Abstract
Introduction
Neoadjuvant chemoradiation therapy has been shown to improve the outcome in patients with rectal cancer and is generally accepted as standard care; however, only selected patients would benefit from this treatment. We aimed to identify predictors of response to neoadjuvant chemoradiation therapy in colorectal cancer using formalin-fixed paraffin-embedded (FFPE) tissues as source of genetic materials and microarray analysis as investigation tool.
Methods
After optimization of RNA extraction methods from FFPE, microarray analysis was carried out on total RNA extracted from 12 pre-treatment FFPE rectal tissues using Megaplex pool A. Microarray data were analysed using an artificial neural network algorithm. Statistical analysis and correlation with clinicopathological data was performed using SPSS software.
Results
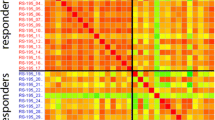
A distinct miRNA expression signature predictive of response to neoadjuvant CRT in 12 FFPE pre-treatment rectal cancer tissue samples was identified. These signatures consisted of three miRNA transcripts (miR-16, miR-590-5p and miR-153) to predict complete vs. incomplete response and two miRNA transcript (miR-519c-3p and miR-561) to predict good vs. poor response with a median accuracy of 100 %.
Conclusion
Using microarray analysis of pretreatment FFPE rectal cancer tissues, we identified for the first time a group of miRNA predictors of response to neoadjuvant CRT. This, indeed, can lead to a significant improvement in patient selection criteria and personalized rectal cancer management.









Similar content being viewed by others
References
Julien LA, Thorson AG (2010) Current neoadjuvant strategies in rectal cancer. J Surg Oncol 101(4):321–326
Galanis E, Alberts SR, O’Connell MJ (2000) New adjuvant therapy for colon cancer: justified hope or commercial hype. Surg Oncol Clin N Am 9(4):813–823
Kapiteijn E, Marijnen CAM, Nagtegaal ID, Putter H, Steup WH, Wiggers T, Rutten HJT, Pahlman L, Glimelius B, van Krieken J, Leer JWH, van de Velde CJH, Dutch Colorectal Canc (2001) Preoperative radiotherapy combined with total mesorectal excision for resectable rectal cancer. N Engl J Med 345(9):638–646
Sauer R, Becker H, Hohenberger W, Rodel C, Wittekind C, Fietkau R, Martus P, Tschmelitsch J, Hager E, Hess CF, Karstens JH, Liersch T, Schmidberger H, Raab R, German Rectal Canc Study (2004) Preoperative versus postoperative chemoradiotherapy for rectal cancer. N Engl J Med 351(17):1731–1740
Mandard AM, Dalibard F, Mandard JC, Marnay J, Henry-Amar M, Pefiot JF, Roussel A, Jacob JH, Segol P, Samama G, Ollivier JM, Bonvalot S, Gignoux M (1994) Pathologic assessment of tumor regression after preoperative chemoradiotherapy of esophageal carcinoma. Clinicopathologic correlations. Cancer 73(11):2680–2688
Dworak O, Keilholz L, Hoffmann A (1997) Pathological features of rectal cancer after preoperative radiochemotherapy. Int J Color Dis 12(1):19–23
Rodel C, Martus P, Papadoupolos T, Fuzesi L, Klimpfinger M, Fietkau R, Liersch T, Hohenberger W, Raab R, Sauer R, Wittekind C (2005) Prognostic significance of tumor regression after preoperative chemoradiotherapy for rectal cancer. J Clin Oncol 23(34):8688–8696
Habr-Gama A, Perez RO, Nadalin W, Sabbaga J, Ribeiro U, Sousa A, Campos FG, Kiss DR, Gama-Rodrigues J (2004) Operative versus nonoperative treatment for stage 0 distal rectal cancer following chemoradiation therapy—long-term results. Ann Surg 240(4):711–717
Habr-Gama A, Perez RO, Nadalin W, Nahas SC, Ribeiro U, Sousa AHS, Campos FG, Kiss DR, Gama-Rodriguez J (2005) Long-term results of preoperative chemoradiation for distal rectal cancer correlation between final stage and survival. J Gastrointest Surg 9(1):90–99
Habr-Gama A (2006) Assessment and management of the complete clinical response of rectal cancer to chemoradiotherapy. Color Dis 8:21–24
Chan JA, Krichevsky AM, Kosik KS (2005) MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res 65(14):6029–6033
Xi YG, Nakajima G, Gavin E, Morris CG, Kudo K, Hayashi KH, Ju JF (2007) Systematic analysis of microRNA expression of RNA extracted from fresh frozen and formalin-fixed paraffin-embedded samples. RNA Pub RNA Soc 13:1668–1674
Lassmann S, Kreutz C, Schoepflin A, Hopt U, Timmer J, Werner M (2009) A novel approach for reliable microarray analysis of microdissected tumor cells from formalin-fixed and paraffin-embedded colorectal cancer resection specimens. J Mol Med (Berlin) 87(2):211–224
Hui ABY, Shi W, Boutros PC, Miller N, Pintilie M, Fyles T, McCready D, Wong D, Gerster K, Jurisica I, Penn LZ, Liu FF (2009) Robust global micro-RNA profiling with formalin-fixed paraffin-embedded breast cancer tissues. Lab Investig 89(5):597–606
Doleshal M, Magotra AA, Choudhury B, Cannon BD, Labourier E, Szafranska AE (2008) Evaluation and validation of total RNA extraction methods for microRNA expression analyses in formalin-fixed, paraffin-embedded tissues. J Mol Diagn 10(3):203–211
Smith FM, Reynolds JV, Miller N, Stephens RB, Kennedy MJ (2006) Pathological and molecular predictors of the response of rectal cancer to neoadjuvant radiochemotherapy. Eur J Surg Oncol 32(1):55–64
O’Neill BDP, Brown G, Heald RJ, Cunningham D, Tait DM (2007) Non-operative treatment after neoadjuvant chemoradiotherapy for rectal cancer. Lancet Oncol 8(7):625–633
Cummins JM, He YP, Leary RJ, Pagliarini R, Diaz LA, Sjoblom T, Barad O, Bentwich Z, Szafranska AE, Labourier E, Raymond CK, Roberts BS, Juhl H, Kinzler KW, Vogelstein B, Velculescu VE (2006) The colorectal microRNAome. Proc Natl Acad Sci U S A 103(10):3687–3692
Negrini M, Calin GA (2008) Breast cancer metastasis: a microRNA story. Breast Canc Res 10(2)
Ma L, Weinberg RA (2008) MicroRNAs in malignant progression. Cell Cycle 7(5):570–572
Hiroki E, Akahira J, Suzuki F, Nagase S, Ito K, Suzuki T, Sasano H, Yaegashi N (2010) Changes in microRNA expression levels correlate with clinicopathological features and prognoses in endometrial serous adenocarcinomas. Cancer Science 101(1):241–249
Lewis F, Maughan NJ, Smith V, Hillan K, Quirke P (2001) Unlocking the archive—gene expression in paraffin-embedded tissue. J Pathol 195(1):66–71
Bishop C (1995) Neural networks for pattern recognition. Oxford University Press, Oxford
Lancashire L, Schmid O, Shah H, Ball G (2005) Classification of bacterial species from proteomic data using combinatorial approaches incorporating artificial neural networks, cluster analysis and principal components analysis. Bioinformatics 21(10):2191–2199
Michael MZ, O’Connor SM, Pellekaan NGV, Young GP, James RJ (2003) Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Canc Res 1(12):882–891
Motoyama K, Inoue H, Takatsuno Y, Tanaka F, Mimori K, Uetake H, Sugihara K, Mori M (2009) Over- and under-expressed microRNAs in human colorectal cancer. Int J Oncol 34(4):1069–1075
Yang LD, Belaguli N, Berger DH (2009) MicroRNA and colorectal cancer. World J Surg 33(4):638–646
Arndt GM, Dossey L, Cullen LM, Lai A, Druker R, Eisbacher M, Zhang CY, Tran N, Fan HT, Retzlaff K, Bittner A, Raponi M (2009) Characterization of global microRNA expression reveals oncogenic potential of miR-145 in metastatic colorectal cancer. BMC Cancer 9:374
Slaby O, Hrstka R, Dubska L, Svoboda M, Ruckova E, Ovesna J, Vyzula R (2008) Significance of microRNA-21 in colorectal cancer pathogenesis. FEBS J 275:465–465
Ng EKO, Chong WWS, Jin H, Lam EKY, Shin VY, Yu J, Poon TCW, Ng SSM, Sung JJY (2009) Differential expression of microRNAs in plasma of patients with colorectal cancer: a potential marker for colorectal cancer screening. Gut 58(10):1375–1381
Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M, Stephens RM, Okamoto A, Yokota J, Tanaka T, Colin GA, Liu CG, Croce CM, Harris CC (2006) Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Canc Cell 9(3):189–198
Yu SL, Chen HY, Chang GC, Chen CY, Chen HW, Singh S, Cheng CL, Yu CJ, Lee YC, Chen HS, Su TJ, Chiang CC, Li HN, Hong QS, Su HY, Chen CC, Chen WJ, Liu CC, Chan WK, Chen WJ, Li KC, Chen JJW, Yang PC (2008) MicroRNA signature predicts survival and relapse in lung cancer. Canc Cell 13(1):48–57
Schetter AJ, Leung SY, Sohn JJ, Zanetti KA, Bowman ED, Yanaihara N, Yuen ST, Chan TL, Kwong DLW, Au GKH, Liu CG, Calin GA, Croce CM, Harris CC (2008) MicroRNA expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. J Am Med Assoc 299(4):425–436
Budhu A, Jia HL, Forgues M, Liu CG, Goldsteir D, Lam A, Zanetti KA, Ye QH, Qin LY, Croce CM, Tang ZY, Wang XW (2008) Identification of metastasis-related microRNAs in hepatocellular carcinoma. Hepatology 47(3):897–907
Guo Y, Chen ZL, Zhang L, Zhou F, Shi SS, Feng XL, Li BZ, Meng X, Ma X, Luo MY, Shao K, Li N, Qiu B, Mitchelson K, Cheng J, He J (2008) Distinctive microRNA profiles relating to patient survival in esophageal squamous cell carcinoma. Cancer Res 68(1):26–33
Farragher SM, Tanney A, Kennedy RD, Harkin DP (2008) RNA expression analysis from formalin fixed paraffin embedded tissues. Histochem Cell Biol 130(3):435–445
Vincek V, Nassiri M, Nadji M, Morales AR (2003) A tissue fixative that protects macromolecules (DNA, RNA, and protein) and histomorphology in clinical samples. Lab Investig 83(10):1427–1435
Chen X, Guo X, Zhang H, Xiang Y, Chen J, Yin Y, Cai X, Wang K, Wang G, Ba Y, Zhu L, Wang J, Yang R, Zhang Y, Ren Z, Zen K, Zhang J, Zhang CY (2009) Role of miR-143 targeting KRAS in colorectal tumorigenesis. Oncogene 28(10):1385–1392
Asangani IA, Rasheed SAK, Nikolova DA, Leupold JH, Colburn NH, Post S, Allgayer H (2008) MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 27(15):2128–2136
Slaby O, Svoboda M, Fabian P, Smerdova T, Knoflickova D, Bednarikova M, Nenutil R, Vyzula R (2007) Altered expression of miR-21, miR-31, miR-143 and miR-145 is related to clinicopathologic features of colorectal cancer. Oncology 72(5–6):397–402
Valentini V, Coco C, Picciocchi A, Morganti AG, Trodella L, Ciabattoni A, Cellini F, Barbaro B, Cogliandolo S, Nuzzo G, Doglietto GB, Ambesi-Impiombato F, Cosimelli M (2002) Does downstaging predict improved outcome after preoperative chemoradiation for extraperitoneal locally advanced rectal cancer? A long-term analysis of 165 patients. Int J Radiat Oncol Biol Phys 53(3):664–674
Reerink O, Karrenbeld A, Plukker JTM, Verschueren RCJ, Szabo BG, Sluiter WJ, Hospers GAP, Mulder NH (2004) Molecular prognostic factors in locally irresectable rectal cancer treated preoperatively by chemo-radiotherapy. Anticancer Res 24(2C):1217–1221
Heriot AG, Tekkis PP, Fazio VW, Neary P, Lavery IC (2005) Adjuvant radiotherapy is associated with increased sexual dysfunction in male patients undergoing resection for rectal cancer—a predictive model. Ann Surg 242(4):502–511
Birgisson H, Pahlman L, Gunnarsson U, Glimelius B (2005) Adverse effects of preoperative radiation therapy for rectal cancer: long-term follow-up of the Swedish rectal cancer trial. J Clin Oncol 23(34):8697–8705
Hummel R, Hussey DJ, Haier J (2010) MicroRNAs: predictors and modifiers of chemo- and radiotherapy in different tumour types. Eur J Cancer 46(2):298–311
Chang KH, Mestdagh P, Vandesompele J, Kerin MJ, Miller N (2010) MicroRNA expression profiling to identify and validate reference genes for relative quantification in colorectal cancer. BMC Cancer 10:173
Davoren PA, McNeill RE, Lowery AJ, Kerin MJ, Miller N (2008) Identification of suitable endogenous control genes for microRNA gene expression analysis in human breast cancer. BMC Mol Biol 9:76
Earle JSL, Luthra R, Romans A, Abraham R, Ensor J, Yao, Hamilton SR (2010) Association of microRNA expression with microsatellite instability status in colorectal adenocarcinoma. J Mole Diagn 12(4):433–440
Schaefer A, Jung M, Mollenkopf HJ, Wagner I, Stephan C, Jentzmik F, Miller K, Lein M, Kristiansen G, Jung K (2010) Diagnostic and prognostic implications of microRNA profiling in prostate carcinoma. Int J Canc 126(5):1166–1176
Nam EJ, Yoon HJ, Kim SW, Kim HG, Kim YT, Kim JH, Kim JW, Kim SH (2008) MicroRNA expression profiles in serous ovarian carcinoma. Clin Cancer Res 14(9):2690–2695
Aqeilan RI, Calin GA, Croce CM. miR-15a and miR-16-1 in cancer: discovery, function and future perspectives. Cell Death Differ 17(2):215–220
Schaefer A, Jung M, Miller K, Lein M, Kristiansen G, Erbersdobler A, Jung K (2010) Suitable reference genes for relative quantification of miRNA expression in prostate cancer. Exp Mol Med 42(11):749–758
Shan HL, Zhang Y, Lu YJ, Zhang Y, Pan ZW, Cai BZ, Wang N, Li XL, Feng TM, Hong Y, Yang BF (2009) Downregulation of miR-133 and miR-590 contributes to nicotine-induced atrial remodelling in canines. Cardiovasc Res 83(3):465–472
Abdelmohsen K, Srikantan S, Kuwano Y, Gorospe M (2008) miR-519 reduces cell proliferation by lowering RNA-binding protein HuR levels. Proc Natl Acad Sci U S A 105(51):20297–20302
Abdelmohsen K, Kim MM, Srikantan S, Mercken EM, Brennan SE, Wilson GM, de Cabo R, Gorospe M (2010) miR-519 suppresses tumor growth by reducing HuR levels. Cell Cycle 9(7):1354–1359
Ristimaki A (2010) Tumor suppressor effect of the microRNA miR-519 is mediated via the mRNA-binding protein HuR. Cell Cycle 9(7):1234–1234
Acknowledgments
We would like to thank the National Breast Cancer Research Institute for their financial support of the study.
Conflict of interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kheirelseid, E.A.H., Miller, N., Chang, K.H. et al. miRNA expressions in rectal cancer as predictors of response to neoadjuvant chemoradiation therapy. Int J Colorectal Dis 28, 247–260 (2013). https://doi.org/10.1007/s00384-012-1549-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00384-012-1549-9