Abstract
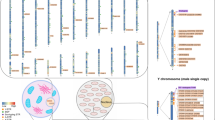
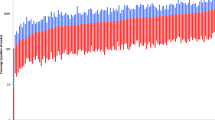
The MiSeq® FGX Forensic system and the HID-Ion AmpliSeq Panel were previously developed for massively parallel sequencing (MPS) for forensic casework. Among the three major sequencing platforms, BGISEQ-500TM, which is based on multiple PCRs, is still lacking in forensics. Here, a novel forensic panel was constructed to detect 186 single-nucleotide polymorphisms (SNPs) and 123 short tandem repeats (STRs) with MPS technology on the BGISEQ-500™ platform. First, the library preparation, sequencing process, and data analysis were performed, focusing on the average depth of coverage and heterozygote balance. We calculated the allelic frequencies and forensic parameters of STR and SNP loci in 73 unrelated Chinese Han individuals. In addition, performance was evaluated with accuracy, uniformity, sensitivity, PCR inhibitor, repeatability and reproducibility, mixtures, degraded samples, case-type samples, and pedigree analyses. The results showed that 100% accurate and concordant genotypes can be obtained, and the loci with an abundance in the interquartile range accounted for 92.90% of the total, suggesting reliable uniformity in this panel. We obtained a locus detection rate that was higher than 98.78% from 78 pg of input DNA, and the optimal amount was 1.25–10 ng. The maximum concentrations of hematin and humic acid were 200 and 100 μM, respectively (the ratios of detected loci were 96.52% and 92.41%), in this panel. As a mixture, compared with those of SNPs, minor-contributor alleles of STRs could be detected at higher levels. For the degraded sample, the ratio of detected loci was 98.41%, and most profiles from case-type samples were not significantly different in abundance in our studies. As a whole, this panel showed high-performance, reliable, robust, repeatable, and reproducible results, which are sufficient for paternity testing, individual identification, and use for potentially degraded samples in forensic science.

Similar content being viewed by others
References
Budowle B, van Daal A (2008) Forensically relevant SNP classes. Biotechniques 44:603–610. https://doi.org/10.2144/000112806
Frégeau CJ, Fourney RM (1993) DNA typing with fluorescently tagged short tandem repeats: a sensitive and accurate approach to human identification. Biotechniques 15:100–119
Hammond HA, Jin L, Zhong Y, Caskey CT, Chakraborty R (1994) Evaluation of 13 short tandem repeat loci for use in personal identification applications. Am J Hum Genet 55:175–189
Jäger AC, Alvarez ML, Davis CP, Guzmán E, Han Y, Way L, Walichiewicz P, Silva D, Pham N, Caves G, Bruand J, Schlesinger F, Pond SJK, Varlaro J, Stephens KM, Holt CL (2017) Developmental validation of the MiSeq FGx forensic genomics system for targeted next generation sequencing in forensic DNA casework and database laboratories. Forensic Sci Int Genet 28:52–70. https://doi.org/10.1016/j.fsigen.2017.01.011
Edwards MC, Gibbs RA (1994) Multiplex PCR: advantages, development, and applications. PCR Methods Appl 3:S65–S75. https://doi.org/10.1101/gr.3.4.s65
Seo SB, King JL, Warshauer DH, Davis CP, Ge J, Budowle B (2013) Single nucleotide polymorphism typing with massively parallel sequencing for human identification. Int J Legal Med 127:1079–1086. https://doi.org/10.1007/s00414-013-0879-7
Bruijns B, Tiggelaar R, Gardeniers H (2018) Massively parallel sequencing techniques for forensics: A review. Electrophoresis 39:2642–2654. https://doi.org/10.1002/elps.201800082
Børsting C, Morling N (2015) Next generation sequencing and its applications in forensic genetics. Forensic Sci Int Genet 18:78–89. https://doi.org/10.1016/j.fsigen.2015.02.002
Daniel R, Santos C, Phillips C, Fondevila M, van Oorschot RAH, Carracedo Á, Lareu MV, McNevin D (2015) A SNaPshot of next generation sequencing for forensic SNP analysis. Forensic Sci Int Genet 14:50–60. https://doi.org/10.1016/j.fsigen.2014.08.013
Pakstis AJ, Speed WC, Fang R, Hyland FCL, Furtado MR, Kidd JR, Kidd KK (2010) SNPs for a universal individual identification panel. Hum Genet 127:315–324. https://doi.org/10.1007/s00439-009-0771-1
Guo F, Zhou Y, Song H, Zhao J, Shen H, Zhao B, Liu F, Jiang X (2016) Next generation sequencing of SNPs using the HID-Ion AmpliSeq™ Identity Panel on the Ion Torrent PGM™ platform. Forensic Sci Int Genet 25:73–84. https://doi.org/10.1016/j.fsigen.2016.07.021
Avila E, Felkl AB, Graebin P, Nunes CP, Alho CS (2019) Forensic characterization of Brazilian regional populations through massive parallel sequencing of 124 SNPs included in HID ion Ampliseq Identity Panel. Forensic Sci Int Genet 40:74–84. https://doi.org/10.1016/j.fsigen.2019.02.012
Grandell I, Samara R, Tillmar AO (2016) A SNP panel for identity and kinship testing using massive parallel sequencing. Int J Legal Med 130:905–914. https://doi.org/10.1007/s00414-016-1341-4
de la Puente M, Phillips C, Santos C, Fondevila M, Carracedo Á, Lareu MV (2017) Evaluation of the Qiagen 140-SNP forensic identification multiplex for massively parallel sequencing. Forensic Sci Int Genet 28:35–43. https://doi.org/10.1016/j.fsigen.2017.01.012
Phillips C, Parson W, Lundsberg B, Santos C, Freire-Aradas A, Torres M, Eduardoff M, Børsting C, Johansen P, Fondevila M, Morling N, Schneider P, Carracedo Á, Lareu MV (2014) Building a forensic ancestry panel from the ground up: The EUROFORGEN Global AIM-SNP set. Forensic Sci Int Genet 11:13–25. https://doi.org/10.1016/j.fsigen.2014.02.012
Al-Asfi M, McNevin D, Mehta B, Power D, Gahan ME, Daniel R (2018) Assessment of the Precision ID Ancestry panel. Int J Legal Med 132:1581–1594. https://doi.org/10.1007/s00414-018-1785-9
Børsting C, Fordyce SL, Olofsson J, Mogensen HS, Morling N (2014) Evaluation of the Ion Torrent™ HID SNP 169-plex: A SNP typing assay developed for human identification by second generation sequencing. Forensic Sci Int Genet 12:144–154. https://doi.org/10.1016/j.fsigen.2014.06.004
Sanchez JJ, Phillips C, Børsting C, Balogh K, Bogus M, Fondevila M, Harrison CD, Musgrave-Brown E, Salas A, Syndercombe-Court D, Schneider PM, Carracedo A, Morling N (2006) A multiplex assay with 52 single nucleotide polymorphisms for human identification. Electrophoresis 27:1713–1724. https://doi.org/10.1002/elps.200500671
Fordyce SL, Mogensen HS, Børsting C, Lagacé RE, Chang CW, Rajagopalan N, Morling N (2015) Second-generation sequencing of forensic STRs using the Ion Torrent™ HID STR 10-plex and the Ion PGM™. Forensic Sci Int Genet 14:132–140. https://doi.org/10.1016/j.fsigen.2014.09.020
Nassir R, Kosoy R, Tian C et al (2009) An ancestry informative marker set for determining continental origin: validation and extension using human genome diversity panels. BMC Genet 10:39. https://doi.org/10.1186/1471-2156-10-39
Phillips C, Devesse L, Ballard D, van Weert L, de la Puente M, Melis S, Álvarez Iglesias V, Freire-Aradas A, Oldroyd N, Holt C, Syndercombe Court D, Carracedo Á, Lareu MV (2018) Global patterns of STR sequence variation: sequencing the CEPH human genome diversity panel for 58 forensic STRs using the Illumina ForenSeq DNA Signature Prep Kit. Electrophoresis 39:2708–2724. https://doi.org/10.1002/elps.201800117
Clayton TM, Whitaker JP, Sparkes R, Gill P (1998) Analysis and interpretation of mixed forensic stains using DNA STR profiling. Forensic Sci Int 91:55–70. https://doi.org/10.1016/s0379-0738(97)00175-8
Chen K, Zhou Y-X, Li K, Qi LX, Zhang QF, Wang MC, Xiao JH (2016) A novel three-round multiplex PCR for SNP genotyping with next generation sequencing. Anal Bioanal Chem 408:4371–4377. https://doi.org/10.1007/s00216-016-9536-6
Churchill JD, Schmedes SE, King JL, Budowle B (2016) Evaluation of the Illumina(®) Beta Version ForenSeq™ DNA Signature Prep Kit for use in genetic profiling. Forensic Sci Int Genet 20:20–29. https://doi.org/10.1016/j.fsigen.2015.09.009
Huang J, Liang X, Xuan Y, Geng C, Li Y, Lu H, Qu S, Mei X, Chen H, Yu T, Sun N, Rao J, Wang J, Zhang W, Chen Y, Liao S, Jiang H, Liu X, Yang Z, Mu F, Gao S (2017) A reference human genome dataset of the BGISEQ-500 sequencer. Gigascience 6:1–9. https://doi.org/10.1093/gigascience/gix024
Xu Y, Lin Z, Tang C et al (2019) A new massively parallel nanoball sequencing platform for whole exome research. BMC Bioinf 20:153. https://doi.org/10.1186/s12859-019-2751-3
Chen K, Liu J, Liu S et al (2017) Methyltransferase SETD2-mediated methylation of STAT1 is critical for interferon antiviral activity. Cell 170:492–506.e14. https://doi.org/10.1016/j.cell.2017.06.042
Fehlmann T, Reinheimer S, Geng C et al (2016) cPAS-based sequencing on the BGISEQ-500 to explore small non-coding RNAs. Clin Epigenetics 8:123. https://doi.org/10.1186/s13148-016-0287-1
Zhou Y, Liu C, Zhou R et al (2019) SEQdata-BEACON: a comprehensive database of sequencing performance and statistical tools for performance evaluation and yield simulation in BGISEQ-500. BioData Min 12:21. https://doi.org/10.1186/s13040-019-0209-9
Goodwin S, McPherson JD, McCombie WR (2016) Coming of age: ten years of next-generation sequencing technologies. Nat Rev Genet 17:333–351. https://doi.org/10.1038/nrg.2016.49
Natarajan KN, Miao Z, Jiang M et al (2019) Comparative analysis of sequencing technologies for single-cell transcriptomics. Genome Biol 20:70. https://doi.org/10.1186/s13059-019-1676-5
Fang C, Zhong H, Lin Y, Chen B, Han M, Ren H, Lu H, Luber JM, Xia M, Li W, Stein S, Xu X, Zhang W, Drmanac R, Wang J, Yang H, Hammarström L, Kostic AD, Kristiansen K, Li J (2018) Assessment of the cPAS-based BGISEQ-500 platform for metagenomic sequencing. Gigascience 7:1–8. https://doi.org/10.1093/gigascience/gix133
Barbisin M, Fang R, O'Shea CE, Calandro LM, Furtado MR, Shewale JG (2009) Developmental validation of the Quantifiler Duo DNA Quantification kit for simultaneous quantification of total human and human male DNA and detection of PCR inhibitors in biological samples. J Forensic Sci 54:305–319. https://doi.org/10.1111/j.1556-4029.2008.00951.x
Mulero JJ, Chang CW, Lagacé RE, Wang DY, Bas JL, McMahon TP, Hennessy LK (2008) Development and validation of the AmpFlSTR MiniFiler PCR Amplification Kit: a MiniSTR multiplex for the analysis of degraded and/or PCR inhibited DNA. J Forensic Sci 53:838–852. https://doi.org/10.1111/j.1556-4029.2008.00760.x
Elwick K, Zeng X, King J, Budowle B, Hughes-Stamm S (2018) Comparative tolerance of two massively parallel sequencing systems to common PCR inhibitors. Int J Legal Med 132:983–995. https://doi.org/10.1007/s00414-017-1693-4
Gettings KB, Kiesler KM, Vallone PM (2015) Performance of a next generation sequencing SNP assay on degraded DNA. Forensic Sci Int Genet 19:1–9. https://doi.org/10.1016/j.fsigen.2015.04.010
Dror IE, Hampikian G (2011) Subjectivity and bias in forensic DNA mixture interpretation. Sci Justice 51:204–208. https://doi.org/10.1016/j.scijus.2011.08.004
Haned H, Benschop CCG, Gill PD, Sijen T (2015) Complex DNA mixture analysis in a forensic context: evaluating the probative value using a likelihood ratio model. Forensic Sci Int Genet 16:17–25. https://doi.org/10.1016/j.fsigen.2014.11.014
Chang L, Yu H, Miao X, Zhang J, Li S Development and comprehensive evaluation of a noninvasive prenatal paternity testing method through a scaled trial
Christiansen SA-O, Jakobsen B, Børsting C et al Non-invasive prenatal paternity testing using a standard forensic genetic massively parallel sequencing assay for amplification of human identification SNPs
Zhang S, Han S, Zhang M, Wang Y Non-invasive prenatal paternity testing using cell-free fetal DNA from maternal plasma: DNA isolation and genetic marker studies
Acknowledgments
The acquisition of the study was inseparable from the efforts of volunteers and laboratory staff.
Funding
This study was funded by China Postdoctoral Science Foundation (Grant No. 2018M633526 and 2019M663745).
Author information
Authors and Affiliations
Contributions
BZ designed experiment and reviewed the manuscript. HYY, BL, and LC collected and prepared the samples. ZHL and BT developed the methods and protocols. YSS, YNW, and JNF performed the experiments. XYM, YSS, and XJG analyzed the data and prepared the original manuscript. XYM and YSS prepared the figures and tables. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics statement
All works were approved by the Institutional Review 98 Board Administration in BGI (No. BGI-IRB 16036, voted at April 29, 2016). All human samples and experiments involved were in line with the ethical standards of the institutional and national research committees and the Helsinki Declaration.
We certify that this manuscript is original and has not been published and will not be submitted elsewhere for publication while being considered by IJLM. And the study is not split up into several parts to increase the quantity of submissions and submitted to various journals or to one journal over time. No data have been fabricated or manipulated (including images) to support your conclusions. No data, text, or theories by others are presented as if they were our own.
The submission has been received explicitly from all co-authors. And authors whose names appear on the submission have contributed sufficiently to the scientific work and therefore share collective responsibility and accountability for the results.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Miao, X., Shen, Y., Gong, X. et al. A novel forensic panel of 186-plex SNPs and 123-plex STR loci based on massively parallel sequencing. Int J Legal Med 135, 709–718 (2021). https://doi.org/10.1007/s00414-020-02403-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-020-02403-z