Abstract
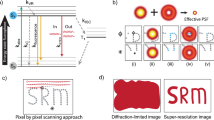
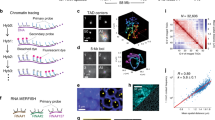
Spatially modulated illumination (SMI) microscopy is a method of widefield fluorescence microscopy featuring interferometric illumination, which delivers structural information about nanoscale features in fluorescently labeled cells. Using this approach, structural changes in the context of gene activation and chromatin remodeling may be revealed. In this paper we present the application of SMI microscopy to size measurements of the 7q22 gene region, giving us a size estimate of 105±16 nm which corresponds to an average compaction ratio of 1:324. The results for the 7q22 domain are compared with the previously measured sizes of other fluorescently labeled gene regions, and to those obtained for transcription factories. The absence of a correlation between the measured and genomic sizes of the various gene regions indicate that a high variability in chromatin folding is present, with factors other than the sequence length contributing to the chromatin compaction. Measurements of the 7q22 region in different preparations and at different excitation wavelengths show a good agreement, thus demonstrating that the technique is robust when applied to biological samples.



Similar content being viewed by others
References
Albrecht B, Failla A, Schweitzer A, Cremer C (2002) Spatially modulated illumination microscopy allows axial distance resolution in the nanometer range. Appl Opt 41:80–87
Bailey B, Farkas DL, Taylor DL, Lanni F (1993) Enhancement of axial resolution in fluorescence microscopy by standing-wave excitation. Nature 366:44–48
Bartova E, Kozubek S, Jirsova P, Kozubek M, Gajova H, Lukasova E, Skalnikova M, Ganova A, Koutna I, Hausmann M (2002) Nuclear topography and gene activity in human differentiated cells. J Struct Biol 139:76–89
Bornfleth H, Sätzler K, Eils R, Cremer C (1998) High-precision distance measurements and volume-conserving segmentation of objects near and below the resolution limit in three-dimensional confocal fluorescence microscopy. J Microsc 189:118–136
Chambeyron S, Bickmore W (2004) Chromatin decondensation and nuclear reorganization of the HoxB locus upon induction of transciption. Genes Dev 18:1119–1130
Cremer T, Cremer C (2001) Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat Rev Genet 2:292–301
Cremer T, Kreth G, Koester H, Fink RHA, Heintzmann R, Solovei I, Zink D, Cremer C (2000) Chromosome territories, interchromatin domain compartment and nuclear matrix: an integrated view of the functional nuclear architecture. Crit Rev Euk Gene Expr 12:179–212
Cremer M, von Hase J, Volm T, Brero A, Kreth G, Walter J, Fischer C, Solovei I, Cremer C, Cremer T (2001) Non-random radial higher-order chromatin arrangements in nuclei of diploid human cells. Chromosome Res 9:541–567
van Driel R, Fransz PF, Verschure PJ (2003) The eukaryotic genome: a system regulated at different hierarchical levels. J Cell Sci 116:4067–4075
Dyba M, Hell SW (2002) Focal spots of size λ/23 open up far-field fluorescence microscopy at 33 nm axial resolution. Phys Rev Lett 88:163901
Ebert BL, Bunn HF (1999) Regulation of the erythropoietin gene. Blood 94:1864–1877
Egner A, Jakobs S, Hell SW (2002) Fast 100-nm resolution three-dimensional microscope reveals structural plasticity of mitochondria in live yeast. PNAS 99:3370–3375
Failla AV, Cavallo A, Cremer C (2002a) Subwavelength size determination using spatially modulated illumination virtual microscopy. Appl Opt 41:6651–6659
Failla AV, Spöri U, Albrecht B, Kroll A, Cremer C (2002b) Nanosizing of fluorescent objects by spatially modulated illumination microscopy. Appl Opt 41:7275–7283
Failla AV, Albrecht B, Spöri U, Schweitzer A, Kroll A, Hildenbrand G, Bach M, Cremer C (2003) Nanostructure analysis using spatially modulated illumination microscopy. ComPlexUs 1:77–88
Fandrey J (2004) Oxygen-dependent and tissue-specific regulation of erythropoietin gene expression. Am J Physiol Regul Integr Comp Physiol 286:R977–R988
Frohn JT, Knapp HF, Stemmer A (2000) True optical resolution beyond the Rayleigh limit achieved by standing wave illumination. PNAS 97:7232–7236
Frohn JT, Knapp HF, Stemmer A (2001) Three-dimensional resolution enhancement in fluorescence microscopy by harmonic excitation. Opt Lett 26:828–830
Griffiths G, Burke B, Lucocq J (1993) Fine structure immunocytochemistry. Springer, Berlin Heidelberg New York
Gustafsson MGL, Agard DA, Sedat JW (1995) Sevenfold improvement of axial resolution in 3D widefield microscopy using two objective lenses. Proc SPIE 2412:147–156
Gustafsson MGL, Agard DA, Sedat JW (1996) 3D widefield microscopy with two objective lenses: experimental verification of improved axial resolution. Proc SPIE 2655:62–65
Hausmann M, Winkler R, Hildenbrand G, Finsterle J, Weisel A, Rapp A, Schmitt E, Janz S, Cremer C (2003) COMBO-FISH: specific labeling of nondenatured chromatin targets by computer-selected DNA oligonucleotide probe combinations. Biotechniques 35:564–577
Heintzmann R, Jovin TM, Cremer C (2002) Saturated patterned excitation microscopy—a concept for optical resolution improvement. J Opt Soc Am A 19:1599–1609
Hell SW, Lindek S, Cremer C, Stelzer EHK (1994) Measurement of 4pi-confocal point spread function proves 75 nm axial resolution. Appl Phys Lett 64:1335–1337
Hildenbrand G, Rapp A, Spoeri U, Wagner C, Cremer C, Hausmann M (2005) Nano-sizing of specific gene domains in intact human cell nuclei by spatially modulated illumination light microscopy. Biophys J 88:4312–4318
Jackson DA, Iborra FJ, Manders EMM, Cook PR (1998) Numbers and organization of RNA polymerases, nascent transcripts, and transcription units in HeLa nuclei. Mol Biol Cell 9:1523–1536
Klar TA, Jakobs S, Dyba M, Egner A, Hell SW (2000) Fluorescence microscopy with diffraction resolution barrier broken by stimulated emission. Proc Natl Acad Sci USA 97:8206–8210
Kozubek S, Lukasova E, Jirsova P, Koutna I, Kozubek M, Ganova A, Bartova E, Falk M, Pasekova R (2002) 3D structure of the human genome: order in randomness. Chromosoma 111:321–331
Martin S, Pombo A (2003) Transcription factories: quantitative studies of nanostructures in the mammalian nucleus. Chrom Res 11:461–470
Martin S, Failla AV, Spöri U, Cremer C, Pombo A (2004) Measuring the size of biological nanostructures with spatially modulated illumination microscopy. Mol Biol Cell 15:2449–2455
Miller OLJ, Bakken AH (1972) Morphological studies of transcription. Acta Endocrinol Suppl Copenh 168:155–177
O’Brien TP, Bult CJ, Cremer C, Grunze M, Knowles BB, Langowski J, McNally J, Pederson T, Politz JC, Pombo A, Schmahl G, Spatz JP, van Driel R (2003) Genome function and nuclear architecture: from gene expression to nanoscience. Genome Res 13:1029–1041
Odenheimer J, Kreth G, Heermann DW (2005) Dynamic simulation of active/inactive chromatin domains. J Biol Phys 31(4):305–310
Pombo A, Hollinshead M, Cook PR (1999a) Bridging the resolution gap: imaging the same transcription factories in cryosections by light and electron microscopy. J Histochem Cytochem 47:471–480
Pombo A, Jackson DA, Hollinshead M, Wang Z, Roeder RG, Cook PR (1999b) Regional specialization in the nucleus: visualization of discrete sites of transcription by RNA polymerase III. EMBO J 18:2241–2253
Schweitzer A, Wagner C, Cremer C (2004) The nanosizing of fluorescent objects by 458 nm spatially modulated illumination microscopy using a simplified size evaluation algorithm. J Phys Condens Matter 16:S2393–S2404
Solovei I, Cavallo A, Schermelleh L, Jaunin F, Scasselati C, Cmarko D, Cremer C, Fakan S, Cremer T (2002) Spatial preservation of nuclear chromatin architecture during three-dimensional fluorescence in situ hybridization (3D-FISH). Exp Cell Res 276:10–23
Spector DL (2003) The dynamics of chromosome organization and gene regulation. Ann Rev Biochem 72:573–608
Spöri U, Failla AV, Cremer C (2004) Superresolution size determination in fluorescence microscopy: a comparison between spatially modulated illumination and confocal laser scanning microscopy. J Appl Phys 95:8436–8443
Tanabe H, Müller S, Neusser M, von Hase J, Calcagno E, Cremer M, Solovei I, Cremer C, Cremer T (2002) Evolutionary conservation of chromosome territory arrangements in cell nuclei from higher primates. PNAS 99:4424–4429
Wagner C, Spöri U, Cremer C (2005) High-precision SMI microscopy size measurements by simultaneous frequency domain reconstruction of the axial point spread function. Optik 116:15–21
Wansink D, Sibon O, Cremers F, van Driel R, de Jong L (1996) Ultrastructural localization of active genes in nuclei of A431 cells. J Cell Biochem 62:10–18
Acknowledgements
We thank Dr. Ana Pombo and Dr. Sonya Martin for providing the HeLa cell cryosections, the measurements, and continuing support. We thank Prof. Michael Hausmann for scientific assistance and Dr. Lars Hildenbrand for stimulating discussions. We gratefully acknowledge financial support within the DFG priority program 1128 ‘Optical analysis of the structure and dynamics of supra-molecular biological complexes’, and by the European Commission, projects LSHG-CT-2003-503259 and LSHG-CT-2003-503441.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mathée, H., Baddeley, D., Wotzlaw, C. et al. Nanostructure of specific chromatin regions and nuclear complexes. Histochem Cell Biol 125, 75–82 (2006). https://doi.org/10.1007/s00418-005-0096-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00418-005-0096-7