Abstract
Telomeres are specialized structures that evolved to protect the end of linear chromosomes from the action of the cell DNA damage machinery. They are composed of tandem arrays of repeated DNA sequences with a specific heterochromatic organization. The length of telomeric repeats is dynamically regulated and can be affected by changes in the telomere chromatin structure. When telomeres are not properly controlled, the resulting chromosomal alterations can induce genomic instability and ultimately the development of human diseases, such as cancer. Therefore, proper establishment, regulation, and maintenance of the telomere chromatin structure are required for cell homeostasis. Here, we review the current knowledge on telomeric chromatin dynamics during cell division and early development in mammals, and how its proper regulation safeguards genome stability.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In eukaryotes, the genome is organized in different compaction states. Schematically, euchromatin, which defines the open compaction state, contains active genes, whereas heterochromatin, which is the repressive compacted chromatin, includes gene-poor regions and different types of repeated elements with a specific chromatin structure (Nishibuchi and Déjardin 2017). The ends of linear chromosomes, or telomeres, are protected from recognition by the DNA damage response (DDR) machinery. This prevents chromosome end-to-end fusion and cell death (Palm and de Lange 2008). In mammals, telomeres are wrapped with nucleosomes and form a heterochromatin structure (Schoeftner and Blasco 2010). Telomeres consist of an array of tandem repetitions of the hexanucleotide TTAGGG motif that spans over several kilobases (10-15 kb in humans and up to 50–100 kb in laboratory mice). Because of the semi-conservative process of DNA replication, telomeric DNA shortens at each cell cycle, a phenomenon known as the “end replication problem” (i.e., the inability of DNA polymerases to fully replicate linear genomes) (Gilson and Géli 2007). Once telomeric DNA shortening reaches a critical point, this triggers the permanent exit from cell cycle, preventing the deleterious loss of genetic material encoded by the chromosome ends. However, telomeric DNA can be elongated by the telomerase enzyme or by alternative mechanisms (Cesare and Reddel 2010) (Fig. 1, upper panel). Here, we review the current knowledge on the role of canonical chromatin factors on telomere dynamics, particularly how the telomere chromatin structure regulates telomere maintenance in meiosis and in early development.
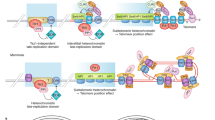
Chromatin structure of human and mouse telomeres. Telomeric chromatin is organized in a closed state with repressive histone posttranslational modifications (PTM), similar to those observed at pericentric heterochromatin (PCH) domains. Telomeric nucleosomes are labeled with H3K9me3 and possibly H4K20me3, while subtelomeric DNA is also methylated at CpG dinucleotides. H3.3-containing nucleosomes are provided by the ATRX/DAXX complex. Although telomeric chromatin is heterochromatic, subtelomeric regions contain promoters that promote the transcription of the long non-coding RNA TERRA. Telomere length is maintained by the action of the enzyme telomerase that catalyzes the addition of extra TTAGGG repeats to chromosome ends. Shelterin complexes bind along the telomere and protect from recognition by the DNA damage pathway. For graphical clarity, only one shelterin complex (TRF1, TRF2, POT1a/b, TPP1, and TIN2) is shown (top). In a subset of cancer cells, telomerase is not used to elongate telomeres. Instead, alternative lengthening of telomere (or ALT), allows recombination-mediated elongation. ATRX is frequently lost in ALT cells. Repressive marks are reduced, and TERRA RNA is more abundant, leading to putative recombinogenic DNA-RNA hybrids (R-loops). Variant repeats in ALT cells can induce binding of NR2C/F transcription factors to telomeres, leading to chromosomal rearrangements and genomic instability (bottom)
Maintenance of telomere chromatin structure in mammalian cells
Components of telomeres that protect against DDR
The Shelterin complex is one of the main components of mammalian telomeres (Palm and de Lange 2008) and consists of six main proteins in mammals: TRF1, TRF2, POT1 (POT1a and POT1b in mice), TPP1, and TIN2 (Fig. 1). TRF1, TRF2, and POT1 bind to telomeric double- and single-stranded DNA, respectively. The current model of Shelterin assembly suggests that each complex binds independently to telomeric DNA repeats according to a “beads on a string” pattern, the length of which is proportional to the telomere length (Erdel et al. 2017). Then, Shelterin protects the telomere end from recognition by the DDR machinery, thus preventing deleterious end-to-end chromosomal fusions (van Steensel et al. 1998). TRF2 is a key player in telomeric DNA protection by regulating T-loop formation at telomeres and by suppressing DDR (reviewed in Feuerhahn et al. 2015). This last function could depend on Shelterin-mediated chromatin compaction, thereby preventing telomere expansion and hindering its recognition by the DDR machinery (Bandaria et al. 2016). However, two recent studies using stochastic optical reconstruction microscopy (STORM) and assay for transposase-accessible chromatin with high-throughput sequencing (ATAC-seq) showed robust DDR localization at telomeres following co-depletion of TRF1 and TRF2, despite the fact that their depletion did not significantly affect telomeric chromatin compaction and accessibility (Timashev et al. 2017; Vancevska et al. 2017). This suggests that telomere recognition by DDR is most likely due to changes in telomeric chromatin structure and composition rather than to decompaction.
Moreover, in mammalian cells, telomeres and subtelomeric regions harbor specific histone posttranslational modifications (PTMs) and proteins that are typically found at pericentric heterochromatin (PCH), such as trimethylation of lysine 9 of histone H3 (H3K9me3), trimethylation of lysine 20 of histone H4 (H4K20me3), hypoacetylation of histone H3 and H4, and HP1 proteins (Schoeftner and Blasco 2010). However, H4K20me3 was not found at telomeres using mass spectrometry approaches (Saksouk et al. 2014). Subtelomeric regions that contain CpG dinucleotides are also methylated. Like PCH, telomeric DNA is considered to be in a closed and repressed heterochromatic environment. Nevertheless, transcription by RNA polymerase II is observed at subtelomeric regions, leading to the production of the long non-coding RNA TERRA that coats telomere ends (López de Silanes et al. 2014). Although its precise function remains enigmatic, TERRA was proposed to play several roles in telomere maintenance (reviewed in Azzalin and Lingner 2015).
ATRX and alternative lengthening of telomeres
By perturbing the chromatin state of telomeres, several studies demonstrated the critical influence of the chromatin structure on telomere homeostasis and length (O’Sullivan and Almouzni 2014). Stem cells and most cancer cells use telomerase to add de novo TTAGGG repeats and prevent telomere shortening in order to maintain high cell proliferation (Fig. 1, upper panel). However, a subset of cancer cells (about 10–15%) do not rely on telomerase activity but use a recombination-mediated alternative lengthening of telomere (ALT) mechanism (ALT cells) (Dunham et al. 2000) (Fig. 1, bottom panel). ALT cells can be identified by the presence of PML bodies that contain telomeric DNA, Shelterin proteins, DNA repair factors and chromatin proteins, and that are known as ALT-associated PML bodies (APBs). Other ALT features include telomere clustering, extrachromosomal DNA of telomeric repeats (both linear and circular DNA), telomere sister-chromatid exchange (T-SCE), telomere length heterogeneity, and absence of telomerase (O’Sullivan and Almouzni 2014). Although these features are often considered as ALT hallmarks, they may be observed also in non-ALT cells in some conditions. For instance, human embryonic stem (ES) cells contain extra-circular telomeric DNA that results from trimming of overly long telomeres (Rivera et al. 2016). Therefore, these characteristics are most likely to be the consequence rather than the causality of recombination occurring at telomeres. However, recombination is a requirement for ALT maintenance, because depletion of homologous recombination proteins impairs ALT and results in telomere shortening (Zhong et al. 2007). ALT cells contain variant telomeric repeats that are recognized by the nuclear orphan receptor NR2C/F that can promote chromosomal rearrangements (Déjardin and Kingston 2009; Marzec et al. 2015). Indeed, ALT-mediated telomere maintenance is associated with increased genomic instability.
Because of its occurrence in cancer cells, it is important to understand the molecular mechanisms and factors involved in ALT induction. Interestingly, absence of ATRX expression or localization at telomeres is a frequent feature of ALT cancer cell lines (Heaphy et al. 2011; Lovejoy et al. 2012) (Fig. 1). ATRX is a chromatin remodeler that contains a SWI/SNF2-type ATPase/helicase motif and a plant homeodomain-like zinc finger (Watson et al. 2015). ATRX forms a complex with the histone H3.3 chaperone DAXX that regulates histone loading at telomeres (Wong et al. 2010; Lewis et al. 2010). Strikingly, when ectopic ATRX is expressed in ATRX-null human osteosarcoma U2OS cells, the ALT pathway is reduced. This effect requires DAXX function (Clynes et al. 2015; Napier et al. 2015). Several hypotheses were proposed to explain how ATRX mutations promote ALT (reviewed in Amorim et al. 2016). ATRX is involved in the formation of repressed heterochromatin structures at several genomic loci, such as intracisternal A particle (IAP) retrotransposons and imprinted loci (Voon et al. 2015; Sadic et al. 2015). Loss of ATRX function leads to reduced incorporation of histone H3.3; therefore, chromatin marks cannot be maintained, presumably leading to telomere derepression and increased TERRA transcription (Udugama et al. 2015; Nguyen et al. 2017). Transcribed telomeric repeats form RNA-DNA hybrids (R-loops) that are supposed to promote the formation and stabilization of G-quadruplexes at telomeric repeats. As G-quadruplexes can impair replication fork progression, the increased rate of replication fork stalling might lead to DNA damage and drive homology search and recombination at telomeric regions (Arora et al. 2014).Another study suggested that in the absence of ATRX, the histone variant macroH2A1.1 binds to the poly(ADP-ribose) polymerase tankyrase 1, thus preventing its binding to telomeres and the resolution of cohesion between sister chromatids. Persistent cohesion would then promote recombination by T-SCE (Ramamoorthy and Smith 2015). However, activation of the ALT pathway might not depend only on ATRX. Indeed, ATRX inactivation in primary or immortalized cell lines does not trigger ALT (Napier et al. 2015). Moreover, ALT is more frequently observed in cancer cells of mesenchymal origin (Jiao et al. 2011; Heaphy et al. 2011). These observations suggest that additional chromatin or cellular events (e.g., DNA damage at telomeres) could be required to promote ALT in the absence of ATRX (Hu et al. 2016). For instance, co-depletion of the two paralogs of anti-silencing function 1 (ASF1a and ASF1b) leads to ALT (O’Sullivan et al. 2014). ASF1 is a histone chaperone of both H3.1/H3.3-H4 dimers that plays a critical role in nucleosome transfer during DNA replication. Thus, destabilization of nucleosomes at telomeres could affect the chromatin state, leading to ALT induction.
Safeguarding telomeric chromatin during DNA replication
DNA replication involves the faithful duplication of the genetic material and proper re-establishment of the parental chromatin states in both daughter strands. Chromatin reassembly after the replication fork passage requires efficient maintenance mechanisms to preserve the epigenetic information and cell identity. For instance, replication fork stalling might affect proper transfer of nucleosomes and maintenance of chromatin marks, thus impairing the binding of readers to their specific histone PTM (Groth et al. 2007; Jasencakova et al. 2010). Purification and analysis of the proteins present on chromatin associated with newly synthesized DNA showed that heterochromatic proteins, such as SUV39H1/2, DNMT1, and HP1, are readily found on nascent chromatin, while the re-establishment of H3K9me3 on parental and new histones occurs over several cell cycles (Alabert et al. 2014, 2015).
As telomeres resemble fragile sites, they may challenge the DNA replication machinery. Moreover, ALT cells undergo replication stress due to the presence of long, recombinogenic telomeres. Therefore, specific proteins are required for proper replication of the chromosome ends and for preserving genomic integrity (Higa et al. 2017). ATRX is critical for resolving DNA structures, such as G4-quadruplexes, that could induce replication fork stalling at repetitive regions of the genome, including telomeres (Law et al. 2010). As ATRX interacts with H3K9me3, through its ADD (ATRX-DNMT3-DNMT3L) domain, and with HP1, through a conserved motif (Iwase et al. 2011), improper maintenance of nucleosomes and histone PTM at telomeres might prevent ATRX binding, thus compromising telomeric DNA stability. Replication defects could impair ATRX loading and affect telomere maintenance. Several Shelterin components also participate in the proper maintenance of telomere stability by recruiting specialized enzymes to resolve deleterious secondary DNA structures that might form at telomeres. For instance, TRF1 and TRF2 associate with the helicases Werner syndrome RecQ helicase (WRN) and Bloom syndrome protein (BLM) that are involved in the resolution of G-quadruplexes and Holliday junctions. The functions of these two helicases do not fully overlap. WRN can promote the restart of collapsed forks by break-induced replication (BIR). WRN associates with SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily A-like 1 (SMARCAL1), another helicase that interacts directly with replication protein A (RPA) and is involved in replication fork restart and genome stability (Ciccia et al. 2009). In the absence of SMARCAL1, collapsed replication forks at telomeres are converted to DNA double-strand breaks that promote end-to-end chromosome fusion and compromise cell viability (Cox et al. 2016). Thus, SMARCAL1, BLM, and WRN are essential factors to prevent replicative stress of both normal and ALT telomeric DNA. T-loops also represent a barrier to replication forks and must be unwound during DNA synthesis. This could be partly mediated by the helicase RTEL1 (Vannier et al. 2012; Sarek et al. 2015) because RTEL1 loss induces replication fork stalling and C-circle accumulation.
Replication of chromosome ends is not only a matter of replicative stress management but also of preservation of the chromatin state. Indeed, subtelomeric regions contain genes and TERRA promoters, the activity of which must be regulated. Modifications in the chromatin status or DNA methylation levels of telomere and subtelomeres can alter TERRA expression (Schoeftner and Blasco 2008). Uncontrolled TERRA expression might favor the accumulation of RNA-DNA hybrid structures that could impair fork progression and stability. In mouse ES cells, the telomeric protein RIF1 associates with multiple H3K9 histone methyltransferases to maintain subtelomeric heterochromatin silencing (Dan et al. 2014). Upon Rif1 knockdown, this heterochromatic structure is lost and the subtelomeric Zscan4 gene is derepressed, leading to an overrecombination phenotype (Zscan4 function is discussed below). Similarly, the epigenetic regulator structural maintenance of chromosomes (SMC) flexible hinge domain-containing 1 (SMCHD1) is involved in the silencing of subtelomeric DUX4 repeats (Lemmers et al. 2012). In human cells, DUX4 is localized in the subtelomeric region of chromosome 4q. This locus is composed of tandem repeats of 10 to 100 D4Z4 units organized in a condensed heterochromatic state in somatic cells. Genetic or chromatin alterations of the repeat array are associated with facioscapulohumeral dystrophy (FSHD) (reviewed in Daxinger et al. 2015). SMCHD1 is a non-canonical SMC protein identified in a screen for genes that induce a variegated expression pattern of a multi-copy transgene in the mouse (Blewitt et al. 2005). In SMCHD1 heterozygous individuals, SMCHD1 levels are reduced and this was correlated with increased DUX4 expression. Although many facets of SMCHD1 regulation are still under investigation, it appears essential for the maintenance of the heterochromatic state of tandem repeats, including telomeric DNA.
Thus, maintenance of the chromatin state of telomeres in daughter cells is vital for genome stability and cell identity. The interplay between chromatin regulators and the replication machinery or Shelterin proteins is an efficient system to perpetuate chromatin states as the cells divide. In the next chapters, we will discuss how chromosome ends are regulated during the drastic chromatin remodeling that occurs during germ cell differentiation and early embryonic development.
Chromatin dynamics at telomeres during early mammalian development
Telomere structure in the germline
The germline has the critical task of producing the male and female gametes (i.e., spermatozoa and oocytes) through a process called gametogenesis. Importantly, during this process, male and female germ cells undergo meiosis, a specialized cell division characterized by one round of DNA synthesis followed by two rounds of cell division, resulting in the production of haploid gametes. During and after meiosis, germ cells undergo chromatin remodeling, leading to specific DNA and histone methylation patterns, and to histone to protamine exchange in male germ cells (Kota and Feil 2010). Perturbation of these processes can result in the reduction or loss of germ cells, leading to infertility, or in aneuploidy, leading to abortive pregnancies or congenital abnormalities in the newborn. Besides their role in protecting the chromosome ends, telomeres also contribute to the dynamic process of germ cell development (Reig-Viader et al. 2016; Keefe 2016). The following section will focus on recent insights on the role of the telomere structure in germ cell function. The chromatin dynamics in germ cells has been described elsewhere (Kota and Feil 2010; Gill et al. 2012).
In male germ cells, high telomerase activity is considered as a hallmark of undifferentiated spermatogonia (Pech et al. 2015). Then, telomerase activity progressively declines during gametogenesis and is absent in spermatozoa (Wright et al. 1996; Eisenhauer et al. 1997; Achi et al. 2000). Nevertheless, telomere length in spermatozoa is among the longest in mammalian cells. Importantly, it has been shown that sperm telomere length correlates with embryo quality and thus with good clinical outcome in in vitro fertilization procedures (Yang et al. 2015). Moreover, in late-generation telomerase-null mice, the germ cell compartment is strongly depleted, leading to almost empty seminiferous tubules (Lee et al. 1998). Thus, telomerase activity appears critical for male germ cell homeostasis, although it cannot be excluded that other lengthening mechanisms might take place during spermatogenesis (Tanemura et al. 2005). Conversely, and despite some discrepancies, telomeres in female germ cells do not elongate throughout oogenesis. Telomere length in oocytes is shorter than in somatic cells, most likely due to low or no telomerase activity and long exposure to reactive oxygen species before ovulation (Keefe et al. 2006; Liu et al. 2007).
Interestingly, in late-generation telomerase-null mice, meiosis is impaired due to altered synapsis of homologous chromosomes and decreased recombination (Liu et al. 2004b). Thus, proper telomere length homeostasis is essential for gametogenesis. During meiosis, telomeres tether chromosomes together to the nuclear envelope, leading to the formation of the “bouquet”, a characteristic structure of clustered telomeres (Scherthan 2007). This “bouquet” conformation promotes alignment and pairing of homologous chromosomes through cytoplasmic microtubule movements. Telomeres are attached to the nuclear membrane by their interaction with the linker of nucleoskeleton and cytoskeleton (LINC) complex that includes the highly conserved transmembrane proteins KASH5 and SUN1 (Shibuya and Watanabe 2014). The LINC complex connects telomeres to cytoplasmic motors (i.e., dynein and dynactin), allowing chromosome pairing (Ding et al. 2007). Telomere association with the LINC complex is mediated by the TERB1/2-MAJIN complex (Shibuya et al. 2015). TERB1 is a meiosis-specific protein with affinity for telomeric DNA (Daniel et al. 2014). In early prophase, a chimeric complex of Shelterin and TERB1/2-MAJIN is bound to telomeres. During meiosis progression, Shelterin is released from the complex, leaving only TERB1/2-MAJIN on the telomere. This meiosis-specific complex is essential for proper chromosome tethering and recombination, while protecting the chromosome ends. Therefore, and despite the fact that the Shelterin complex is considered as a hallmark of mammalian telomeres (Palm and de Lange 2008), in mature sperm cells, telomeres are specifically anchored to the nuclear periphery (Haaf and Ward 1995) through a sperm-specific protein complex that does not include TRF1 and TRF2, but contains a sperm-specific variant of histone H2B (Gineitis et al. 2000).
The higher order chromatin structure of sperm in mice specifies regulatory information to be used during embryonic development (Jung et al. 2017). However, it is not known whether there is structural information embedded in telomeric DNA that could be transmitted to the embryo. Indeed, only few data are available on the composition of telomeric chromatin in germ cells. TERRA is highly abundant in meiotic prophase I of both male and female human and mouse germ cells (Reig-Viader et al. 2014). TERRA precise role in this context is unclear, but it could be involved in telomerase regulation during gametogenesis or chromosomal movements, particularly homologous chromosome synapsis and pairing during meiosis. Indeed, in interphasic cancer cells, TERRA transcription correlates with higher telomere movements (Arora et al. 2012). Moreover, in somatic and ES cells, short telomeres or altered telomeric DNA structure can promote TERRA transcription (Schoeftner and Blasco 2008). Interestingly, in human somatic cells, H3K9me3 enrichment at telomeres might repress TERRA transcription, presumably through the action of SUV39H1 (Arnoult et al. 2012), while in human ES cells, reduction of TERRA expression impairs SUV39H1 recruitment and promotes telomere lengthening (Zeng et al. 2017). Thus, certain TERRA-dependent targeting of chromatin regulator might be cell specific. In mouse germ cells, loss of Suv39h1 and Suv39h2 results in impaired synapsis between homologous chromosomes, leading to spermatogenesis failure and infertility (Peters et al. 2001). However, this seems to be related to the loss of H3K9me3 at PCH and aberrant centromere clustering (Takada et al. 2011). Similarly, loss of Setdb1, another H3K9 methyltransferase, in the female germline leads to altered kinetochore-spindle interactions, bipolar spindle organization, and chromosome segregation defects (Kim et al. 2016; Eymery et al. 2016). However, the telomeric structure of Setdb1 −/− germ cells was not specifically evaluated. Therefore, the precise molecular composition of telomeric chromatin in germ cells awaits future investigations.
Telomere dynamics during early development
Mammalian development starts by the fusion of two highly differentiated cells, the sperm and the oocyte. This event, commonly referred to as fertilization, marks the beginning of early embryonic development and is characterized by the formation of a totipotent 1-cell zygote. This unique cell will further divide and give rise to the whole embryo and extra-embryonic tissues, such as the placenta. As the embryo develops, the DNA methylation and histone PTM profiles at chromatin domains undergo extensive maturation, associated with changes in the expression of genes and repeated elements (Albert and Peters 2009). Proper and timed reorganization of the chromatin landscape in early embryos is required for their successful development (Beaujean 2014). Due to the limited number of available cells, the first studies were predominantly performed by microscopic analysis using indirect immunofluorescence or tagged proteins in combination with mouse genetic models (Santos et al. 2005; Burton and Torres-Padilla 2010). These studies revealed that broad genome domains show highly dynamic DNA methylation and histone PTM patterns during preimplantation development and that impairing the establishment of such patterns is deleterious for the embryo (Posfai et al. 2012). Due to their role in chromosomal stability and cellular homeostasis, it is important to understand telomere contribution and regulation during preimplantation embryo development.
First of all, the establishment of proper telomere length is of crucial importance for reproduction/fertility and embryo development. Knockout mouse models gave the first insights of telomere elongation role in mammalian development. Mice lacking the telomerase RNA component (mTR −/−) do not show any obvious phenotype and can be maintained for at least six generations, suggesting that the overly long telomeres of laboratory mice could compensate for the telomeric sequence reduction. Nevertheless, telomeric DNA is gradually lost at each generation with increasing incidence of chromosomal abnormalities (Blasco et al. 1997), particularly in highly proliferative cells. Indeed, late-generation mTR −/− mice have hematopoietic defects, impaired mitogen-induced proliferation of primary splenocytes, and severe germ cell depletion (Lee et al. 1998). Male germ cells, which require a high level of telomerase activity for their efficient maintenance, are strongly depleted in these animals, resulting in decreased fertility (Lee et al. 1998; Pech et al. 2015). Although gametes from earlier generation mTR −/− mice (generation 2 or 3) can be used for in vivo or in vitro fertilization, they are less efficient than wild-type gametes and lead to aberrant embryos with pronucleus formation defects, cytofragmentation, and impaired preimplantation development (Liu et al. 2002). Of note, if short telomeres increase the risk of genomic instability, too long telomeric DNA also can compromise telomere stability and should similarly be avoided. In human and mouse ES cells, telomeric DNA trimming is performed by NBS1/XRCC3 (Rivera et al. 2016). Whether these complexes also regulate telomeric length during early embryonic development is currently not known.
Work from Liu and colleagues suggests that telomeres are elongated dramatically in the absence of any detectable telomerase activity after egg fertilization and activation (Liu et al. 2007). This surprising result warrants further investigations, as it implies the activation of an ALT-like mechanism at this developmental stage. During the first cell cycles, telomere length increases by several kilobases. This is incompatible with the sole activity of telomerase because it allows only the addition of a few hundred base pairs per division. Interestingly, early embryos show high T-SCE rate and co-localization of recombination proteins with TRF1 foci, a common phenotype of ALT cancer cells. In embryos, about half of the telomeres are localized close to PCH, while the other half is dispersed in the nucleoplasm, which is consistent with the acrocentric nature of mouse chromosomes (Aguirre-Lavin et al. 2012). These studies did not report any strong clustering of telomeres, which could be indicative of an ALT phenotype (Marzec et al. 2015). However, as embryonic chromatin is highly dynamic, these events might not be easily captured when using fixed material and might require live-imaging techniques. On the other hand, telomerase activity is clearly detected at the morula-blastocyst transition in mammals, when telomere length is stabilized (Wright et al. 1996; Schaetzlein et al. 2004). These intriguing observations suggest that the telomere maintenance mechanism switches from recombination-based “ALT-like” elongation after fertilization to telomerase activation prior to the blastocyst stage (Fig. 2). In human cancer cells, these two mechanisms are not always mutually exclusive. Perrem et al. (2001) and Cerone et al. (2001) describe co-occurrence of ALT with telomerase. In cancer cells, the ALT pathway generates heterogenous telomere length with very long and also very short telomeres. Such short telomeres could be generated by breakage in the telomere tract, which could either be extended by break-induced replication (BIR) or by the telomerase to ensure genome stability (Ford et al. 2001; Dilley et al. 2016). In contrast, it does not seem that very short telomeres are generated during early embryonic development, suggesting that the mechanism at play is not a typical ALT pathway. It could be that the transition between ALT-like and telomerase telomere lengthening might not be abrupt and that both mechanisms could co-exist during a short period in order to restore a homogeneous telomere length at the end of the preimplantation development. Moreover, it is not clear why and how telomeres are elongated by a telomerase-independent pathway in early embryos. Studies in cultured mammalian cells suggest that alteration of the telomere chromatin state could favor the ALT pathway (O’Sullivan and Almouzni 2014). Indeed, the chromatin landscape during early embryonic development is highly dynamic and more open than that of somatic cells (Wu et al. 2016). Single-nuclei Hi-C studies suggest that the chromatin of mouse preimplantation embryos exist in a “relaxed” state after fertilization, while higher-order structures are progressively re-established as embryonic development proceed (Du et al. 2017; Ke et al. 2017). Whether such “openness” of the genome at the early stages of development could favor the ALT pathway is unknown. Based on the present knowledge from studies of the ALT pathway in cancer cells and early embryonic development, several factors could contribute to telomere lengthening control in preimplantation embryos. Here, we will focus on the ATRX/DAXX complex and DUX/ZSCAN4 pathway (Fig. 2).
Telomere dynamics during mouse embryo preimplantation development. Extensive chromatin remodeling occurs at the onset of embryo development, with a histone-to-protamine replacement after fertilization, formation of the two parental pronuclei and major zygotic genome activation (ZGA) at the two-cell stage. In this environment, telomeres are elongated by a recombination-mediated mechanism that is telomerase-independent at the one- and two-cell stage. It is not known whether this mechanism is also used at later stages. Then, telomerase activity is readily detected starting from the morula-blastocyst transition. In mouse zygotes, ATRX labels PCH, and by the morula stage, ATRX is clearly targeted to telomeres. Telomere chromatin maturation might allow ATRX targeting by yet unknown mechanisms, such as recognition of histone PTM, a specific chromatin factor or binding to TERRA that is expressed at the 4-cell stage (TERRA dynamics at later stages are still unclear). DUX, a major driver of ZGA in both mouse and human zygotes, promotes expression of endogenous retroviruses (MERVL), H3K9 lysine demethylases from the Kdm4 family and Zscan4 that favors telomere elongation by recombination in mouse ES cells
The ATRX/DAXX complex
As mentioned before, the ATRX/DAXX complex is a key repressor of the ALT pathway. Mouse Atrx or Daxx genetic ablation is embryonic lethal at E9.5 (Michaelson et al. 1999; Garrick et al. 2006). Daxx −/− embryos are growth-retarded and show massive apoptosis, while embryos in which Atrx is inactivated at the 8–16-cell stage grow and implant normally, but die later because of defects in extraembryonic trophectoderm formation. The presence of defects in telomere maintenance in these embryos was not determined. Atrx and Daxx mRNA are maternally provided in the embryo (RNA sequencing data from Park et al. 2013). Interestingly, ATRX and DAXX undergo a spatiotemporal re-localization during preimplantation embryo development. After fertilization, ATRX is localized at maternal condensed chromosomes, then it labels predominantly PCH domains (which assembles around prenucleolar bodies) of maternal origin where it represses transcription of the underlying major satellite repeats (De La Fuente et al. 2015). Conversely, DAXX is concentrated on paternal PCH, defining an interesting epigenetic asymmetry that contrasts with the classic view of ATRX and DAXX co-localization at these regions, as observed in oocytes and most somatic cells. Interestingly DAXX targeting at IAP retrotransposons in ES cells does not rely on ATRX (Hoelper et al. 2017). Thus, ATRX and DAXX could operate in different complexes during development and assemble on chromatin with different kinetics. Strikingly, by the morula stage, ATRX and DAXX are mostly co-localized at telomeres (He et al. 2015). Therefore, ATRX/DAXX absence from telomeres in the early stages of development is compatible with an ALT-like mechanism and suggests that telomeric chromatin undergoes changes during development, thus allowing the specific loading of ATRX and DAXX at the end of preimplantation. The reasons for ATRX and DAXX absence at telomeres in the early stages of embryo development are unclear. Specific requirements for ATRX targeting (i.e., H3K9me3 or HP1 proteins) could be missing on telomeric chromatin just after fertilization and be re-established only later during development. The paternal genome undergoes massive protamine-to-histone exchange right after fertilization and this could impair ATRX loading. At the one- and two-cell stage, H3K9me2/3 is low on the paternal pronucleus and global levels on both pronuclei start to increase at the four-cell stage onward (Liu et al. 2004a; Yeo et al. 2005). Such dynamics in the H3K9 methylation could thus favor ATRX loading at telomeres only latter during development after histone methylation levels are restored. Also, it would be interesting to test whether the kinetics of ATRX and DAXX loading at telomeres differs according to the parental origin (telomere transgenerational inheritance and parental origin are discussed below).
Similarly, it is not known how ATRX and DAXX are loaded at telomeres later in development. It was suggested that in mouse ES cells, DNA hypomethylation promotes ATRX/DAXX loading at tandem repeats (e.g., IAP or telomeres). This could lead to SUV39H1 recruitment to mediate H3K9me3 and heterochromatinization (He et al. 2015). Such model is appealing because the genome undergoes global hypomethylation during preimplantation and this could favor ATRX binding (Messerschmidt et al. 2014). However, telomeres and IAP are not regulated by DNA methylation in ES cells, and IAP retains high levels of DNA methylation throughout preimplantation (Maksakova et al. 2013; Arand et al. 2015). Furthermore, subtelomeric DNA methylation is not altered in induced pluripotent stem (iPS) cells generated from mouse embryo fibroblasts, suggesting that, at least during in vitro reprogramming, DNA methylation is maintained at these loci (Marion et al. 2009). Whether this also applies to preimplantation embryos is not clear and will require further investigations.
During early embryo development, several transposable elements and tandem repeats undergo massive transcription (Peaston et al. 2004; Probst et al. 2010; Fadloun et al. 2013). As exemplified with PCH, transcription of major satellites is required for efficient maturation of these loci and embryo development progression (Casanova et al. 2013). TERRA signals are detected by RNA FISH by the 4-cell stage (Probst et al. 2010). We could hypothesize that TERRA transcription in the embryo represents a starting signal for recruiting ATRX and/or other chromatin factors to telomeres and establishing a canonical H3K9me3/HP1 heterochromatic state (Nguyen et al. 2017). Thus, deciphering the regulation of ATRX/DAXX recruitment at telomeres during natural embryo development could help better understanding heterochromatin establishment at telomeric DNA.
The DUX/ZSCAN4 pathway
After fertilization, maternally inherited transcripts control development before their rapid clearance. The embryonic genome is then activated (zygotic genome activation (ZGA)), marking the maternal-to-zygotic transition (Svoboda et al. 2015). In mice, a minor ZGA occurs at the one-cell stage followed by a major ZGA at the two-cell stage (four- to eight-cell stage in humans). Zinc finger and SCAN domain-containing 4 (Zscan4) is among the genes that are specifically expressed during ZGA (Falco et al. 2007) (Fig. 2, bottom panel). In mice, the Zscan4 cluster includes six genes and three pseudogenes, while humans only have one copy of ZSCAN4. In preimplantation embryos, Zscan4 expression peaks at the two-cell stage in mice and at the four- to eight-cell stage in humans (Vassena et al. 2011; Ishiguro et al. 2017). Interestingly, a subpopulation of proliferating mouse ES cells (about 1–5% of all cells) expresses transiently transcripts that are detected in two-cell embryos, such as the MERVL endogenous retrovirus and the Zscan4 cluster. Accordingly, these cells are referred to as “2C-like” cells (Macfarlan et al. 2012). ES cells that express Zscan4 (“Zscan4-associated event” or “Z4 event”) are characterized by rapid and telomerase-independent telomere lengthening. Telomeres are bound by recombination proteins and undergo T-SCE (Zalzman et al. 2010). In ALT cancer cells these features are associated with genomic instability. Conversely, Zscan4 expression in ES cells is associated with improved karyotypes and developmental potency and is required for long-term culture of ES cells (Amano et al. 2013). Similarly, Zscan4 improves iPS cell reprogramming efficiency, when expressed for only a few days (Hirata et al. 2012; Jiang et al. 2013). However, uncontrolled Zscan4 expression leads to over-recombination, telomere length heterogeneity, and chromosome fusions (Dan et al. 2014). In preimplantation mouse embryos, Zscan4 genetic ablation or sustained expression impairs development progression and blastocyst implantation (Falco et al. 2007). Thus, fine-tuned Zscan4 expression is required for proper genome stability in ES cells and early embryo development.
Interestingly, the chromatin state of ES cells that express Zscan4 is characterized by global DNA hypomethylation, histone hyperacetylation, and transcription of heterochromatin domains (pericentromeres, telomeres, and retrotransposons) (Akiyama et al. 2015; Eckersley-Maslin et al. 2016). Hypomethylation of subtelomeric DNA is associated with recombination-mediated telomere lengthening, while hypermethylation, as observed in Tet −/− ES cells, is associated with reduced recombination and telomere shortening (Gonzalo et al. 2006; Yang et al. 2016). Tet1 knockdown in epiblast-like cells (EpiLC) results in Zscan4 decrease and telomere shortening, while Tet TKO in ES cells cultured in 2i conditions (“ground state”) induces increased Zscan4 expression and telomere elongation (Lu et al. 2014; Khoueiry et al. 2017). Thus, Tet enzymes may regulate Zscan4 levels with different outcomes depending on the cell type or pluripotency status. Accordingly, the hypomethylated state of Zscan4-expressing cells should favor recombination-mediated telomere elongation. The expression and localization of activating and repressive chromatin regulators in ZSCAN4-positive cells suggest that ZSCAN4 and, most likely, other factors promote first an open and then repressed chromatin state, presumably to allow the transient nature of the “2c-like” state in ES cells (Akiyama et al. 2015). Indeed, a prolonged “2c-like” state in ES cells might lead to the irreversible erasure of specific chromatin marks, such as parental imprints (Eckersley-Maslin et al. 2016).
How are Zscan4 genes induced during ZGA? In ES cells, the transcription factor TBX3 induces Zscan4 expression through modulation of DNMT3b and TET2 levels, thereby reducing DNA methylation at the Zscan4 locus (Dan et al. 2013). However, single-cell transcriptomic data in preimplantation mouse embryos suggest that Tbx3 expression is restricted to the inner cell mass (Nestorov et al. 2015). Therefore, a Tbx3 role in Zscan4 expression at the two-cell stage seems unlikely. Several studies identified human DUX4 and mouse Dux genes as master regulators of many genes and retrotransposons, including Zscan4, during ZGA (De Iaco et al. 2017; Hendrickson et al. 2017; Whiddon et al. 2017). What drives the expression of Dux/DUX4 is still unclear. Its expression appears to be associated with chromatin relaxation or destabilization, as observed in mouse ES cells in which the chromatin assembly factor (CAF-1) Chaf1a is downregulated and in myoblasts from patients with FSHD (Geng et al. 2012; Ishiuchi et al. 2015). The early embryo genome is characterized by an open chromatin structure and dynamic histone mobility (Bošković et al. 2014; Wu et al. 2016). This environment could favor Dux/DUX4 upregulation and consequently Zscan4 gene transcription to promote a rapid, but transient telomere lengthening and karyotype correction. Maternally and/or zygotically expressed factors that repress the 2c-like state in ES cells (such as SETDB1, HP1, and TRIM28) could participate in Dux and Zscan4 repression and allow the establishment of a canonical heterochromatin state at later developmental stages (Macfarlan et al. 2012; Maksakova et al. 2013; De Iaco et al. 2017).
Chromatin marks at telomeres and genome reprogramming
After fertilization, the embryo genome undergoes a rapid reorganization or “reprogramming” that is essential for proper ZGA, totipotency acquisition, and development. Nuclear reprogramming can by studied by somatic cell nuclear transfer (SCNT) in which a donor nucleus (usually from a somatic cell) is injected into an enucleated oocyte. The oocyte will reprogram the donor nucleus in a similar way as the reprogramming of the maternal and paternal genomes after natural fertilization (Sepulveda-Rincon et al. 2016). Another method consists in expressing keys transcription factors (Oct4, Sox2, Klf4, and Myc) in somatic cells to induce pluripotency (Takahashi and Yamanaka 2016). In this way, iPS cells have been successfully generated in different mammalian species. However, experimental reprogramming is not equivalent to what observed following natural fertilization, suggesting that somatic cells contain specific factors or chromatin determinants that are refractory to artificial reprogramming. Telomere rejuvenation is a feature of successful reprogramming and its efficiency can be enhanced by specific alterations of the chromatin state (Liu 2017). Strikingly, H3K9 methylation appears to be a roadblock to efficient reprogramming in both SCNT and iPS cells (Chen et al. 2013; Matoba et al. 2014; Chung et al. 2015). During iPS cell induction, telomeres adopt a chromatin structure similar to that of ES cells with reduced H3K9me3 (Marion et al. 2009). Reduction of H3K9me3 levels by ectopic expression of the H3K9 demethylase KDM4B in SCNT-derived embryos helps to restore transcriptional reprogramming and induces Zscan4 expression (Matoba et al. 2014). Moreover, Kdm4b or Zscan4 expression enhances telomere elongation and the successful formation of iPS cells (Jiang et al. 2013; Sridharan et al. 2013). In pluripotent cells, SUV39H1/2 and SETDB1 could control H3K9me3 levels at telomeres (García-Cao et al. 2004; Udugama et al. 2015). Both maternally inherited Suv39h2 and Setdb1 transcripts are found in the zygote (Park et al. 2013). However, the absence of ATRX binding to telomeres, which relies predominantly on H3K9me3 and HP1, suggests that H3K9me3 methyltransferases are not targeted to these regions in the zygote. Genome-wide studies of the chromatin landscape in early embryos will surely provide a better understanding of the dynamic organization of telomeres during preimplantation development.
Inheritance and parental origin
The discovery that after the histone to protamine exchange, the fraction of histones retained in mature sperm carries PTM led to the idea that male germ cells could transmit epigenetic information to the progeny (Gill et al. 2012). Moreover, the retained histones are not randomly positioned in the genome. Indeed, ChIP-sequencing analysis of nucleosomes and histone PTM in human and mouse mature sperm cells led to the identification of a specific chromatin structure at developmentally regulated genes (Hammoud et al. 2009; Brykczynska et al. 2010; Erkek et al. 2013). Removal of histone marks from sperm impairs the expression of embryonic genes in a paternal-specific manner and leads to impaired embryo development (Teperek et al. 2016). Chromatin marks and more generally higher-order chromatin structures in sperm cells (i.e., 3D organization, enhancers) are supposed to influence genome organization and gene expression in the embryo (Jung et al. 2017). Several studies reported that telomeres in human mature sperm cells contain nucleosomes (Zalenskaya et al. 2000), and that nucleosomes can be found in gene-poor regions and at several repeated elements in sperm (Carone et al. 2014; Samans et al. 2014). However, bioinformatics analysis of repeated elements is to be considered very carefully because the obvious repetitive nature of these elements requires proper data treatment to prevent misleading conclusions (Royo et al. 2016). Moreover, chromatin higher-order structure of both parental genomes is drastically remodeled from fertilization until blastocyst stage (Du et al. 2017). Therefore, it is not clear whether in gametes, telomeres have a specific chromatin structure that could be inherited and with a role in early development.
The parental origin influence, if any, on telomeric chromatin regulation is also quite enigmatic. A parental-specific chromatin structure is visible at PCH domains in the zygote. H3K9me3 and HP1 are absent from paternal PCH, while they are highly enriched at maternal PCH. Conversely, Polycomb complexes bind to these domains and repress expression of major satellites from the paternal pronucleus (Puschendorf et al. 2008). To analyze the influence of the parental origin on telomere regulation, Liu and colleagues analyzed telomere lengthening in mouse parthenotes (embryos with a genome of only maternal origin). Telomere elongation was comparable in parthenotes and in in vitro fertilized zygotes, suggesting that the factors controlling telomere lengthening in embryos are provided by the oocyte (Liu et al. 2007). Interestingly, ATAC-seq and DNAse-seq genomic analyses, which provide allelic information in early embryos, suggest that chromatin accessibility is comparable in the two parental genomes with the exception of imprinted genes (Wu et al. 2016; Lu et al. 2016). Therefore, paternal chromatin accessibility is reprogrammed shortly after fertilization to similar levels as for maternal chromatin. This suggests that the parental-specific differences in chromatin states of telomeres, if they exist, do not exert a strong influence on their regulation at the onset of development.
Conclusions
Telomeres are essential structure to ensure genome stability, tissue homeostasis, and embryo development. Telomeric chromatin plasticity is a striking feature that allows the dynamic regulation of chromosome ends following developmental cues, cell cycle, or metabolism. These plasticity features are crucial during natural development or experimental reprogramming of somatic nuclei. Thus, understanding the mechanisms that operate at telomeric DNA in these conditions will greatly help the development and optimization of assisted reproductive technologies or therapeutic reprogramming techniques (Sepulveda-Rincon et al. 2016). Furthermore, chromatin modifications at telomeric repeats can alter the telomere lengthening mode, switching to the ALT pathway (O’Sullivan and Almouzni 2014). Although telomerase is the most common mechanism of telomere maintenance in cancer cells, treatment with telomerase inhibitors can lead to the appearance of ALT (Hu et al. 2012). Thus, understating how the chromatin state can promote one or the other telomere maintenance mode could lead to the development of therapeutic strategies to enhance anti-telomerase treatments. Interestingly, ALT or ALT-like lengthening also occurs in physiological conditions (Liu et al. 2007; Neumann et al. 2013). Deciphering the role of such mechanisms during evolution and comparing the mechanisms used in other organisms should provide clues on the advantages of using a specific telomere-lengthening mode during the development and lifespan of an organism.
References
Achi MV, Ravindranath N, Dym M (2000) Telomere length in male germ cells is inversely correlated with telomerase activity. Biol Reprod 63(2):591–598. https://doi.org/10.1095/biolreprod63.2.591
Aguirre-Lavin T, Adenot P, Bonnet-Garnier A, Lehmann G, Fleurot R, Boulesteix C, Debey P, Beaujean N (2012) 3D-FISH analysis of embryonic nuclei in mouse highlights several abrupt changes of nuclear organization during preimplantation development. BMC Dev Biol 12(1):30. https://doi.org/10.1186/1471-213X-12-30
Akiyama T, Xin L, Oda M, Sharov AA, Amano M, Piao Y, Cadet JS, Dudekula DB, Qian Y, Wang W, Ko SBH, Ko MSH (2015) Transient bursts of Zscan4 expression are accompanied by the rapid derepression of heterochromatin in mouse embryonic stem cells. DNA Res Int J Rapid Publ Rep Genes Genomes 22(5):307–318. https://doi.org/10.1093/dnares/dsv013
Alabert C, Barth TK, Reverón-Gómez N, Sidoli S, Schmidt A, Jensen ON, Imhof A, Groth A (2015) Two distinct modes for propagation of histone PTMs across the cell cycle. Genes Dev 29(6):585–590. https://doi.org/10.1101/gad.256354.114
Alabert C, Bukowski-Wills J-C, Lee S-B, Kustatscher G, Nakamura K, de Lima Alves F, Menard P, Mejlvang J, Rappsilber J, Groth A (2014) Nascent chromatin capture proteomics determines chromatin dynamics during DNA replication and identifies unknown fork components. Nat Cell Biol 16(3):281–293. https://doi.org/10.1038/ncb2918
Albert M, Peters AHFM (2009) Genetic and epigenetic control of early mouse development. Curr Opin Genet Dev 19(2):113–121. https://doi.org/10.1016/j.gde.2009.03.004
Amano T, Hirata T, Falco G, Monti M, Sharova LV, Amano M, Sheer S, Hoang HG, Piao Y, Stagg CA, Yamamizu K, Akiyama T, Ko MSH (2013) Zscan4 restores the developmental potency of embryonic stem cells. Nat Commun 4:1966. https://doi.org/10.1038/ncomms2966
Amorim JP, Santos G, Vinagre J, Soares P (2016) The role of ATRX in the alternative lengthening of telomeres (ALT) phenotype. Genes 7: . doi: https://doi.org/10.3390/genes7090066
Arand J, Wossidlo M, Lepikhov K, Peat JR, Reik W, Walter J (2015) Selective impairment of methylation maintenance is the major cause of DNA methylation reprogramming in the early embryo. Epigenetics Chromatin 8(1):1. https://doi.org/10.1186/1756-8935-8-1
Arnoult N, Van Beneden A, Decottignies A (2012) Telomere length regulates TERRA levels through increased trimethylation of telomeric H3K9 and HP1α. Nat Struct Mol Biol 19(9):948–956. https://doi.org/10.1038/nsmb.2364
Arora R, Brun CM, Azzalin CM (2012) Transcription regulates telomere dynamics in human cancer cells. RNA N Y N 18(4):684–693. https://doi.org/10.1261/rna.029587.111
Arora R, Lee Y, Wischnewski H, Brun CM, Schwarz T, Azzalin CM (2014) RNaseH1 regulates TERRA-telomeric DNA hybrids and telomere maintenance in ALT tumour cells. Nat Commun 5:5220. https://doi.org/10.1038/ncomms6220
Azzalin CM, Lingner J (2015) Telomere functions grounding on TERRA firma. Trends Cell Biol 25(1):29–36. https://doi.org/10.1016/j.tcb.2014.08.007
Bandaria JN, Qin P, Berk V, Chu S, Yildiz A (2016) Shelterin protects chromosome ends by compacting telomeric chromatin. Cell 164(4):735–746. https://doi.org/10.1016/j.cell.2016.01.036
Beaujean N (2014) Epigenetics, embryo quality and developmental potential. Reprod Fertil Dev 27(1):53–62. https://doi.org/10.1071/RD14309
Blasco MA, Lee HW, Hande MP, Samper E, Lansdorp PM, DePinho RA, Greider CW (1997) Telomere shortening and tumor formation by mouse cells lacking telomerase RNA. Cell 91(1):25–34. https://doi.org/10.1016/S0092-8674(01)80006-4
Blewitt ME, Vickaryous NK, Hemley SJ, Ashe A, Bruxner TJ, Preis JI, Arkell R, Whitelaw E (2005) An N-ethyl-N-nitrosourea screen for genes involved in variegation in the mouse. Proc Natl Acad Sci U S A 102(21):7629–7634. https://doi.org/10.1073/pnas.0409375102
Bošković A, Eid A, Pontabry J et al (2014) Higher chromatin mobility supports totipotency and precedes pluripotency in vivo. Genes Dev 28(10):1042–1047. https://doi.org/10.1101/gad.238881.114
Brykczynska U, Hisano M, Erkek S, Ramos L, Oakeley EJ, Roloff TC, Beisel C, Schübeler D, Stadler MB, Peters AHFM (2010) Repressive and active histone methylation mark distinct promoters in human and mouse spermatozoa. Nat Struct Mol Biol 17(6):679–687. https://doi.org/10.1038/nsmb.1821
Burton A, Torres-Padilla M-E (2010) Epigenetic reprogramming and development: a unique heterochromatin organization in the preimplantation mouse embryo. Brief Funct Genomics 9(5-6):444–454. https://doi.org/10.1093/bfgp/elq027
Carone BR, Hung J-H, Hainer SJ, Chou MT, Carone DM, Weng Z, Fazzio TG, Rando OJ (2014) High-resolution mapping of chromatin packaging in mouse embryonic stem cells and sperm. Dev Cell 30(1):11–22. https://doi.org/10.1016/j.devcel.2014.05.024
Casanova M, Pasternak M, El Marjou F et al (2013) Heterochromatin reorganization during early mouse development requires a single-stranded noncoding transcript. Cell Rep 4(6):1156–1167. https://doi.org/10.1016/j.celrep.2013.08.015
Cerone MA, Londono-Vallejo JA, Bacchetti S (2001) Telomere maintenance by telomerase and by recombination can coexist in human cells. Hum Mol Genet 10(18):1945–1952. https://doi.org/10.1093/hmg/10.18.1945
Cesare AJ, Reddel RR (2010) Alternative lengthening of telomeres: models, mechanisms and implications. Nat Rev Genet 11(5):319–330. https://doi.org/10.1038/nrg2763
Chen J, Liu H, Liu J, Qi J, Wei B, Yang J, Liang H, Chen Y, Chen J, Wu Y, Guo L, Zhu J, Zhao X, Peng T, Zhang Y, Chen S, Li X, Li D, Wang T, Pei D (2013) H3K9 methylation is a barrier during somatic cell reprogramming into iPSCs. Nat Genet 45(1):34–42. https://doi.org/10.1038/ng.2491
Chung YG, Matoba S, Liu Y, Eum JH, Lu F, Jiang W, Lee JE, Sepilian V, Cha KY, Lee DR, Zhang Y (2015) Histone demethylase expression enhances human somatic cell nuclear transfer efficiency and promotes derivation of pluripotent stem cells. Cell Stem Cell 17(6):758–766. https://doi.org/10.1016/j.stem.2015.10.001
Ciccia A, Bredemeyer AL, Sowa ME, Terret ME, Jallepalli PV, Harper JW, Elledge SJ (2009) The SIOD disorder protein SMARCAL1 is an RPA-interacting protein involved in replication fork restart. Genes Dev 23(20):2415–2425. https://doi.org/10.1101/gad.1832309
Clynes D, Jelinska C, Xella B, Ayyub H, Scott C, Mitson M, Taylor S, Higgs DR, Gibbons RJ (2015) Suppression of the alternative lengthening of telomere pathway by the chromatin remodelling factor ATRX. Nat Commun 6:7538. https://doi.org/10.1038/ncomms8538
Cox KE, Maréchal A, Flynn RL (2016) SMARCAL1 resolves replication stress at ALT telomeres. Cell Rep 14(5):1032–1040. https://doi.org/10.1016/j.celrep.2016.01.011
Dan J, Li M, Yang J, Li J, Okuka M, Ye X, Liu L (2013) Roles for Tbx3 in regulation of two-cell state and telomere elongation in mouse ES cells. Sci Rep 3(1):3492. https://doi.org/10.1038/srep03492
Dan J, Liu Y, Liu N, Chiourea M, Okuka M, Wu T, Ye X, Mou C, Wang L, Wang L, Yin Y, Yuan J, Zuo B, Wang F, Li Z, Pan X, Yin Z, Chen L, Keefe DL, Gagos S, Xiao A, Liu L (2014) Rif1 maintains telomere length homeostasis of ESCs by mediating heterochromatin silencing. Dev Cell 29(1):7–19. https://doi.org/10.1016/j.devcel.2014.03.004
Daniel K, Tränkner D, Wojtasz L, Shibuya H, Watanabe Y, Alsheimer M, Tóth A (2014) Mouse CCDC79 (TERB1) is a meiosis-specific telomere associated protein. BMC Cell Biol 15(1):17. https://doi.org/10.1186/1471-2121-15-17
Daxinger L, Tapscott SJ, van der Maarel SM (2015) Genetic and epigenetic contributors to FSHD. Curr Opin Genet Dev 33:56–61. https://doi.org/10.1016/j.gde.2015.08.007
De Iaco A, Planet E, Coluccio A, Verp S, Duc J, Trono D (2017) DUX-family transcription factors regulate zygotic genome activation in placental mammals. Nat Genet 49(6):941–945. https://doi.org/10.1038/ng.3858
De La Fuente R, Baumann C, Viveiros MM (2015) ATRX contributes to epigenetic asymmetry and silencing of major satellite transcripts in the maternal genome of the mouse embryo. Dev Camb Engl 142(10):1806–1817. https://doi.org/10.1242/dev.118927
Déjardin J, Kingston RE (2009) Purification of proteins associated with specific genomic loci. Cell 136(1):175–186. https://doi.org/10.1016/j.cell.2008.11.045
Dilley RL, Verma P, Cho NW, Winters HD, Wondisford AR, Greenberg RA (2016) Break-induced telomere synthesis underlies alternative telomere maintenance. Nature 539(7627):54–58. https://doi.org/10.1038/nature20099
Ding X, Xu R, Yu J, Xu T, Zhuang Y, Han M (2007) SUN1 is required for telomere attachment to nuclear envelope and gametogenesis in mice. Dev Cell 12(6):863–872. https://doi.org/10.1016/j.devcel.2007.03.018
Du Z, Zheng H, Huang B, Ma R, Wu J, Zhang X, He J, Xiang Y, Wang Q, Li Y, Ma J, Zhang X, Zhang K, Wang Y, Zhang MQ, Gao J, Dixon JR, Wang X, Zeng J, Xie W (2017) Allelic reprogramming of 3D chromatin architecture during early mammalian development. Nature 547(7662):232–235. https://doi.org/10.1038/nature23263
Dunham MA, Neumann AA, Fasching CL, Reddel RR (2000) Telomere maintenance by recombination in human cells. Nat Genet 26(4):447–450. https://doi.org/10.1038/82586
Eckersley-Maslin MA, Svensson V, Krueger C, Stubbs TM, Giehr P, Krueger F, Miragaia RJ, Kyriakopoulos C, Berrens RV, Milagre I, Walter J, Teichmann SA, Reik W (2016) MERVL/Zscan4 network activation results in transient genome-wide DNA demethylation of mESCs. Cell Rep 17(1):179–192. https://doi.org/10.1016/j.celrep.2016.08.087
Eisenhauer KM, Gerstein RM, Chiu CP, Conti M, Hsueh AJW (1997) Telomerase activity in female and male rat germ cells undergoing meiosis and in early embryos. Biol Reprod 56(5):1120–1125. https://doi.org/10.1095/biolreprod56.5.1120
Erdel F, Kratz K, Willcox S, Griffith JD, Greene EC, de Lange T (2017) Telomere recognition and assembly mechanism of mammalian shelterin. Cell Rep 18(1):41–53. https://doi.org/10.1016/j.celrep.2016.12.005
Erkek S, Hisano M, Liang C-Y, Gill M, Murr R, Dieker J, Schübeler D, Vlag J, Stadler MB, Peters AHFM (2013) Molecular determinants of nucleosome retention at CpG-rich sequences in mouse spermatozoa. Nat Struct Mol Biol 20(7):868–875. https://doi.org/10.1038/nsmb.2599
Eymery A, Liu Z, Ozonov EA, Stadler MB, Peters AHFM (2016) The methyltransferase Setdb1 is essential for meiosis and mitosis in mouse oocytes and early embryos. Dev Camb Engl 143(15):2767–2779. https://doi.org/10.1242/dev.132746
Fadloun A, Le Gras S, Jost B, Ziegler-Birling C, Takahashi H, Gorab E, Carninci P, Torres-Padilla ME (2013) Chromatin signatures and retrotransposon profiling in mouse embryos reveal regulation of LINE-1 by RNA. Nat Struct Mol Biol 20(3):332–338. https://doi.org/10.1038/nsmb.2495
Falco G, Lee S-L, Stanghellini I, Bassey UC, Hamatani T, Ko MSH (2007) Zscan4: a novel gene expressed exclusively in late 2-cell embryos and embryonic stem cells. Dev Biol 307(2):539–550. https://doi.org/10.1016/j.ydbio.2007.05.003
Feuerhahn S, Chen L, Luke B, Porro A (2015) No DDRama at chromosome ends: TRF2 takes centre stage. Trends Biochem Sci 40(5):275–285. https://doi.org/10.1016/j.tibs.2015.03.003
Ford LP, Zou Y, Pongracz K, Gryaznov SM, Shay JW, Wright WE (2001) Telomerase can inhibit the recombination-based pathway of telomere maintenance in human cells. J Biol Chem 276(34):32198–32203. https://doi.org/10.1074/jbc.M104469200
García-Cao M, O’Sullivan R, Peters AHFM et al (2004) Epigenetic regulation of telomere length in mammalian cells by the Suv39h1 and Suv39h2 histone methyltransferases. Nat Genet 36(1):94–99. https://doi.org/10.1038/ng1278
Garrick D, Sharpe JA, Arkell R, Dobbie L, Smith AJH, Wood WG, Higgs DR, Gibbons RJ (2006) Loss of Atrx affects trophoblast development and the pattern of X-inactivation in extraembryonic tissues. PLoS Genet 2(4):e58. https://doi.org/10.1371/journal.pgen.0020058
Geng LN, Yao Z, Snider L, Fong AP, Cech JN, Young JM, van der Maarel SM, Ruzzo WL, Gentleman RC, Tawil R, Tapscott SJ (2012) DUX4 activates germline genes, retroelements, and immune mediators: implications for facioscapulohumeral dystrophy. Dev Cell 22(1):38–51. https://doi.org/10.1016/j.devcel.2011.11.013
Gill ME, Erkek S, Peters AHFM (2012) Parental epigenetic control of embryogenesis: a balance between inheritance and reprogramming? Curr Opin Cell Biol 24(3):387–396. https://doi.org/10.1016/j.ceb.2012.03.002
Gilson E, Géli V (2007) How telomeres are replicated. Nat Rev Mol Cell Biol 8(10):825–838. https://doi.org/10.1038/nrm2259
Gineitis AA, Zalenskaya IA, Yau PM, Bradbury EM, Zalensky AO (2000) Human sperm telomere-binding complex involves histone H2B and secures telomere membrane attachment. J Cell Biol 151(7):1591–1598. https://doi.org/10.1083/jcb.151.7.1591
Gonzalo S, Jaco I, Fraga MF, Chen T, Li E, Esteller M, Blasco MA (2006) DNA methyltransferases control telomere length and telomere recombination in mammalian cells. Nat Cell Biol 8(4):416–424. https://doi.org/10.1038/ncb1386
Groth A, Corpet A, Cook AJL, Roche D, Bartek J, Lukas J, Almouzni G (2007) Regulation of replication fork progression through histone supply and demand. Science 318(5858):1928–1931. https://doi.org/10.1126/science.1148992
Haaf T, Ward DC (1995) Higher order nuclear structure in mammalian sperm revealed by in situ hybridization and extended chromatin fibers. Exp Cell Res 219(2):604–611. https://doi.org/10.1006/excr.1995.1270
Hammoud SS, Nix DA, Zhang H, Purwar J, Carrell DT, Cairns BR (2009) Distinctive chromatin in human sperm packages genes for embryo development. Nature 460:473–478. https://doi.org/10.1038/nature08162
He Q, Kim H, Huang R, Lu W, Tang M, Shi F, Yang D, Zhang X, Huang J, Liu D, Songyang Z (2015) The Daxx/Atrx complex protects tandem repetitive elements during DNA hypomethylation by promoting H3K9 trimethylation. Cell Stem Cell 17(3):273–286. https://doi.org/10.1016/j.stem.2015.07.022
Heaphy CM, de Wilde RF, Jiao Y, Klein AP, Edil BH, Shi C, Bettegowda C, Rodriguez FJ, Eberhart CG, Hebbar S, Offerhaus GJ, McLendon R, Rasheed BA, He Y, Yan H, Bigner DD, Oba-Shinjo SM, Marie SKN, Riggins GJ, Kinzler KW, Vogelstein B, Hruban RH, Maitra A, Papadopoulos N, Meeker AK (2011) Altered telomeres in tumors with ATRX and DAXX mutations. Science 333(6041):425. https://doi.org/10.1126/science.1207313
Hendrickson PG, Doráis JA, Grow EJ, Whiddon JL, Lim JW, Wike CL, Weaver BD, Pflueger C, Emery BR, Wilcox a, Nix d, Peterson CM, Tapscott SJ, Carrell DT, Cairns BR (2017) Conserved roles of mouse DUX and human DUX4 in activating cleavage-stage genes and MERVL/HERVL retrotransposons. Nat Genet 49(6):925–934. https://doi.org/10.1038/ng.3844
Higa M, Fujita M, Yoshida K (2017) DNA replication origins and fork progression at mammalian telomeres. Genes 8(4). https://doi.org/10.3390/genes8040112
Hirata T, Amano T, Nakatake Y, Amano M, Piao Y, Hoang HG, Ko MSH (2012) Zscan4 transiently reactivates early embryonic genes during the generation of induced pluripotent stem cells. Sci Rep 2(1):208. https://doi.org/10.1038/srep00208
Hoelper D, Huang H, Jain AY, Patel DJ, Lewis PW (2017) Structural and mechanistic insights into ATRX-dependent and -independent functions of the histone chaperone DAXX. Nat Commun 8(1):1193. https://doi.org/10.1038/s41467-017-01206-y
Hu J, Hwang SS, Liesa M, Gan B, Sahin E, Jaskelioff M, Ding Z, Ying H, Boutin AT, Zhang H, Johnson S, Ivanova E, Kost-Alimova M, Protopopov A, Wang YA, Shirihai OS, Chin L, DePinho RA (2012) Antitelomerase therapy provokes ALT and mitochondrial adaptive mechanisms in cancer. Cell 148(4):651–663. https://doi.org/10.1016/j.cell.2011.12.028
Hu Y, Shi G, Zhang L, Li F, Jiang Y, Jiang S, Ma W, Zhao Y, Songyang Z, Huang J (2016) Switch telomerase to ALT mechanism by inducing telomeric DNA damages and dysfunction of ATRX and DAXX. Sci Rep 6(1):32280. https://doi.org/10.1038/srep32280
Ishiguro K-I, Nakatake Y Expression analysis of the endogenous Zscan4 locus and its coding proteins in mouse ES cells and preimplantation embryos, Chikazawa-Nohtomi N, et al (2017). In Vitro Cell Dev Biol Anim 53(2):179–190. https://doi.org/10.1007/s11626-016-0097-y
Ishiuchi T, Enriquez-Gasca R, Mizutani E, Bošković A, Ziegler-Birling C, Rodriguez-Terrones D, Wakayama T, Vaquerizas JM, Torres-Padilla ME (2015) Early embryonic-like cells are induced by downregulating replication-dependent chromatin assembly. Nat Struct Mol Biol 22(9):662–671. https://doi.org/10.1038/nsmb.3066
Iwase S, Xiang B, Ghosh S, Ren T, Lewis PW, Cochrane JC, Allis CD, Picketts DJ, Patel DJ, Li H, Shi Y (2011) ATRX ADD domain links an atypical histone methylation recognition mechanism to human mental-retardation syndrome. Nat Struct Mol Biol 18(7):769–776. https://doi.org/10.1038/nsmb.2062
Jasencakova Z, Scharf AND, Ask K, Corpet A, Imhof A, Almouzni G, Groth A (2010) Replication stress interferes with histone recycling and predeposition marking of new histones. Mol Cell 37(5):736–743. https://doi.org/10.1016/j.molcel.2010.01.033
Jiang J, Lv W, Ye X, Wang L, Zhang M, Yang H, Okuka M, Zhou C, Zhang X, Liu L, Li J (2013) Zscan4 promotes genomic stability during reprogramming and dramatically improves the quality of iPS cells as demonstrated by tetraploid complementation. Cell Res 23(1):92–106. https://doi.org/10.1038/cr.2012.157
Jiao Y, Shi C, Edil BH, de Wilde RF, Klimstra DS, Maitra A, Schulick RD, Tang LH, Wolfgang CL, Choti MA, Velculescu VE, Diaz l, Vogelstein B, Kinzler KW, Hruban RH, Papadopoulos N (2011) DAXX/ATRX, MEN1, and mTOR pathway genes are frequently altered in pancreatic neuroendocrine tumors. Science 331(6021):1199–1203. https://doi.org/10.1126/science.1200609
Jung YH, Sauria MEG, Lyu X, Cheema MS, Ausio J, Taylor J, Corces VG (2017) Chromatin states in mouse sperm correlate with embryonic and adult regulatory landscapes. Cell Rep 18(6):1366–1382. https://doi.org/10.1016/j.celrep.2017.01.034
Ke Y, Xu Y, Chen X et al (2017) 3D chromatin structures of mature gametes and structural reprogramming during mammalian embryogenesis. Cell 170:367–381.e20. https://doi.org/10.1016/j.cell.2017.06.029
Keefe DL (2016) Telomeres, reproductive aging, and genomic instability during early development. Reprod Sci Thousand Oaks Calif 23(12):1612–1615. https://doi.org/10.1177/1933719116676397
Keefe DL, Marquard K, Liu L (2006) The telomere theory of reproductive senescence in women. Curr Opin Obstet Gynecol 18(3):280–285. https://doi.org/10.1097/01.gco.0000193019.05686.49
Khoueiry R, Sohni A, Thienpont B, Luo X, Velde JV, Bartoccetti M, Boeckx B, Zwijsen A, Rao A, Lambrechts D, Koh KP (2017) Lineage-specific functions of TET1 in the postimplantation mouse embryo. Nat Genet 49(7):1061–1072. https://doi.org/10.1038/ng.3868
Kim J, Zhao H, Dan J, Kim S, Hardikar S, Hollowell D, Lin K, Lu Y, Takata Y, Shen J, Chen T (2016) Maternal Setdb1 is required for meiotic progression and preimplantation development in mouse. PLoS Genet 12(4):e1005970. https://doi.org/10.1371/journal.pgen.1005970
Kota SK, Feil R (2010) Epigenetic transitions in germ cell development and meiosis. Dev Cell 19(5):675–686. https://doi.org/10.1016/j.devcel.2010.10.009
Law MJ, Lower KM, Voon HPJ, Hughes JR, Garrick D, Viprakasit V, Mitson M, de Gobbi M, Marra M, Morris A, Abbott A, Wilder SP, Taylor S, Santos GM, Cross J, Ayyub H, Jones S, Ragoussis J, Rhodes D, Dunham I, Higgs DR, Gibbons RJ (2010) ATR-X syndrome protein targets tandem repeats and influences allele-specific expression in a size-dependent manner. Cell 143(3):367–378. https://doi.org/10.1016/j.cell.2010.09.023
Lee HW, Blasco MA, Gottlieb GJ et al (1998) Essential role of mouse telomerase in highly proliferative organs. Nature 392(6676):569–574. https://doi.org/10.1038/33345
Lemmers RJLF, Tawil R, Petek LM, Balog J, Block GJ, Santen GWE, Amell AM, van der Vliet PJ, Almomani R, Straasheijm KR, Krom YD, Klooster R, Sun Y, den Dunnen JT, Helmer Q, Donlin-Smith CM, Padberg GW, van Engelen BGM, de Greef JC, Aartsma-Rus AM, Frants RR, de Visser M, Desnuelle C, Sacconi S, Filippova GN, Bakker B, Bamshad MJ, Tapscott SJ, Miller DG, van der Maarel SM (2012) Digenic inheritance of an SMCHD1 mutation and an FSHD-permissive D4Z4 allele causes facioscapulohumeral muscular dystrophy type 2. Nat Genet 44(12):1370–1374. https://doi.org/10.1038/ng.2454
Lewis PW, Elsaesser SJ, Noh K-M, Stadler SC, Allis CD (2010) Daxx is an H3.3-specific histone chaperone and cooperates with ATRX in replication-independent chromatin assembly at telomeres. Proc Natl Acad Sci U S A 107(32):14075–14080. https://doi.org/10.1073/pnas.1008850107
Liu H, Kim J-M, Aoki F (2004a) Regulation of histone H3 lysine 9 methylation in oocytes and early pre-implantation embryos. Dev Camb Engl 131(10):2269–2280. https://doi.org/10.1242/dev.01116
Liu L (2017) Linking telomere regulation to stem cell pluripotency. Trends Genet TIG 33(1):16–33. https://doi.org/10.1016/j.tig.2016.10.007
Liu L, Bailey SM, Okuka M, Muñoz P, Li C, Zhou L, Wu C, Czerwiec E, Sandler L, Seyfang A, Blasco MA, Keefe DL (2007) Telomere lengthening early in development. Nat Cell Biol 9(12):1436–1441. https://doi.org/10.1038/ncb1664
Liu L, Blasco M, Trimarchi J, Keefe D (2002) An essential role for functional telomeres in mouse germ cells during fertilization and early development. Dev Biol 249(1):74–84. https://doi.org/10.1006/dbio.2002.0735
Liu L, Franco S, Spyropoulos B, Moens PB, Blasco MA, Keefe DL (2004b) Irregular telomeres impair meiotic synapsis and recombination in mice. Proc Natl Acad Sci U S A 101(17):6496–6501. https://doi.org/10.1073/pnas.0400755101
López de Silanes I, Graña O, De Bonis ML et al (2014) Identification of TERRA locus unveils a telomere protection role through association to nearly all chromosomes. Nat Commun 5:4723. https://doi.org/10.1038/ncomms5723
Lovejoy CA, Li W, Reisenweber S, Thongthip S, Bruno J, de Lange T, de S, Petrini JHJ, Sung PA, Jasin M, Rosenbluh J, Zwang Y, Weir BA, Hatton C, Ivanova E, Macconaill L, Hanna M, Hahn WC, Lue NF, Reddel RR, Jiao Y, Kinzler K, Vogelstein B, Papadopoulos N, Meeker AK, for the ALT Starr Cancer Consortium (2012) Loss of ATRX, genome instability, and an altered DNA damage response are hallmarks of the alternative lengthening of telomeres pathway. PLoS Genet 8(7):e1002772. https://doi.org/10.1371/journal.pgen.1002772
Lu F, Liu Y, Inoue A, Suzuki T, Zhao K, Zhang Y (2016) Establishing chromatin regulatory landscape during mouse preimplantation development. Cell 165(6):1375–1388. https://doi.org/10.1016/j.cell.2016.05.050
Lu F, Liu Y, Jiang L, Yamaguchi S, Zhang Y (2014) Role of Tet proteins in enhancer activity and telomere elongation. Genes Dev 28(19):2103–2119. https://doi.org/10.1101/gad.248005.114
Macfarlan TS, Gifford WD, Driscoll S, Lettieri K, Rowe HM, Bonanomi D, Firth A, Singer O, Trono D, Pfaff SL (2012) Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 487:57–63. https://doi.org/10.1038/nature11244
Maksakova IA, Thompson PJ, Goyal P, Jones SJM, Singh PB, Karimi MM, Lorincz MC (2013) Distinct roles of KAP1, HP1 and G9a/GLP in silencing of the two-cell-specific retrotransposon MERVL in mouse ES cells. Epigenetics Chromatin 6(1):15. https://doi.org/10.1186/1756-8935-6-15
Marion RM, Strati K, Li H, Tejera A, Schoeftner S, Ortega S, Serrano M, Blasco MA (2009) Telomeres acquire embryonic stem cell characteristics in induced pluripotent stem cells. Cell Stem Cell 4(2):141–154. https://doi.org/10.1016/j.stem.2008.12.010
Marzec P, Armenise C, Pérot G, Roumelioti FM, Basyuk E, Gagos S, Chibon F, Déjardin J (2015) Nuclear-receptor-mediated telomere insertion leads to genome instability in ALT cancers. Cell 160(5):913–927. https://doi.org/10.1016/j.cell.2015.01.044
Matoba S, Liu Y, Lu F, Iwabuchi KA, Shen L, Inoue A, Zhang Y (2014) Embryonic development following somatic cell nuclear transfer impeded by persisting histone methylation. Cell 159(4):884–895. https://doi.org/10.1016/j.cell.2014.09.055
Messerschmidt DM, Knowles BB, Solter D (2014) DNA methylation dynamics during epigenetic reprogramming in the germline and preimplantation embryos. Genes Dev 28(8):812–828. https://doi.org/10.1101/gad.234294.113
Michaelson JS, Bader D, Kuo F, Kozak C, Leder P (1999) Loss of Daxx, a promiscuously interacting protein, results in extensive apoptosis in early mouse development. Genes Dev 13(15):1918–1923. https://doi.org/10.1101/gad.13.15.1918
Napier CE, Huschtscha LI, Harvey A, Bower K, Noble JR, Hendrickson EA, Reddel RR (2015) ATRX represses alternative lengthening of telomeres. Oncotarget 6(18):16543–16558. https://doi.org/10.18632/oncotarget.3846
Nestorov P, Hotz H-R, Liu Z, Peters AHFM (2015) Dynamic expression of chromatin modifiers during developmental transitions in mouse preimplantation embryos. Sci Rep 5(1):14347. https://doi.org/10.1038/srep14347
Neumann AA, Watson CM, Noble JR, Pickett HA, Tam PPL, Reddel RR (2013) Alternative lengthening of telomeres in normal mammalian somatic cells. Genes Dev 27(1):18–23. https://doi.org/10.1101/gad.205062.112
Nguyen DT, Voon HPJ, Xella B et al (2017) The chromatin remodelling factor ATRX suppresses R-loops in transcribed telomeric repeats. EMBO Rep 18(6):914–928. https://doi.org/10.15252/embr.201643078
Nishibuchi G, Déjardin J (2017) The molecular basis of the organization of repetitive DNA-containing constitutive heterochromatin in mammals. Chromosome Res Int J Mol Supramol Evol Asp Chromosome Biol 25(1):77–87. https://doi.org/10.1007/s10577-016-9547-3
O’Sullivan RJ, Almouzni G (2014) Assembly of telomeric chromatin to create ALTernative endings. Trends Cell Biol 24(11):675–685. https://doi.org/10.1016/j.tcb.2014.07.007
O’Sullivan RJ, Arnoult N, Lackner DH et al (2014) Rapid induction of alternative lengthening of telomeres by depletion of the histone chaperone ASF1. Nat Struct Mol Biol 21(2):167–174. https://doi.org/10.1038/nsmb.2754
Palm W, de Lange T (2008) How shelterin protects mammalian telomeres. Annu Rev Genet 42(1):301–334. https://doi.org/10.1146/annurev.genet.41.110306.130350
Park S-J, Komata M, Inoue F, Yamada K, Nakai K, Ohsugi M, Shirahige K (2013) Inferring the choreography of parental genomes during fertilization from ultralarge-scale whole-transcriptome analysis. Genes Dev 27(24):2736–2748. https://doi.org/10.1101/gad.227926.113
Peaston AE, Evsikov AV, Graber JH, de Vries WN, Holbrook AE, Solter D, Knowles BB (2004) Retrotransposons regulate host genes in mouse oocytes and preimplantation embryos. Dev Cell 7(4):597–606. https://doi.org/10.1016/j.devcel.2004.09.004
Pech MF, Garbuzov A, Hasegawa K, Sukhwani M, Zhang RJ, Benayoun BA, Brockman SA, Lin S, Brunet A, Orwig KE, Artandi SE (2015) High telomerase is a hallmark of undifferentiated spermatogonia and is required for maintenance of male germline stem cells. Genes Dev 29(23):2420–2434. https://doi.org/10.1101/gad.271783.115
Perrem K, Colgin LM, Neumann AA, Yeager TR, Reddel RR (2001) Coexistence of alternative lengthening of telomeres and telomerase in hTERT-transfected GM847 cells. Mol Cell Biol 21(12):3862–3875. https://doi.org/10.1128/MCB.21.12.3862-3875.2001
Peters AH, O’Carroll D, Scherthan H et al (2001) Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 107(3):323–337. https://doi.org/10.1016/S0092-8674(01)00542-6
Posfai E, Kunzmann R, Brochard V, Salvaing J, Cabuy E, Roloff TC, Liu Z, Tardat M, van Lohuizen M, Vidal M, Beaujean N, Peters AHFM (2012) Polycomb function during oogenesis is required for mouse embryonic development. Genes Dev 26(9):920–932. https://doi.org/10.1101/gad.188094.112
Probst AV, Okamoto I, Casanova M, el Marjou F, le Baccon P, Almouzni G (2010) A strand-specific burst in transcription of pericentric satellites is required for chromocenter formation and early mouse development. Dev Cell 19(4):625–638. https://doi.org/10.1016/j.devcel.2010.09.002
Puschendorf M, Terranova R, Boutsma E, Mao X, Isono K, Brykczynska U, Kolb C, Otte AP, Koseki H, Orkin SH, van Lohuizen M, Peters AHFM (2008) PRC1 and Suv39h specify parental asymmetry at constitutive heterochromatin in early mouse embryos. Nat Genet 40(4):411–420. https://doi.org/10.1038/ng.99
Ramamoorthy M, Smith S (2015) Loss of ATRX suppresses resolution of telomere cohesion to control recombination in ALT cancer cells. Cancer Cell 28(3):357–369. https://doi.org/10.1016/j.ccell.2015.08.003
Reig-Viader R, Garcia-Caldés M, Ruiz-Herrera A (2016) Telomere homeostasis in mammalian germ cells: a review. Chromosoma 125(2):337–351. https://doi.org/10.1007/s00412-015-0555-4
Reig-Viader R, Vila-Cejudo M, Vitelli V, Buscà R, Sabaté M, Giulotto E, Caldés MG, Ruiz-Herrera A (2014) Telomeric repeat-containing RNA (TERRA) and telomerase are components of telomeres during mammalian gametogenesis. Biol Reprod 90(5):103. https://doi.org/10.1095/biolreprod.113.116954
Rivera T, Haggblom C, Cosconati S, Karlseder J (2016) A balance between elongation and trimming regulates telomere stability in stem cells. Nat Struct Mol Biol 24(1):30–39. https://doi.org/10.1038/nsmb.3335
Royo H, Stadler MB, Peters AHFM (2016) Alternative computational analysis shows no evidence for nucleosome enrichment at repetitive sequences in mammalian spermatozoa. Dev Cell 37(1):98–104. https://doi.org/10.1016/j.devcel.2016.03.010
Sadic D, Schmidt K, Groh S, et al (2015) Atrx promotes heterochromatin formation at retrotransposons. EMBO rep. Doi: https://doi.org/10.15252/embr.201439937
Saksouk N, Barth TK, Ziegler-Birling C, Olova N, Nowak A, Rey E, Mateos-Langerak J, Urbach S, Reik W, Torres-Padilla ME, Imhof A, Déjardin J, Simboeck E (2014) Redundant mechanisms to form silent chromatin at pericentromeric regions rely on BEND3 and DNA methylation. Mol Cell 56(4):580–594. https://doi.org/10.1016/j.molcel.2014.10.001
Samans B, Yang Y, Krebs S, Sarode GV, Blum H, Reichenbach M, Wolf E, Steger K, Dansranjavin T, Schagdarsurengin U (2014) Uniformity of nucleosome preservation pattern in mammalian sperm and its connection to repetitive DNA elements. Dev Cell 30(1):23–35. https://doi.org/10.1016/j.devcel.2014.05.023
Santos F, Peters AH, Otte AP, Reik W, Dean W (2005) Dynamic chromatin modifications characterise the first cell cycle in mouse embryos. Dev Biol 280(1):225–236. https://doi.org/10.1016/j.ydbio.2005.01.025
Sarek G, Vannier J-B, Panier S, Petrini JHJ, Boulton SJ (2015) TRF2 recruits RTEL1 to telomeres in S phase to promote t-loop unwinding. Mol Cell 57(4):622–635. https://doi.org/10.1016/j.molcel.2014.12.024
Schaetzlein S, Lucas-Hahn A, Lemme E, Kues WA, Dorsch M, Manns MP, Niemann H, Rudolph KL (2004) Telomere length is reset during early mammalian embryogenesis. Proc Natl Acad Sci U S A 101(21):8034–8038. https://doi.org/10.1073/pnas.0402400101
Scherthan H (2007) Telomere attachment and clustering during meiosis. Cell Mol Life Sci CMLS 64(2):117–124. https://doi.org/10.1007/s00018-006-6463-2
Schoeftner S, Blasco MA (2010) Chromatin regulation and non-coding RNAs at mammalian telomeres. Semin Cell Dev Biol 21(2):186–193. https://doi.org/10.1016/j.semcdb.2009.09.015
Schoeftner S, Blasco MA (2008) Developmentally regulated transcription of mammalian telomeres by DNA-dependent RNA polymerase II. Nat Cell Biol 10(2):228–236. https://doi.org/10.1038/ncb1685
Sepulveda-Rincon LP, Solanas EDL, Serrano-Revuelta E et al (2016) Early epigenetic reprogramming in fertilized, cloned, and parthenogenetic embryos. Theriogenology 86(1):91–98. https://doi.org/10.1016/j.theriogenology.2016.04.022
Shibuya H, Hernández-Hernández A, Morimoto A, Negishi L, Höög C, Watanabe Y (2015) MAJIN links telomeric DNA to the nuclear membrane by exchanging telomere cap. Cell 163(5):1252–1266. https://doi.org/10.1016/j.cell.2015.10.030
Shibuya H, Watanabe Y (2014) The meiosis-specific modification of mammalian telomeres. Cell Cycle Georget Tex 13(13):2024–2028. https://doi.org/10.4161/cc.29350
Sridharan R, Gonzales-Cope M, Chronis C, Bonora G, McKee R, Huang C, Patel S, Lopez D, Mishra N, Pellegrini M, Carey M, Garcia BA, Plath K (2013) Proteomic and genomic approaches reveal critical functions of H3K9 methylation and heterochromatin protein-1γ in reprogramming to pluripotency. Nat Cell Biol 15(7):872–882. https://doi.org/10.1038/ncb2768
Svoboda P, Franke V, Schultz RM (2015) Sculpting the transcriptome during the oocyte-to-embryo transition in mouse. Curr Top Dev Biol 113:305–349. https://doi.org/10.1016/bs.ctdb.2015.06.004
Takada Y, Naruse C, Costa Y, Shirakawa T, Tachibana M, Sharif J, Kezuka-Shiotani F, Kakiuchi D, Masumoto H, Shinkai Y, Ohbo K, Peters AHFM, Turner JMA, Asano M, Koseki H (2011) HP1γ links histone methylation marks to meiotic synapsis in mice. Dev Camb Engl 138(19):4207–4217. https://doi.org/10.1242/dev.064444
Takahashi K, Yamanaka S (2016) A decade of transcription factor-mediated reprogramming to pluripotency. Nat Rev Mol Cell Biol 17(3):183–193. https://doi.org/10.1038/nrm.2016.8
Tanemura K, Ogura A, Cheong C, Gotoh H, Matsumoto K, Sato E, Hayashi Y, Lee HW, Kondo T (2005) Dynamic rearrangement of telomeres during spermatogenesis in mice. Dev Biol 281(2):196–207. https://doi.org/10.1016/j.ydbio.2005.02.025
Teperek M, Simeone A, Gaggioli V, Miyamoto K, Allen GE, Erkek S, Kwon T, Marcotte EM, Zegerman P, Bradshaw CR, Peters AHFM, Gurdon JB, Jullien J (2016) Sperm is epigenetically programmed to regulate gene transcription in embryos. Genome Res 26(8):1034–1046. https://doi.org/10.1101/gr.201541.115
Timashev LA, Babcock H, Zhuang X, de Lange T (2017) The DDR at telomeres lacking intact shelterin does not require substantial chromatin decompaction. Genes Dev 31(6):578–589. https://doi.org/10.1101/gad.294108.116
Udugama M, Chang FTM, Chan FL et al (2015) Histone variant H3.3 provides the heterochromatic H3 lysine 9 tri-methylation mark at telomeres. Nucleic acids res. https://doi.org/10.1093/nar/gkv847
van Steensel B, Smogorzewska A, de Lange T (1998) TRF2 protects human telomeres from end-to-end fusions. Cell 92(3):401–413. https://doi.org/10.1016/S0092-8674(00)80932-0
Vancevska A, Douglass KM, Pfeiffer V, Manley S, Lingner J (2017) The telomeric DNA damage response occurs in the absence of chromatin decompaction. Genes Dev 31(6):567–577. https://doi.org/10.1101/gad.294082.116
Vannier J-B, Pavicic-Kaltenbrunner V, Petalcorin MIR, Ding H, Boulton SJ (2012) RTEL1 dismantles T loops and counteracts telomeric G4-DNA to maintain telomere integrity. Cell 149(4):795–806. https://doi.org/10.1016/j.cell.2012.03.030
Vassena R, Boué S, González-Roca E et al (2011) Waves of early transcriptional activation and pluripotency program initiation during human preimplantation development. Dev Camb Engl 138(17):3699–3709. https://doi.org/10.1242/dev.064741
Voon HPJ, Hughes JR, Rode C, de la Rosa-Velázquez IA, Jenuwein T, Feil R, Higgs DR, Gibbons RJ (2015) ATRX plays a key role in maintaining silencing at interstitial heterochromatic loci and imprinted genes. Cell Rep 11(3):405–418. https://doi.org/10.1016/j.celrep.2015.03.036
Watson LA, Goldberg H, Bérubé NG (2015) Emerging roles of ATRX in cancer. Epigenomics 7(8):1365–1378. https://doi.org/10.2217/epi.15.82
Whiddon JL, Langford AT, Wong C-J, Zhong JW, Tapscott SJ (2017) Conservation and innovation in the DUX4-family gene network. Nat Genet 49(6):935–940. https://doi.org/10.1038/ng.3846
Wong LH, McGhie JD, Sim M, Anderson MA, Ahn S, Hannan RD, George AJ, Morgan KA, Mann JR, Choo KHA (2010) ATRX interacts with H3.3 in maintaining telomere structural integrity in pluripotent embryonic stem cells. Genome Res 20(3):351–360. https://doi.org/10.1101/gr.101477.109
Wright WE, Piatyszek MA, Rainey WE, Byrd W, Shay JW (1996) Telomerase activity in human germline and embryonic tissues and cells. Dev Genet 18(2):173–179. https://doi.org/10.1002/(SICI)1520-6408(1996)18:2<173::AID-DVG10>3.0.CO;2-3
Wu J, Huang B, Chen H, Yin Q, Liu Y, Xiang Y, Zhang B, Liu B, Wang Q, Xia W, Li W, Li Y, Ma J, Peng X, Zheng H, Ming J, Zhang W, Zhang J, Tian G, Xu F, Chang Z, Na J, Yang X, Xie W (2016) The landscape of accessible chromatin in mammalian preimplantation embryos. Nature 534(7609):652–657. https://doi.org/10.1038/nature18606
Yang J, Guo R, Wang H, Ye X, Zhou Z, Dan J, Wang H, Gong P, Deng W, Yin Y, Mao SQ, Wang L, Ding J, Li J, Keefe DL, Dawlaty MM, Wang J, Xu GL, Liu L (2016) Tet enzymes regulate telomere maintenance and chromosomal stability of mouse ESCs. Cell Rep 15(8):1809–1821. https://doi.org/10.1016/j.celrep.2016.04.058
Yang Q, Zhao F, Dai S, Zhang N, Zhao W, Bai R, Sun Y (2015) Sperm telomere length is positively associated with the quality of early embryonic development. Hum Reprod Oxf Engl 30(8):1876–1881. https://doi.org/10.1093/humrep/dev144
Yeo S, Lee K-K, Han Y-M, Kang Y-K (2005) Methylation changes of lysine 9 of histone H3 during preimplantation mouse development. Mol Cells 20(3):423–428
Zalenskaya IA, Bradbury EM, Zalensky AO (2000) Chromatin structure of telomere domain in human sperm. Biochem Biophys Res Commun 279(1):213–218. https://doi.org/10.1006/bbrc.2000.3917
Zalzman M, Falco G, Sharova LV, Nishiyama A, Thomas M, Lee SL, Stagg CA, Hoang HG, Yang HT, Indig FE, Wersto RP, Ko MSH (2010) Zscan4 regulates telomere elongation and genomic stability in ES cells. Nature 464(7290):858–863. https://doi.org/10.1038/nature08882
Zeng S, Liu L, Sun Y, Lu G, Lin G (2017) Role of telomeric repeat-containing RNA in telomeric chromatin remodeling during the early expansion of human embryonic stem cells. FASEB J Off Publ Fed Am Soc Exp Biol 31(11):4783–4795. https://doi.org/10.1096/fj.201600939RR
Zhong Z-H, Jiang W-Q, Cesare AJ, Neumann AA, Wadhwa R, Reddel RR (2007) Disruption of telomere maintenance by depletion of the MRE11/RAD50/NBS1 complex in cells that use alternative lengthening of telomeres. J Biol Chem 282(40):29314–29322. https://doi.org/10.1074/jbc.M701413200
Funding
Mathieu Tardat’s research on telomere chromatin is funded by a H2020 Marie Skłodowska-Curie Actions (LS150091). Research in the Déjardin’s lab is funded by a H2020 European Research Council Grant (CoG#724835), by the Association pour la Recherche contre le cancer labelisation, and by the Institut National du Cancer.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interests
The Déjardin’s lab is also funded by the Merck Sharp & Dome Avenir programme (#Gnostic).
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Tardat, M., Déjardin, J. Telomere chromatin establishment and its maintenance during mammalian development. Chromosoma 127, 3–18 (2018). https://doi.org/10.1007/s00412-017-0656-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-017-0656-3