zuberer, and Bernard De Ligondes) and Christophe Lejeusne for their help in sampling. We also thank Alexander Ereskovsky for helpful comments, Christian Marschal for his help in the use of the microscope and binocular magnifier and also Dr Clive Wilkinson and Michael Paul for help with the English revision.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by: M.Q. Martindale
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material S1
Sequences of the primers used. (DOC 27.0 KB)
Supplementary material S2
(DOC 65.5 KB)
Supplementary material S3
Homeodomain sequences used for phylogenetic analysis, with accession numbers. (RTF 148 KB)
Rights and permissions
About this article
Cite this article
Gazave, E., Lapébie, P., Renard, E. et al. NK homeobox genes with choanocyte-specific expression in homoscleromorph sponges. Dev Genes Evol 218, 479–489 (2008). https://doi.org/10.1007/s00427-008-0242-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-008-0242-z