Abstract
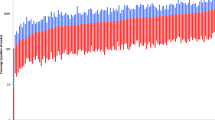
Within forensic genetics, there is still a need for supplementary DNA marker typing in order to increase the power to solve cases for both identity testing and complex kinship issues. One major disadvantage with current capillary electrophoresis (CE) methods is the limitation in DNA marker multiplex capability. By utilizing massive parallel sequencing (MPS) technology, this capability can, however, be increased. We have designed a customized GeneRead DNASeq SNP panel (Qiagen) of 140 previously published autosomal forensically relevant identity SNPs for analysis using MPS. One single amplification step was followed by library preparation using the GeneRead Library Prep workflow (Qiagen). The sequencing was performed on a MiSeq System (Illumina), and the bioinformatic analyses were done using the software Biomedical Genomics Workbench (CLC Bio, Qiagen). Forty-nine individuals from a Swedish population were genotyped in order to establish genotype frequencies and to evaluate the performance of the assay. The analyses showed to have a balanced coverage among the included loci, and the heterozygous balance showed to have less than 0.5 % outliers. Analyses of dilution series of the 2800M Control DNA gave reproducible results down to 0.2 ng DNA input. In addition, typing of FTA samples and bone samples was performed with promising results. Further studies and optimizations are, however, required for a more detailed evaluation of the performance of degraded and PCR-inhibited forensic samples. In summary, the assay offers a straightforward sample-to-genotype workflow and could be useful to gain information in forensic casework, for both identity testing and in order to solve complex kinship issues.




Similar content being viewed by others
Notes
For these estimate, haplotype frequencies were used for the loci in LD instead of allele frequencies
References
Butler JM (2006) Genetics and genomics of core short tandem repeat loci used in human identity testing. J Forensic Sci 51(2):253–265. doi:10.1111/j.1556-4029.2006.00046.x
Butler JM, Hill CR (2012) Biology and genetics of new autosomal STR loci useful for forensic DNA analysis. Forensic Sci Rev 24(1):15–26
Parsons TJ, Huel R, Davoren J, Katzmarzyk C, Milos A, Selmanovic A, Smajlovic L, Coble MD, Rizvic A (2007) Application of novel “mini-amplicon” STR multiplexes to high volume casework on degraded skeletal remains. Forensic Sci Int Genet 1(2):175–179. doi:10.1016/j.fsigen.2007.02.003
Phillips C, Gelabert-Besada M, Fernandez-Formoso L, Garcia-Magarinos M, Santos C, Fondevila M, Ballard D, Syndercombe Court D, Carracedo A, Lareu MV (2014) “New turns from old STaRs”: enhancing the capabilities of forensic short tandem repeat analysis. Electrophoresis 35(21–22):3173–3187. doi:10.1002/elps.201400095
Ge J, Budowle B, Chakraborty R (2011) Choosing relatives for DNA identification of missing persons. J Forensic Sci 56(Suppl 1):S23–28. doi:10.1111/j.1556-4029.2010.01631.x
Nothnagel M, Schmidtke J, Krawczak M (2010) Potentials and limits of pairwise kinship analysis using autosomal short tandem repeat loci. Int J Leg Med 124(3):205–215. doi:10.1007/s00414-009-0413-0
Phillips C, Fondevila M, Garcia-Magarinos M, Rodriguez A, Salas A, Carracedo A, Lareu MV (2008) Resolving relationship tests that show ambiguous STR results using autosomal SNPs as supplementary markers. Forensic Sci Int Genet 2(3):198–204. doi:10.1016/j.fsigen.2008.02.002
Børsting C, Morling N (2015) Next generation sequencing and its applications in forensic genetics. Forensic Sci Int Genet 18:78–89. doi:10.1016/j.fsigen.2015.02.002
Daniel R, Santos C, Phillips C, Fondevila M, van Oorschot RA, Carracedo A, Lareu MV, McNevin D (2015) A SNaPshot of next generation sequencing for forensic SNP analysis. Forensic Sci Int Genet 14:50–60. doi:10.1016/j.fsigen.2014.08.013
Børsting C, Morling N (2011) Mutations and/or close relatives? Six case work examples where 49 autosomal SNPs were used as supplementary markers. Forensic Sci Int Genet 5(3):236–241. doi:10.1016/j.fsigen.2010.02.007
Pakstis AJ, Speed WC, Fang R, Hyland FC, Furtado MR, Kidd JR, Kidd KK (2010) SNPs for a universal individual identification panel. Hum Genet 127(3):315–324. doi:10.1007/s00439-009-0771-1
Tillmar AO, Mostad P (2014) Choosing supplementary markers in forensic casework. Forensic Sci Int Genet 13:128–133. doi:10.1016/j.fsigen.2014.06.019
Børsting C, Fordyce SL, Olofsson J, Mogensen HS, Morling N (2014) Evaluation of the Ion Torrent™ HID SNP 169-plex: a SNP typing assay developed for human identification by second generation sequencing. Forensic Sci Int Genet 12:144–154. doi:10.1016/j.fsigen.2014.06.004
Churchill JD, Schmedes SE, King JL, Budowle B (2015) Evaluation of the Illumina(®) beta version ForenSeq™ DNA signature prep kit for use in genetic profiling. Forensic Sci Int Genet 20:20–29. doi:10.1016/j.fsigen.2015.09.009
Warshauer DH, Davis CP, Holt C, Han Y, Walichiewicz P, Richardson T, Stephens K, Jager A, King J, Budowle B (2015) Massively parallel sequencing of forensically relevant single nucleotide polymorphisms using TruSeq™ forensic amplicon. Int J Legal Med 129(1):31–36. doi:10.1007/s00414-014-1108-8
Sanchez JJ, Phillips C, Børsting C, Balogh K, Bogus M, Fondevila M, Harrison CD, Musgrave-Brown E, Salas A, Syndercombe-Court D, Schneider PM, Carracedo A, Morling N (2006) A multiplex assay with 52 single nucleotide polymorphisms for human identification. Electrophoresis 27(9):1713–1724. doi:10.1002/elps.200500671
Lindblom B, Holmlund G (1988) Rapid DNA purification for restriction fragment length polymorphism analysis. Gene Anal Tech 5(5):97–101
Holmlund G, Lodestad I, Nilsson H, Lindblom B (2006) Experiences from DNA analysis in Sweden for the identification of tsunami victims. Int Congr Ser 1288:744–746
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10(3):564–567. doi:10.1111/j.1755-0998.2010.02847.x
Holm S (1979) A simple sequentially rejective multiple test procedure. Scand J Stat 6:65–70
Chakraborty R, Stivers DN (1996) Paternity exclusion by DNA markers: effects of paternal mutations. J Forensic Sci 41(4):671–677
Jones DA (1972) Blood samples: probability of discrimination. J Forensic Sci Soc 12(2):355–359
Arezi B, Xing W, Sorge JA, Hogrefe HH (2003) Amplification efficiency of thermostable DNA polymerases. Anal Biochem 321(2):226–235
Eduardoff M, Santos C, de la Puente M, Gross TE, Fondevila M, Strobl C, Sobrino B, Ballard D, Schneider PM, Carracedo Á, Lareu MV, Parson W, Phillips C (2015) Inter-laboratory evaluation of SNP-based forensic identification by massively parallel sequencing using the Ion PGM™. Forensic Sci Int Genet 17:110–121. doi:10.1016/j.fsigen.2015.04.007
1000 Genomes Project Consortium, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, Korbel JO, Marchini JL, McCarthy S, McVean GA, Abecasis GR (2015) A global reference for human genetic variation. Nature 526(7571):68–74. doi:10.1038/nature15393
Buckleton J, Triggs C (2006) The effect of linkage on the calculation of DNA match probabilities for siblings and half siblings. Forensic Sci Int 160(2–3):193–199. doi:10.1016/j.forsciint.2005.10.004
Gill P, Phillips C, McGovern C, Bright JA, Buckleton J (2012) An evaluation of potential allelic association between the STRs vWA and D12S391: implications in criminal casework and applications to short pedigrees. Forensic Sci Int Genet 6(4):477–486. doi:10.1016/j.fsigen.2011.11.001
O’Connor KL, Tillmar AO (2012) Effect of linkage between vWA and D12S391 in kinship analysis. Forensic Sci Int Genet 6(6):840–844. doi:10.1016/j.fsigen.2012.03.008
Kling D, Egeland T, Tillmar AO (2012) FamLink—a user friendly software for linkage calculations in family genetics. Forensic Sci Int Genet 6(5):616–620. doi:10.1016/j.fsigen.2012.01.012
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 371 kb)
Rights and permissions
About this article
Cite this article
Grandell, I., Samara, R. & Tillmar, A.O. A SNP panel for identity and kinship testing using massive parallel sequencing. Int J Legal Med 130, 905–914 (2016). https://doi.org/10.1007/s00414-016-1341-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-016-1341-4