Abstract
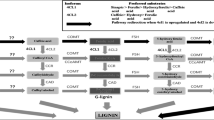
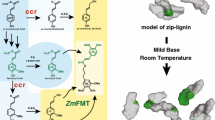
Red clover (Trifolium pratense) leaves accumulate several μmol of phaselic acid [2-O-caffeoyl-l-malate] per gram fresh weight. Post-harvest oxidation of such o-diphenols to o-quinones by endogenous polyphenol oxidases (PPO) prevents breakdown of forage protein during storage. Forages like alfalfa (Medicago sativa) lack both foliar PPO activity and o-diphenols. Consequently, breakdown of their protein upon harvest and storage results in economic losses and release of excess nitrogen into the environment. Understanding how red clover synthesizes o-diphenols such as phaselic acid will help in the development of forages utilizing this natural system of protein protection. We have proposed biosynthetic pathways in red clover for phaselic acid that involve a specific hydroxycinnamoyl-CoA:malate hydroxycinnamoyl transferase. It is unclear whether the transfer reaction to malate to form phaselic acid involves caffeic acid or p-coumaric acid and subsequent hydroxylation of the resulting p-coumaroyl-malate. The latter would require a coumarate 3′-hydroxylase (C3′H) capable of hydroxylating p-coumaroyl-malate, an activity not previously described. Here, a cytochrome P450 C3′H (CYP98A44) was identified and its gene cloned from red clover. CYP98A44 shares 96 and 79% amino acid identity with Medicago truncatula and Arabidopsis thaliana C3′H proteins that are capable of hydroxylating p-coumaroyl-shikimate and have been implicated in monolignol biosynthesis. CYP98A44 mRNA is expressed in stems and flowers and to a lesser extent in leaves. Immune serum raised against CYP98A44 recognizes a membrane-associated protein in red clover stems and leaves and cross-reacts with C3′H proteins from other species. CYP98A44 expressed in Saccharomyces cerevisiae is capable of hydroxylating p-coumaroyl-shikimate, but not p-coumaroyl-malate. This finding indicates that in red clover, phaselic acid is likely formed by transfer of a caffeoyl moiety to malic acid, although the existence of a second C3′H capable of hydroxylating p-coumaroyl-malate cannot be definitively ruled out.





Similar content being viewed by others
Abbreviations
- 4CL:
-
4-Coumarate:CoA ligase
- C3′H:
-
Coumarate 3′-hydroxylase
- C4H:
-
Cinnamate-4-hydroxylase
- CYP:
-
Cytochrome P450
- HCT:
-
Hydroxycinnamoyl-CoA hydroxycinnamoyl transferase
- PAL:
-
Phenylalanine ammonia lyase
- PCR:
-
Polymerase chain reaction
- PPO:
-
Polyphenol oxidase
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
- SDS-PAGE:
-
Sodium dodecyl sulfate-polyacrylamide gel electrophoresis
- SMT:
-
Sinapoyl-glucose:malate synapoyl transferase
References
Ausubel FM, Brent R, Kingston RE, Moore DD, Deidman JG, Smith JA, Struhl K (eds) (1998) Current protocols in molecular biology. Wiley, New York
Franke R, Humphreys JM, Hemm MR, Denault JW, Ruegger MO, Cusumano JC, Chapple C (2002) The Arabidopsis REF8 gene encodes the 3-hydroxylase of phenylpropanoid metabolism. Plant J 30:33–45
Gang DR, Beuerle T, Ullmann P, Werck-Reichart D, Pichersky E (2002) Differential production of meta-hydroxylated phenlypropanoids in sweet basil peltate gladular trichomes and leaves is controlled by the activities of specific acyltransferases and hydroxylases. Plant Physiol 130:1536–1544
Gietz RD, Woods RA (2002) Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. Methods Enzymol 350:87–96
Grawe W, Bachhuber P, Mock HP, Strack D (1992) Purification and characterization of sinapolyglucose—malate sinapoyltransferase from Raphanus sativus L. Planta 187:236–241
Harlow E, Lane D (1988) Antibodies: a laboratory manual. Cold Spring Harbor Laboratory, New York, p 508
Hemmerle H, Burger HJ, Below P, Schubert G, Rippel R, Schindler PW, Paulus E, Herling AW (1997) Chlorogenic acid and synthetic chlorogenic acid derivatives: novel inhibitors of hepatic glucose-6-phosphate translocase. J Med Chem 40:137–145
Hulo N, Bairoch A, Bulliard V, Cerutti L, Cuche BA, de Castro E, Lachaize C, Langendijk-Genevaux PS, Sigrist CJA (2008) The 20 years of PROSITE. Nucleic Acids Res 36:D245–D249
Lee MRF, Winters AL, Scollan ND, Dewhurst RJ, Theodorou MK, Minchin FR (2004) Plant-mediated lipolysis and proteolysis in red clover with different polyphenol oxidase activities. J Sci Food Agric 84:1639–1645
Lehfeldt C, Shirley AM, Meyer K, Ruegger MO, Cusumano JC, Viitanen PV, Strack D, Chapple C (2000) Cloning of the SNG1 gene of arabidopsis reveals a role for a serine carboxypeptidase-like protein as an acyltransferase in secondary metabolism. Plant Cell 12:1295–1306
Mahesh V, Million-Rousseau R, Ullmann P, Chabrillange N, Bustamante J, Mondolot L, Morant M, Noirot M, Hamon S, de Kochko A, Werck-Reichhart D, Campa C (2007) Functional characterization of two p-coumaroyl ester 3′-hydroxylase genes from coffee tree: evidence of a candidate for chlorogenic acid biosynthesis. Plant Mol Biol 64:145–159
Matsuno M, Nagatsu A, Ogihara Y, Ellis BE, Mizukami H (2002) CYP98A6 from Lithospermum erythrorhizon encodes 4-coumaroyl-4′-hydroxyphenyllactic acid 3-hydroxylase involved in rosmarinic acid biosynthesis. FEBS Lett 514:219–224
Morant M, Schoch GA, Ullmann P, Ertunc T, Little D, Olsen CE, Petersen M, Negrel J, Werck-Reichhart D (2007) Catalytic activity, duplication and evolution of the CYP98 cytochrome P450 family in wheat. Plant Mol Biol 63:1–19
Niggeweg R, Michael AJ, Martin C (2004) Engineering plants with increased levels of the antioxidant chlorogenic acid. Nat Biotechnol 22:746–754
Pompon D, Louerat B, Bronine A, Urban P (1996) Yeast expression of animal and plant P450s in optimized redox environments. Methods Enzymol 272:51–64
Reddy MSS, Chen F, Shadle G, Jackson L, Aljoe H, Dixon RA (2005) Targeted down-regulation of cytochrome P450 enzymes for forage quality improvement in alfalfa (Medicago sativa L.). Proc Natl Acad Sci USA 102:16573–16578
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning, a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Sato S, Isobe S, Asamizu E, Ohmido N, Kataoka R, Nakamura Y, Kaneko T, Sakurai N, Okumura K, Klimenko I, Sasamoto S, Wada T, Watanabe A, Kohara M, Fujishiro T, Tabata S (2005) Comprehensive structural analysis of the genome of red clover (Trifolium pratense L.). DNA Res 12:301–364
Schaller GE, Dewitt ND (1995) Analysis of the H+-ATPase and other proteins of the arabidopsis plasma membrane. Methods Cell Biol 50:129–148
Schoch G, Goepfert S, Morant M, Hehn A, Meyer D, Ullmann P, Werck-Reichhart D (2001) CYP98A3 from Arabidopsis thaliana is a 3′-hydroxylase of phenolic esters, a missing link in the phenylpropanoid pathway. J Biol Chem 276:36566–36574
Schoch GA, Morant M, Abdulrazzak N, Asnaghi C, Goepfert S, Petersen M, Ullmann P, Werck-Reichhart D (2006) The meta-hydroxylation step in the phenylpropanoid pathway: a new level of complexity in the pathway and its regulation. Environ Chem Lett 4:127–136
Shadle G, Chen F, Reddy MSS, Jackson L, Nakashima J, Dixon RA (2007) Down-regulation of hydroxycinnamoyl CoA: shikimate hydroxycinnamoyl transferase in transgenic alfalfa affects lignification, development and forage quality. Phytochemistry 68:1521–1529
Sharma V, Strack D (1985) Vacuolar localization of 1-sinapoylglucose—l-malate sinapoyltransferase in protoplasts from cotyledons of Raphanus sativus. Planta 163:563–568
Smith RR, Maxwell DP (1980) Registration of WI-1 and WI-2 red clover. Crop Sci 20:831
Steffens JC, Harel E, Hunt MD (1994) Polyphenol oxidase. In: Ellis BE, Kuroki GW, Stafford HA (eds) Genetic engineering of plant secondary metabolism. Plenum Press, New York, pp 275–312
Stukkens Y, Bultreys A, Grec S, Trombik T, Vanham D, Boutry M (2005) NpPDR1, a pleiotropic drug resistance-type ATP-binding cassette transporter from Nicotiana plumbaginifolia, plays a major role in plant pathogen defense. Plant Physiol 139:341–352
Sullivan ML (2009a) A novel red clover hydroxycinnamoyl transferase has enzymatic activities consistent with a role in phaselic acid biosynthesis. Plant Physiol 150:1866–1879
Sullivan ML (2009b) Phenylalanine ammonia lyase genes in red clover: expression in whole plants and in response to yeast fungal elicitor. Biol Plant 53:301–306
Sullivan ML, Hatfield RD (2006) Polyphenol oxidase and o-diphenols inhibit postharvest proteolysis in red clover and alfalfa. Crop Sci 46:662–670
Sullivan ML, Hatfield RD, Thoma SL, Samac DA (2004) Cloning and characterization of red clover polyphenol oxidase cDNAs and expression of active protein in Escherichia coli and transgenic alfalfa. Plant Physiol 136:3234–3244
Taylor CB, Bariola PA, Delcardayre SB, Raines RT, Green PJ (1993) Rns2—a senescence-associated RNase of arabidopsis that diverged from the S-RNases before speciation. Proc Natl Acad Sci USA 90:5118–5122
Zdobnov EM, Apweiler R (2001) InterProScan—an integration platform for the signature-recognition methods in InterPro. Bioinformatics 17:847–848
Acknowledgments
This work was supported by the US Department of Agriculture-Cooperative State Research, Education, and Extension Service-National Research Initiative Competitive Grants Program (Grant no. 2009-35318-05048). We wish to thank Sara Zerbel, Lisa Koch, and Kendal Cooper for excellent technical assistance; Paul Schatz and John Ralph for providing hydroxycinnamoyl ester standards; Clint Chapple, Jing-Ke Weng, Dave Gang, and Anna Berim for providing P450 microsome samples; Philippe Urban and Denis Pompon for providing pYeDP60 and WAT11; Sheryl Rakowski and Marcin Filutowicz for access to ultracentrifugation equipment; and Jane Marita and Ron Hatfield for technical advice and helpful discussions regarding this work. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the US Department of Agriculture.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sullivan, M.L., Zarnowski, R. Red clover coumarate 3′-hydroxylase (CYP98A44) is capable of hydroxylating p-coumaroyl-shikimate but not p-coumaroyl-malate: implications for the biosynthesis of phaselic acid. Planta 231, 319–328 (2010). https://doi.org/10.1007/s00425-009-1054-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-009-1054-8