Abstract
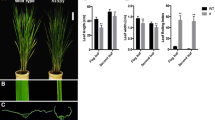
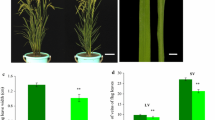
Appropriate leaf shape has proved to be useful in improving photosynthesis and increasing grain yield. To understand the molecular mechanism of leaf morphogenesis, we identified a rice mutant nrl1, which was characterized by a phenotype of narrow and rolled leaves. Microscopic observation showed that the mutation significantly decreased the number of vascular bundles of leaf and stem. Genetic analysis revealed that the mutation was controlled by a single nuclear-encoded recessive gene. To isolate the nrl1 gene, 756 F2 and F3 mutant individuals from a cross of the nrl1 mutant with Longtepu were used and a high-resolution physical map of the chromosomal region around the nrl1 gene was made. Finally, the gene was mapped in 16.5 kb region between marker RL21 and marker RL36 within the BAC clone OSJNBa0027H05. Cloning and sequencing of the target region from the mutant showed that there was a 58 bp deletion within the second exon of the cellulose synthase-like D4 gene (TIGR locus Os12g36890). The nrl1 mutation was rescued by transformation with the wild-type cellulose synthase-like D4 gene. Accordingly, the cellulose synthase-like D4 gene was identified as the NRL1 gene. NRL1 was transcribed in various tissues and was mainly expressed in panicles and internodes. NAL7 and SLL1 were found to be upregulated, whereas OsAGO7 were downregulated in the nrl1 mutant. These findings suggested that there might be a functional association between these genes in regulating leaf development.







Similar content being viewed by others
Abbreviations
- CSL:
-
Cellulose synthase like
- InDel:
-
Insertion–deletion
- nrl1 :
-
Narrow and rolled leaves 1
References
Bernal AJ, Jensen JK, Harholt J, Sorensen S, Moller I, Blaukopf C, Johansen B, de Lotto R, Pauly M, Scheller HV, Willats WG (2007) Disruption of ATCSLD5 results in reduced growth, reduced xylan and homogalacturonan synthase activity and altered xylan occurrence in Arabidopsis. Plant J 52:791–802
Bernal AJ, Yoo CM, Mutwil M, Jensen JK, Hou G, Blaukopf C, Sorensen I, Blancaflor EB, Scheller HV, Willats WGT (2008) Functional analysis of the cellulose synthase like genes CSLD1, CSLD2 and CSLD4 in tip growing Arabidopsis cells. Plant Physiol 148:1238–1253
Byrne ME, Barley R, Curtis M, Arroyo JM, Dunham M, Hudson A, Martienssen RA (2000) Asymmetric leaves 1 mediates leaf patterning and stem cell function in Arabidopsis. Nature 408:967–971
Cocuron JC, Lerouxel O, Drakakaki G, Alonso AP, Liepman AH, Keegstra K, Raikhel N, Wilkerson C (2007) A gene from the cellulose synthase-like C family encodes β-1, 4 glucan synthase. Proc Natl Acad Sci USA 104:8550–8555
Dhugga KS, Barreiro R, Whitten B, Stecca K, Hazebroek J, Randhawa GS, Dolan M, Kinney AJ, Tomes D, Nichols S, Anderson P (2004) Guar seed β-mannan synthase is a member of the cellulose synthase super gene family. Science 303:363–366
Eshed Y, Baum SF, Perea JV, Bowman JL (2001) Establishment of polarity in lateral organs of plants. Curr Biol 11:1251–1260
Favery B, Ryan E, Foreman J, Linstead P, Boudonck K, Steer M, Shaw P, Dolan L (2001) KOJAK encodes a cellulose synthase-like protein required for root hair cell morphogenesis in Arabidopsis. Genes Dev 15:79–89
Fujino K, Matsuda Y, Ozawa K, Nishimura T, Koshiba T, Fraaije MW, Sekiguchi H (2008) NARROW LEAF 7 controls leaf shape mediated by auxin in rice. Mol Genet Genomics 279:499–507
Hibara KI, Obara M, Hayashida E, Abe M, Ishimaru T, Satoh H, Itoh JI, Nagato Y (2009) The ADAXIALIZED LEAF1 gene functions in leaf and embryonic pattern formation in rice. Dev Biol 334:345–354
Horiguchi G, Kim GT, Tsukaya H (2005) The transcription factor AtGRF5 and the transcription coactivator AN3 regulate cell proliferation in leaf primordia of Arabidopsis thaliana. Plant J 43:68–78
Huang W, Pi L, Liang W, Xu B, Wang H, Cai R, Huang H (2006) The proteolytic function of the Arabidopsis 26S proteasome is required for specifying leaf adaxial identity. Plant Cell 18:2479–2492
Kerstetter RA, Bollman K, Taylor RA, Bomblies K, Poethig RS (2001) KANADI regulates organ polarity in Arabidopsis. Nature 411:706–709
Kim GT, Tsukaya H, Uchimiya H (1998) The ROTUNDIFOLIA3 gene of Arabidopsis thaliana encodes a new member of the cytochrome P-450 family that is required for the regulated polar elongation of leaf cells. Genes Dev 12:2381–2391
Kim GT, Shoda K, Tsuge T, Cho KH, Uchimiya H, Yokoyama R, Nishitani K, Tsukaya H (2002) The ANGUSTIFOLIA gene of Arabidopsis, a plant CtBP gene, regulates leaf-cell expansion, the arrangement of cortical microtubules in leaf cells and expression of a gene involved in cell-wall formation. EMBO J 21:1267–1279
Kim CM, Park SH, Je BI, Park SH, Park SJ, Piao HL, Eun MY, Dolan L, Han C (2007) OsCSLD1, a cellulose synthase-like D1 gene, is required for root hair morphogenesis in rice. Plant Physiol 143:1220–1230
Liepman AH, Wilkerson CG, Keegstra K (2005) Expression of cellulose synthase-like (Csl) genes in insect cells reveals that CslA family members encode mannan synthases. Proc Natl Acad Sci USA 102:2221–2226
Liu W, Fu Y, Hu G, Si H, Zhu L, Wu C, Sun Z (2007) Identification and fine mapping of a thermo-sensitive chlorophyll deficient mutant in rice (Oryza sativa L.). Planta 226:785–795
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408
Long JA, Moan EI, Medford JI, Barton MK (1996) A member of the KNOTTED class of homeodomain proteins encoded by the STM gene of Arabidopsis. Nature 379:66–69
McConnell JR, Emery J, Eshed Y, Bao N, Bowman J, Barton MK (2001) Role of PHABULOSA and PHAVOLUTA in determining radial patterning in shoots. Nature 411:709–713
McCouch SR, Teytelman L, Xu YB, Lobos KB, Clare K, Walton M, Fu B, Maghirang R, Li Z, Xing Y, Zhang Q, Kono I, Yano M, Fjellstrom R, DeClerck G, Schneider D, Cartinhour S, Ware D, Stein L (2002) Development and mapping of 2240 new SSR markers for rice (Oryza sativa L.). DNA Res 9:199–207
Micol JL (2009) Leaf development: time to turn over a new leaf? Curr Opin Plant Biol 12:9–16
Nakai K, Horton P (1999) PSORT: a program for detecting sorting signals in proteins and predicting their subcellular localization. Trends Biochem Sci 24:34–36
Narita NN, Moore S, Horiguchi G, Kubo M, Demura T, Fukuda H, Goodrich J, Tsukaya H (2004) Overexpression of a novel small peptide ROTUNDIFOLIA4 decreases cell proliferation and alters leaf shape in Arabidopsis thaliana. Plant J 38:699–713
Otsuga D, DeGuzman B, Prigge MJ, Drews GN, Clark SE (2001) REVOLUTA regulates meristem initiation at lateral positions. Plant J 25:223–236
Panaud O, Chen X, McCouch SR (1996) Development of microsatellite markers and characterization of simple sequence length polymorphism (SSLP) in rice (Oryza sativa L.). Mol Genet Genomics 252:597–607
Pekker I, Alvarez JP, Eshed Y (2005) Auxin response factors mediate Arabidopsis organ asymmetry via modulation of KANADI activity. Plant Cell 17:2899–2910
Qi J, Qian Q, Bu Q, Li S, Chen Q, Sun J, Liang W, Zhou Y, Chu C, Li X, Ren F, Palme K, Zhao B, Chen J, Chen M, Li C (2008) Mutation of the rice Narrow leaf1 gene, which encodes a novel protein, affects vein patterning and polar auxin transport. Plant Physiol 147:1947–1959
Shi Z, Wang J, Wan X, Shen G, Wang X, Zhang J (2007) Over-expression of rice OsAGO7 gene induces upward curling of the leaf blade that enhanced erect-leaf habit. Planta 226:99–108
Small I, Peeters N, Legeai F, Lurin C (2004) Predotar: a tool for rapidly screening proteomes for N-terminal targeting sequences. Proteomics 4:1581–1590
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Temnykh S, DeClerck G, Lukashova A, Lipovich L, Cartinhour S, McCouch S (2001) Computational and experimental analysis of microsatellites in rice (Oryza sativa L.): frequency, length, variation, transposon associations, and genetic marker potential. Genome Res 11:1441–1452
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tsiantis M, Schneeberger R, Golz JF, Freeling M, Langdale JA (1999) The maize rough sheath2 gene and leaf development programs in monocot and dicot plants. Science 284:154–156
Tsukaya H (2006) Mechanism of leaf-shape determination. Annu Rev Plant Biol 57:477–496
Waites R, Selvadurai HR, Oliver IR, Hudson A (1998) The PHANTASTICA gene encodes a MYB transcription factor involved in growth and dorsoventrality of lateral organs in Antirrhinum. Cell 93:779–789
Wang MB, Waterhouse PM (1997) A rapid and simple method of assaying plants transformed with hygromycin or PPT resistance genes. Plant Mol Biol Rep 15:209–215
Wang X, Cnops G, Vanderhaeghen R, De Block S, Van Montagu M, Van Lijsebettens M (2001) AtCSLD3, a cellulose synthase-like gene important for root hair growth in Arabidopsis. Plant Physiol 126:575–586
Woo YM, Park HJ, Su’udi M, Yang JI, Park JJ, Back K, Park YM, An G (2007) Constitutively wilted 1, a member of the rice YUCCA gene family, is required for maintaining water homeostasis and an appropriate root to shoot ratio. Plant Mol Biol 65:125–136
Xu L, Xu Y, Dong A, Sun Y, Pi L, Xu Y, Huang H (2003) Novel as1 and as2 defects in leaf adaxial-abaxial polarity reveal the requirement for ASYMMETRIC LEAVES1 and 2 and ERECTA functions in specifying leaf adaxial identity. Development 130:4097–4107
Yan S, Yan CJ, Zeng XH, Yang YC, Fang YW, Tian CY, Sun YW, Cheng ZK, Gu MH (2008) ROLLED LEAF 9, encoding a GARP protein, regulates the leaf abaxial cell fate in rice. Plant Mol Biol 68:239–250
Yin Y, Huang J, Xu Y (2009) The cellulose synthase superfamily in fully sequenced plants and algae. BMC Plant Biol 9:99
Zhang GH, Xu Q, Zhu XD, Qian Q, Xue HW (2009) SHALLOT-LIKE1 is a KANADI transcription factor that modulates rice leaf rolling by regulating leaf abaxial cell development. Plant Cell 21:719–735
Zhou GK, Kubo M, Zhong R, Demura T, Ye ZH (2007) Overexpression of miR165 affects apical meristem formation, organ polarity establishment and vascular development in Arabidopsis. Plant Cell Physiol 48:391–404
Zhu ZG, Xiao H, Fu YP, Hu GC, Yu YH, Si HM, Zhang JL, Sun ZX (2001) Construction of transgenic rice populations by inserting the maize transposon Ac/Ds and genetic analysis for several mutants. Chinese J Biotech 17:288–292 (in Chinese with English abstract)
Acknowledgments
This project was supported by National Natural Science Foundation of China (30900790), the National High Technology Research and Development Program of China (2006AA10Z1E8) and Zhejiang Natural Science Foundation (Y307070 and 2008C22077). We are grateful to Mrs Honglan Yan (China National Rice Research Institute, China) for taking pictures for the article.
Author information
Authors and Affiliations
Corresponding authors
Additional information
The sequences in this article can be found in NCBI under the following accession numbers: OsACT1 (NM_001057621), NRL1 (AK242601), SLL1 (FJ268748) NAL1 (EU093963), NAL7 (AB354302) and OsAGO7 (EF486281). The accession numbers of proteins used in the phylogeny are shown in Supplemental Data Set 1.
C. Wu, Y. Fu contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wu, C., Fu, Y., Hu, G. et al. Isolation and characterization of a rice mutant with narrow and rolled leaves. Planta 232, 313–324 (2010). https://doi.org/10.1007/s00425-010-1180-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-010-1180-3