Abstract
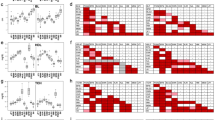
Few simple, easy-to-score PCR markers are available for studying genetic variation in wild mice populations belonging to Mus musculus at the population and subspecific levels. In this study, we show the abundant B1 family of short interspersed DNA elements (SINEs) is a very promising source of such markers. Thirteen Bl sequences from different regions of the genome were retrieved on the basis of their high degree of homology to a mouse consensus sequence, and the presence of these elements was screened for in wild derived mice representing M. spretus, macedonicus and spicilegus and the different subspecies of M. musculus. At five of these loci, varying degrees of insertion polymorphism were found in M. m. domesticus mice. These insertions were almost totally absent in the mice representing the other subspecies and species. Six other Bl elements were fixed in all the Mus species tested. At these loci, polymorphism associated with three restriction sites in the Bl consensus sequence was found in M. musculus. Most of these polymorphisms appear to be ancestral as they are shared by at least one of the other Mus species tested. Both insertion and restriction polymorphism revealed differences between five inbred laboratory strains considered to be of mainly domesticus origin, and at the six restriction loci a surprising number of these strains carried restriction variants that were either not found or very infrequent in domesticus. This suggests that in this particular group of loci, allcles of far Eastern origin are more frequent than expected.
Similar content being viewed by others
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z et al. (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25, 3389–3402
Batzer MA, Arcot SS, Phinney JW, Alegria-Hartman M, Kass DH et al. (1996) Genetic variation of recent Alu insertions in human populations. J Mol Evol 42, 22–29
Beck JA, Lloyd S, Hafezparast M, Lennon-Pierce M, Eppig JT et al. (2000) Genealogies of mouse inbred strains. Nat Genet 24, 23–25
Bishop CE, Boursot P, Bonhomme F, Hatat D (1985) Most classical Mus musculus domesticus laboratory mouse strains carry a Mus musculus musculus Y chromosome. Nature 315, 70–72
Bonhomme F, Guénet J-L, Dod B, Moriwaki K, Bulfield G (1987) The polyphyletic origin of laboratory inbred mice and their rate of evolution. Biol J Linn Soc 30, 51–58
Hardies SC, Wang L, Zhou L, Zhao Y, Casavant NC et al. (2000) LINE-1 (L1) lineages in the mouse. Mol Biol Evol 17, 616–628
Kass DH, Raynor ME, Williams TM (2000) Evolutionary history of B1 retroposons in the genus Mus. J Mol Evol 51, 256–264
Loeb DD, Padgett RW, Hardies SC, Shehee WR, Comer MB et al. (1986) The sequence of a large L1Md element reveals a tandemly repeated 5’ end and several features found in retrotransposons. Mol Cell Biol 6, 168–182
Nagamine CM, Nishioka Y, Moriwaki K, Boursot P, Bonhomme F et al. (1992) The musculus-type Y chromosome of the laboratory mouse is of Asian origin. Mamm Genome 3, 84–91
Quentin Y (1989) Successive waves of fixation of Bl variants in rodent lineage history. J Mol Evol 28, 299–305
Rogers JH (1985) The origin and evolution of retroposons. Int Rev Cytol 93, 187–279
Roy-Engel AM, Carroll ML, Vogel E, Garber RK, Nguyen SV et al. (2001) Alu insertion polymorphisms for the study of human genomic diversity. Genetics 159, 279–290
Rozen S, Skaletsky HJ (2000) Primer3 on the WWW for general users and for biologist programmers. In: Kra-wetz S, Misener S, (eds.) Bioinformatics Methods and Protocols: Methods in Molecular Biology. (Totowa, NJ: Humana Press), pp 365–386
Shedlock AM, Okada N (2000) SINE insertions: powerful tools for molecular systematics. BioEssays 22, 148–160
Smit AF (1999) Interspersed repeats and other mementos of transposable elements in mammalian genomes. Curr Opin Genet Dev 9, 657–663
Tucker PK, Lee BK, Lundrigan BL, Eicher EM (1992) Geographic origin of the Y chromosomes in “old” inbred strains of mice. Mamm Genome 3, 254–261
Ullu E, Tschudi C (1984) Alu sequences are processed 7SL RNA genes. Nature 312, 171–172
Wakasugi N, Tomita T, Kondo K (1967) Differences of fertility in reciprocal crosses between inbred strains of mice. J Reprod Fertil 13, 41–50
Zhao WD, Ishikawa A, Yamagata T, Bolor H, Wakasugi N (2002) Female mice of DDK strain are fully fertile in the intersubspecific crosses with Mus musculus molossinus and M. m. castaneus. Mamm Genome 13, 345–351
Zietkiewicz E, Labuda D (1996) Mosaic evolution of rodent Bl elements. J Mol Evol 42, 66–72
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Munclinger, P., Boursot, P. & Dod, B. B1 insertions as easy markers for mouse population studies. Mammalian Genome 14, 359–366 (2003). https://doi.org/10.1007/s00335-002-3065-7
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00335-002-3065-7