Abstract
Histone acetylation is involved in the regulation of chromatin structure and gene function. We reported previously that histone H3 acetylation pattern is subject to dynamic changes and limited to certain stages of germ cell differentiation during murine spermatogenesis, suggesting a crucial role for acetylation in the process. In the present study, we investigated the effects of hyper- and hypo-acetylation on spermatogenesis. Changes in acetylation level were induced by either in vivo administration of sodium phenylbutyrate, a histone deacetylase inhibitor, or by knockdown of histone acetyltransferases using short hairpin RNA plasmids transfection. Administration of sodium phenylbutyrate induced accumulation of acetylated histone H3 at lysine 9 and lysine 18 in round spermatids, together with spermatid morphological abnormalities and induction of apoptosis through a Bax-related pathway. Knockdown of steroid receptor coactivator 1, a member of histone acetyltransferases, but not general control of amino acid synthesis 5 nor elongator protein 3 by in vivo electroporation of shRNA plasmids, reduced acetylated histone H3 at lysine 9 in round spermatids, and induced morphological abnormalities. We concluded that the proper regulation of histone H3 acetylation levels is important for spermatid differentiation and complex chromatin remodeling during spermiogenesis.











Similar content being viewed by others
References
Berger SL (2002) Histone modifications in transcriptional regulation. Curr Opin Genet Dev 12(2):142–148. doi:10.1016/S0959-437X(02)00279-4
Boivin AJ, Momparler LF, Hurtubise A, Momparler RL (2002) Antineoplastic action of 5-aza-2′-deoxycytidine and phenylbutyrate on human lung carcinoma cells. Anticancer Drugs 13(8):869–874. doi:10.1097/00001813-200209000-00013
Butler LM, Zhou X, Xu WS, Scher HI, Rifkind RA, Marks PA, Richon VM (2002) The histone deacetylase inhibitor SAHA arrests cancer cell growth, up-regulates thioredoxin-binding protein-2, and down-regulates thioredoxin. Proc Natl Acad Sci USA 99(18):11700–11705. doi:10.1073/pnas.182372299
Calvaruso G, Carabillo M, Giuliano M, Lauricella M, D’Anneo A, Vento R, Tesoriere G (2001) Sodium phenylbutyrate induces apoptosis in human retinoblastoma Y79 cells: the effect of combined treatment with the topoisomerase I-inhibitor topotecan. Int J Oncol 18(6):1233–1237. doi:10.3892/ijo.18.6.1233
Cervoni N, Szyf M (2001) Demethylase activity is directed by histone acetylation. J Biol Chem 276(44):40778–40787. doi:10.1074/jbc.M103921200
Creasy DM, Ford GR, Gray TJ (1990) The morphogenesis of cyclohexylamine-induced testicular atrophy in the rat: in vivo and in vitro studies. Exp Mol Pathol 52(2):155–169. doi:10.1016/0014-4800(90)90001-T
Damavandi E, Hishikawa Y, Izumi S-I, Shin M, Koji T (2002) Involvement of Bax redistribution in the induction of germ cell apoptosis in neonatal mouse testes. Acta Histochem Cytochem 35(6):449–459. doi:10.1679/aohc.66.1
de Ruijter AJ, van Gennip AH, Caron HN, Kemp S, van Kuilenburg AB (2003) Histone deacetylases (HDACs): characterization of the classical HDAC family. Biochem J 370(Pt 3):737–749. doi:10.1042/BJ20021321
Delfino F, Walker WH (1998) Stage-specific nuclear expression of NF-kappaB in mammalian testis. Mol Endocrinol 12(11):1696–1707. doi:10.1210/mend.12.11.0194
Delmas V, van der Hoorn F, Mellstrom B, Jegou B, Sassone-Corsi P (1993) Induction of CREM activator proteins in spermatids: down-stream targets and implications for haploid germ cell differentiation. Mol Endocrinol 7(11):1502–1514. doi:10.1210/mend.7.11.8114765
Detich N, Bovenzi V, Szyf M (2003) Valproate induces replication-independent active DNA demethylation. J Biol Chem 278(30):27586–27592. doi:10.1074/jbc.M303740200
Feinman R, Clarke KO, Harrison LE (2002) Phenylbutyrate-induced apoptosis is associated with inactivation of NF-kappaB IN HT-29 colon cancer cells. Cancer Chemother Pharmacol 49(1):27–34. doi:10.1007/s00280-001-0390-6
Gavrieli Y, Sherman Y, Ben-Sasson SA (1992) Identification of programmed cell death in situ via specific labeling of nuclear DNA fragmentation. J Cell Biol 119(3):493–501. doi:10.1083/jcb.119.3.493
Giles RH, Peters DJ, Breuning MH (1998) Conjunction dysfunction: CBP/p300 in human disease. Trends Genet 14(5):178–183. doi:10.1016/S0168-9525(98)01438-3
Govin J, Lestrat C, Caron C, Pivot-Pajot C, Rousseaux S, Khochbin S (2006) Histone acetylation-mediated chromatin compaction during mouse spermatogenesis. Ernst Schering Res Found Workshop 57:155–172. doi:10.1007/3-540-37633-X_9
Hazzouri M, Pivot-Pajot C, Faure AK, Usson Y, Pelletier R, Sele B, Khochbin S, Rousseaux S (2000) Regulated hyperacetylation of core histones during mouse spermatogenesis: involvement of histone deacetylases. Eur J Cell Biol 79(12):950–960. doi:10.1078/0171-9335-00123
Iannitti T, Palmieri B (2011) Clinical and experimental applications of sodium phenylbutyrate. Drugs R D 11(3):227–249. doi:10.2165/11591280-000000000-00000
Kapushesky M, Adamusiak T, Burdett T, Culhane A, Farne A, Filippov A, Holloway E, Klebanov A, Kryvych N, Kurbatova N, Kurnosov P, Malone J, Melnichuk O, Petryszak R, Pultsin N, Rustici G, Tikhonov A, Travillian R, Williams E, Zorin A, Parkinson H, Brazma A (2012) Gene expression atlas update—a value-added database of microarray and sequencing-based functional genomics experiments. Nucleic Acids Res 40:D1077–D1081. doi:10.1093/nar/gkr913
Koji T, Kondo S, Hishikawa Y, An S, Sato Y (2008) In situ detection of methylated DNA by histo endonuclease-linked detection of methylated DNA sites: a new principle of analysis of DNA methylation. Histochem Cell Biol 130(5):917–925. doi:10.1007/s00418-008-0487-7
Kondo Y, Shen L, Yan PS, Huang TH, Issa JP (2004) Chromatin immunoprecipitation microarrays for identification of genes silenced by histone H3 lysine 9 methylation. Proc Natl Acad Sci USA 101(19):7398–7403. doi:10.1073/pnas.0306641101
Kopera IA, Bilinska B, Cheng CY, Mruk DD (2010) Sertoli-germ cell junctions in the testis: a review of recent data. Philos Trans R Soc Lond B Biol Sci 365(1546):1593–1605. doi:10.1098/rstb.2009.0251
Kuo MH, Allis CD (1998) Roles of histone acetyltransferases and deacetylases in gene regulation. BioEssays 20(8):615–626. doi:10.1002/(SICI)1521-1878(199808)20:8<615:AID-BIES4>3.0.CO;2-H
Lee B, Rhead W, Diaz GA, Scharschmidt BF, Mian A, Shchelochkov O, Marier JF, Beliveau M, Mauney J, Dickinson K, Martinez A, Gargosky S, Mokhtarani M, Berry SA (2010) Phase 2 comparison of a novel ammonia scavenging agent with sodium phenylbutyrate in patients with urea cycle disorders: safety, pharmacokinetics and ammonia control. Mol Genet Metab 100(3):221–228. doi:10.1016/j.ymgme.2010.03.014
Lin YN, Matzuk MM (2005) High-throughput discovery of germ-cell-specific genes. Semin Reprod Med 23(3):201–212. doi:10.1055/s-2005-872448
Louis M, Rosato RR, Brault L, Osbild S, Battaglia E, Yang XH, Grant S, Bagrel D (2004) The histone deacetylase inhibitor sodium butyrate induces breast cancer cell apoptosis through diverse cytotoxic actions including glutathione depletion and oxidative stress. Int J Oncol 25(6):1701–1711. doi:10.3892/ijo.25.6.1701
Meister G, Tuschl T (2004) Mechanisms of gene silencing by double-stranded RNA. Nature 431(7006):343–349. doi:10.1038/nature02873
Milkevitch M, Shim H, Pilatus U, Pickup S, Wehrle JP, Samid D, Poptani H, Glickson JD, Delikatny EJ (2005) Increases in NMR-visible lipid and glycerophosphocholine during phenylbutyrate-induced apoptosis in human prostate cancer cells. Biochim Biophys Acta 1734(1):1–12. doi:10.1016/j.bbalip.2005.01.008
Mizzen CA, Yang XJ, Kokubo T, Brownell JE, Bannister AJ, Owen-Hughes T, Workman J, Wang L, Berger SL, Kouzarides T, Nakatani Y, Allis CD (1996) The TAF(II)250 subunit of TFIID has histone acetyltransferase activity. Cell 87(7):1261–1270. doi:10.1016/S0092-8674(00)81821-8
Munshi A, Shafi G, Aliya N, Jyothy A (2009) Histone modifications dictate specific biological readouts. J Genet Genomics 36(2):75–88. doi:10.1016/S1673-8527(08)60094-6
Nair M, Nagamori I, Sun P, Mishra DP, Rheaume C, Li B, Sassone-Corsi P, Dai X (2008) Nuclear regulator Pygo2 controls spermiogenesis and histone H3 acetylation. Dev Biol 320(2):446–455. doi:10.1016/j.ydbio.2008.05.553
Natsume-Kitatani Y, Shiga M, Mamitsuka H (2011) Genome-wide integration on transcription factors, histone acetylation and gene expression reveals genes co-regulated by histone modification patterns. PLoS ONE 6(7):e22281. doi:10.1371/journal.pone.0022281
Neal KC, Pannuti A, Smith ER, Lucchesi JC (2000) A new human member of the MYST family of histone acetyl transferases with high sequence similarity to Drosophila MOF. Biochim Biophys Acta 1490(1–2):170–174. doi:10.1016/S0167-4781(99)00211-0
Neuwald AF, Landsman D (1997) GCN5-related histone N-acetyltransferases belong to a diverse superfamily that includes the yeast SPT10 protein. Trends Biochem Sci 22(5):154–155. doi:10.1016/S0968-0004(97)01034-7
Penttila TL, Yuan L, Mali P, Hoog C, Parvinen M (1995) Haploid gene expression: temporal onset and storage patterns of 13 novel transcripts during rat and mouse spermiogenesis. Biol Reprod 53(3):499–510. doi:10.1095/biolreprod53.3.499
Petri S, Kiaei M, Kipiani K, Chen J, Calingasan NY, Crow JP, Beal MF (2006) Additive neuroprotective effects of a histone deacetylase inhibitor and a catalytic antioxidant in a transgenic mouse model of amyotrophic lateral sclerosis. Neurobiol Dis 22(1):40–49. doi:10.1016/j.nbd.2005.09.013
Roosen-Runge EC (1962) The process of spermatogenesis in mammals. Biol Rev Camb Philos Soc 37:343–377. doi:10.1111/j.1469-185X.1962.tb01616.x
Roy A, Ghosh A, Jana A, Liu X, Brahmachari S, Gendelman HE, Pahan K (2012) Sodium phenylbutyrate controls neuroinflammatory and antioxidant activities and protects dopaminergic neurons in mouse models of Parkinson’s disease. PLoS ONE 7(6):e38113. doi:10.1371/journal.pone.0038113
Ruefli AA, Ausserlechner MJ, Bernhard D, Sutton VR, Tainton KM, Kofler R, Smyth MJ, Johnstone RW (2001) The histone deacetylase inhibitor and chemotherapeutic agent suberoylanilide hydroxamic acid (SAHA) induces a cell-death pathway characterized by cleavage of Bid and production of reactive oxygen species. Proc Natl Acad Sci USA 98(19):10833–10838. doi:10.1073/pnas.191208598
Saito M, Kumamoto K, Robles AI, Horikawa I, Furusato B, Okamura S, Goto A, Yamashita T, Nagashima M, Lee TL, Baxendale VJ, Rennert OM, Takenoshita S, Yokota J, Sesterhenn IA, Trivers GE, Hussain SP, Harris CC (2010) Targeted disruption of Ing2 results in defective spermatogenesis and development of soft-tissue sarcomas. PLoS ONE 5(11):e15541. doi:10.1371/journal.pone.0015541
Shah P, Gau Y, Sabnis G (2014) Histone deacetylase inhibitor entinostat reverses epithelial to mesenchymal transition of breast cancer cells by reversing the repression of E-cadherin. Breast Cancer Res Treat 143(1):99–111. doi:10.1007/s10549-013-2784-7
Soderstrom KO, Parvinen M (1976) RNA synthesis in different stages of rat seminiferous epithelial cycle. Mol Cell Endocrinol 5(3–4):181–199. doi:10.1016/0303-7207(76)90082-4
Song N, Liu J, An S, Nishino T, Hishikawa Y, Koji T (2011) Immunohistochemical analysis of histone H3 modifications in germ cells during mouse spermatogenesis. Acta Histochem Cytochem 44(4):183–190. doi:10.1267/ahc.11027
Spencer TE, Jenster G, Burcin MM, Allis CD, Zhou J, Mizzen CA, McKenna NJ, Onate SA, Tsai SY, Tsai MJ, O’Malley BW (1997) Steroid receptor coactivator-1 is a histone acetyltransferase. Nature 389(6647):194–198. doi:10.1038/38304
Suka N, Suka Y, Carmen AA, Wu J, Grunstein M (2001) Highly specific antibodies determine histone acetylation site usage in yeast heterochromatin and euchromatin. Mol Cell 8(2):473–479. doi:10.1016/S1097-2765(01)00301-X
Uhlen M, Oksvold P, Fagerberg L, Lundberg E, Jonasson K, Forsberg M, Zwahlen M, Kampf C, Wester K, Hober S, Wernerus H, Björling L, Ponten F (2010) Towards a knowledge-based Human Protein Atlas. Nat Biotechnol 28(12):1248–1250. doi:10.1038/nbt1210-1248
Ungerstedt JS, Sowa Y, Xu WS, Shao Y, Dokmanovic M, Perez G, Ngo L, Holmgren A, Jiang X, Marks PA (2005) Role of thioredoxin in the response of normal and transformed cells to histone deacetylase inhibitors. Proc Natl Acad Sci USA 102(3):673–678. doi:10.1073/pnas.0408732102
Vempati RK, Jayani RS, Notani D, Sengupta A, Galande S, Haldar D (2010) p300-mediated acetylation of histone H3 lysine 56 functions in DNA damage response in mammals. J Biol Chem 285(37):28553–28564. doi:10.1074/jbc.M110.149393
Wang RA, Nakane PK, Koji T (1998) Autonomous cell death of mouse male germ cells during fetal and postnatal period. Biol Reprod 58(5):1250–1256. doi:10.1095/biolreprod58.5.1250
Wittschieben BO, Otero G, de Bizemont T, Fellows J, Erdjument-Bromage H, Ohba R, Li Y, Allis CD, Tempst P, Svejstrup JQ (1999) A novel histone acetyltransferase is an integral subunit of elongating RNA polymerase II holoenzyme. Mol Cell 4(1):123–128. doi:10.1016/S1097-2765(00)80194-X
Yan W, Si Y, Slaymaker S, Li J, Zheng H, Young DL, Aslanian A, Saunders L, Verdin E, Charo IF (2010) Zmynd15 encodes a histone deacetylase-dependent transcriptional repressor essential for spermiogenesis and male fertility. J Biol Chem 285(41):31418–31426. doi:10.1074/jbc.M110.116418
Yomogida K, Yagura Y, Nishimune Y (2002) Electroporated transgene-rescued spermatogenesis in infertile mutant mice with a sertoli cell defect. Biol Reprod 67(3):712–717. doi:10.1095/biolreprod.101.001743
Yu KH, Weng LJ, Fu S, Piantadosi S, Gore SD (1999) Augmentation of phenylbutyrate-induced differentiation of myeloid leukemia cells using all-trans retinoic acid. Leukemia 13(8):1258–1265. doi:10.1038/sj.leu.2401468
Zhang Y, Kwon S, Yamaguchi T, Cubizolles F, Rousseaux S, Kneissel M, Cao C, Li N, Cheng HL, Chua K, Lombard D, Mizeracki A, Matthias G, Alt FW, Khochbin S, Matthias P (2008) Mice lacking histone deacetylase 6 have hyperacetylated tubulin but are viable and develop normally. Mol Cell Biol 28(5):1688–1701. doi:10.1128/MCB.01154-06
Zhang Z, Liu J, Kaur M, Krantz ID (2012) Characterization of DNA methylation and its association with other biological systems in lymphoblastoid cell lines. Genomics 99(4):209–219. doi:10.1016/j.ygeno.2012.01.002
Zheng J, Xia X, Ding H, Yan A, Hu S, Gong X, Zong S, Zhang Y, Sheng HZ (2008) Erasure of the paternal transcription program during spermiogenesis: the first step in the reprogramming of sperm chromatin for zygotic development. Dev Dyn 237(5):1463–1476. doi:10.1002/dvdy.21499
Zhou Q, Agoston AT, Atadja P, Nelson WG, Davidson NE (2008) Inhibition of histone deacetylases promotes ubiquitin-dependent proteasomal degradation of DNA methyltransferase 1 in human breast cancer cells. Mol Cancer Res 6(5):873–883. doi:10.1158/1541-7786.MCR-07-0330
Acknowledgments
This study was supported in part by a Grant-in-Aid for Scientific Research from the Japan Society for the Promotion of Science (No. 18390060 to Koji, T.).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.

418_2014_1283_MOESM1_ESM.jpg
Fig. S1 TUNEL staining and immunohistochemical detection of 8-OHdG in adult NaPB (a-d)-treated mouse testes at 6 h after treatment and SAHA (e-h)-treated mouse testes at 12 h after treatment. Serial sections were used for H&E staining (a, e), TUNEL staining (b, f), immunohistochemical staining for 8-OHdG (c, g). Arrowheads TUNEL-positive cells without 8-OHdG signals; arrows TUNEL-positive cells with 8-OHdG signals. The insets (b′) in b and (c′) in c are enlarged. As a negative control, sections of NaPB (d)- and SAHA (h)-treated testes were reacted with normal mouse IgG instead of the specific antibody. Scale bars 50 μm. (JPEG 1463 kb)

418_2014_1283_MOESM2_ESM.jpg
Fig. S2 Immunohistochemical detection of histone H3K9ac, H3K18ac and H3K23ac in paraffin-embedded sections of adult DMSO (a-e)- and SAHA (f-j)-treated mouse testes at 12 h after treatment. Serial sections were used for H&E staining (a, f), immunohistochemical staining for H3K9ac (b, g), H3K18ac (c, h), and H3K23ac (d, i). As a negative control, DMSO (e)- and SAHA (j)-treated sections of testes were reacted with normal rabbit IgG instead of specific antibodies. Seminiferous tubules at stage VI-VII are shown. Scale bars 50 μm. (JPEG 2003 kb)

418_2014_1283_MOESM3_ESM.jpg
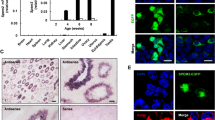
Fig. S3 Immunofluorescence detection of GFP in LacZ (a-c)- , SRC-1 (d-f)-shRNA transfected and non-transfected (g-i) 28-day-old neonatal mouse testes. (a, d, g) immunofluorescence staining of GFP. (b, e, h) As a negative control, sections were reacted with normal mouse IgG instead of the specific antibody. (c, f, i) DAPI staining. Arrowheads GFP-positive cells; asterisks nonspecific fluorescence signals from the secondary antibody in interstitial Leydig cells. Scale bars 50 μm. (JPEG 2039 kb)

418_2014_1283_MOESM4_ESM.jpg
Fig. S4 Double immunofluorescence staining of GFP and GCN5 in LacZ- and GCN5-shRNA plasmid transfected 28-day-old neonatal mouse testes at day 10 post-transfection. (a-f) Double immunofluorescence staining of GFP (a, d) and GCN5 (b, e) in LacZ (a-c)- and GCN5 (d-f)-shRNA transfected testes. Arrows GFP-positive round spermatids in LacZ-shRNA transfected testes; arrowheads GFP-positive round spermatids in GCN5-shRNA transfected testes. Scale bars 10 μm (JPEG 2089 kb)

418_2014_1283_MOESM5_ESM.jpg
Fig. S5 Double immunofluorescence staining of GFP and ELP3 in LacZ- and ELP3-shRNA plasmid transfected 28-day-old neonatal mouse testes at day 10 post-transfection. (a-f) Double immunofluorescence staining of GFP (a, d) and ELP3 (b, e) in LacZ (a-c)- and ELP3 (d-f)-shRNA transfected testes. Arrows GFP-positive round spermatids in LacZ-shRNA transfected testes; arrowheads GFP-positive round spermatids in ELP3-shRNA transfected testes. Scale bars 10 μm. (JPEG 1935 kb)

418_2014_1283_MOESM6_ESM.jpg
Fig. S6 Double immunofluorescence staining of GFP and H3K9ac; GFP and H3K18ac; GFP and H3K23ac in shRNA plasmid transfected 28-day-old neonatal mouse testes at day 10 post-transfection. (a-d) Double immunofluorescence staining of GFP and H3K9ac in LacZ (a)- , SRC-1 (b)-, GCN5 (c)- and ELP3 (d)-shRNA transfected testes. (e-h) Double immunofluorescence staining of GFP and H3K18ac in LacZ (e)- , SRC-1 (f)- , GCN5 (g)- and ELP3 (h)-shRNA transfected testes. (i-l) Double immunofluorescence staining of GFP and H3K23ac in LacZ (i)- , SRC-1 (j)- , GCN5 (k)- and ELP3 (l)-shRNA transfected testes. Arrowheads GFP-positive round spermatid in SRC-1-shRNA transfected testes with reduced H3K9ac staining. Scale bar 20 μm. (JPEG 2750 kb)

418_2014_1283_MOESM7_ESM.jpg
Fig. S7 Methylation level of CCGG sites in adjacent section of NaPB-treated testis at 6 h after treatment by HELMET method using DAB staining. Adjacent sections of paraffin-embedded mouse testes were used for H&E staining, TUNEL, non-methylated CCGG sites and methylated CCGG sites. (a) H&E staining. (b) TUNEL staining. (c) Blockade of 3′-OH ends with dideoxynucleotides by TdT. After blockade, the section was labeled with biotin-16-dUTP by TdT and the incorporated biotin was detected with HRP-anti-biotin. No signals were observed. (d) Staining for non-methylated CCGG sites. After the blockade procedure described in (c), the section was digested with HpaII, labeled with biotin-16-dUTP and visualized by enzyme immunohistochemistry with HRP-anti-biotin. (e) Blockade of HpaII cutting sites with dideoxynucleotides by TdT. The section was digested with HpaII, and the cutting sites were blocked with a dideoxynucleotide mixture. Then, the section was processed in a manner similar to that described in (c). (f) Staining for methylated CCGG sites. After blockade of HpaII cutting sites with dideoxynucleotides, the section was digested with MspI and the cutting sites were labeled with biotin-16-dUTP and visualized with HRP-anti biotin. Arrows TUNEL-positive round spermatids with elevated non-methylated CCGG sites; arrowheads TUNEL-negative round spermatids with elevated non-methylated CCGG sites. Scale bars 50 μm. (JPEG 1996 kb)
Rights and permissions
About this article
Cite this article
Dai, L., Endo, D., Akiyama, N. et al. Aberrant levels of histone H3 acetylation induce spermatid anomaly in mouse testis. Histochem Cell Biol 143, 209–224 (2015). https://doi.org/10.1007/s00418-014-1283-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00418-014-1283-1