Abstract
Key message
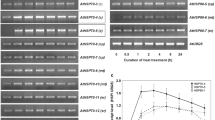
The study determined the tolerance of Aloe vera to high temperature, focusing on the expression of hsp70 , hsp100 and ubiquitin genes. These were highly expressed in plants acclimated at 35 °C prior to a heat shock of 45 °C.
Abstract
Aloe barbadensis Miller (Aloe vera), a CAM plant, was introduced into Chile in the semiarid IV and III Regions, which has summer diurnal temperature fluctuations of 25 to 40 °C and annual precipitation of 40 mm (dry years) to 170 mm (rainy years). The aim of this study was to investigate how Aloe vera responds to water and heat stress, focusing on the expression of heat shock genes (hsp70, hsp100) and ubiquitin, which not studied before in Aloe vera. The LT50 of Aloe vera was determined as 53.2 °C. To study gene expression by semi-quantitative RT-PCR, primers were designed against conserved regions of these genes. Sequencing the cDNA fragments for hsp70 and ubiquitin showed a high identity, over 95 %, with the genes from cereals. The protein sequence of hsp70 deduced from the sequence of the cDNA encloses partial domains for binding ATP and the substrate. The protein sequence of ubiquitin deduced from the cDNA encloses a domain for interaction with the enzymes E2, UCH and CUE. The expression increased with temperature and water deficit. Hsp70 expression at 40–45 °C increased 50 % over the controls, while the expression increased by 150 % over the controls under a water deficit of 50 % FC. The expression of all three genes was also studied under 2 h of acclimation at 35 or 40 °C prior to a heat shock at 45 °C. Under these conditions, the plants showed greater expression of all genes than when they were subjected to direct heat stress.







Similar content being viewed by others
References
Abràmoff MD, Magalhães PJ, Ram SJ (2004) Image Processing with ImageJ. Biophoton Int 11:36–42
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Altschul SF, Madden T, Schaffer A, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucl Acid Res 25:3389–3402
Biedermann S, Hellmann H (2011) WD40and CUL4-based ligases: lubricating aspects of life. Trends Biochem Sci 16:38–46
Borland AM, Taybi T (2004) Synchronization of metabolic processes in plants with Crassulacean acid metabolism. J Exp Bot 55:1255–1265
Callis J, Carpenter TR, Sun CW, Vierstra RD (1995) Structure and evolution of genes encoding polyubiquitin and ubiquitin-like proteins in Arabidopsis thaliana ecotype Columbia. Genetics 139:921–939
Cardemil L (2008) La asimilación de CO2 y síntesis de azúcares en las plantas. In: Cardemil L, Squeo F (eds) Fisiología vegetal, chapter 9. Edición de La Universidad de La Serena. Plant, Cell Environ 29:2113–2123
Ceusters J, Borland AM (2011) Impacts of elevated CO(2) on the growth and physiology of plants with crassulacean acid metabolism. Prog Bot 72:163–181
Ceusters J, Borland AM, Londers E, Verdoodt V, Godts C, De Proft MP (2009) Differential usage of storage carbohydrates in the CAM bromeliad Aechmea ‘Maya’ during acclimation to drought and recovery from dehydration. Physiol Plant 135:174–184
Delatorre-Herrera J, Delfino I, Salinas C, Silva H, Cardemil L (2010) Irrigation restriction effects on water use efficiency and osmotic adjustment in Aloe Vera plants (Aloe barbadensis Miller). Agric Water Manage 97:1564–1570
Gagne JM, Downes BP, Shiu SH, Durski AM, Vierstra RD (2002) The F-box subunit of the SCF E3 complex is encoded by a diverse superfamily of genes in Arabidopsis. Proc Natl Acad Sci USA 99:11519–11524
Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A (2003) ExPASy—the proteomics server for in-depth protein knowledge and analysis. Nucl Acid Res 31:3784–3788
Goidin D, Mammessier A, Staquet MJ, Schmitt D, Berthier-Vergnes O (2001) Ribosomal 18S RNA prevails over glyceraldehyde-3-phosphate dehydrogenase and b-actin genes as internal standard for quantitative comparison of mRNA levels in invasive and noninvasive human melanoma cell subpopulations. Anal Biochem 295:17–21
Groettrup M, Pelzer C, Schmidtke G, Hofmann K (2008) Activating the ubiquitin family: UBA6 challenges the field. Trends Biochem Sci 33:230–237
Gulli M, Corradi M, Rampino P, Marmiroli N, Perrotta C (2007) Four members of the HSP101 gene family are differently regulated in Triticum durum Desf. FEBS Lett 581:4841–4849
Hall AE (2001) Crop responses to environment. CRC Press LLC, Boca Raton
Herrera A (2009) Crassulacean acid metabolism and fitness under water deficits stress: if not for carbon gain, what is facultative CAM good for? Ann Bot 103:645–653
Hicke L, Schubert HL, Hill CP (2005) Ubiquitin-binding domains. Nature 6:610–621
Hisano H, Kanazawa A, Yoshida M, Humphreys MO, Lizuka M, Kitamura K, Yamada T (2008) Coordinated expression of functionally diverse fructosyltransferase genes is associated with fructan accumulation in response to low temperature in perennial ryegrass. New Phytol 178:766–780
Labarga A, Valentin F, Andersson M, Lopez R (2007) Web services at the European Bioinformatics Institute. Nucl Acid Res 35(suppl S):SW6–SW11
Luján R, Lledías F, Martínez LM, Barreto R, Cassab GI, Nieto-Sotelo J (2009) Small heat-shock proteins and leaf cooling capacity account for the unusual heat tolerance of the central spike leaves in Agave tequilana var Weber. Plant Cell Environ 32:1791–1803
Martineau JR, Specht JE, Williams JH, Sullivan CY (1979) Temperature tolerance in soybeans. I. Evaluation of a technique for assessing cellular membrane thermostability. Crop Sci 19:75–78
Melnikov EE, Rotanova TV (2010) Molecular chaperones. Russ J Bioorgan Chem 36:1–10
Mittler R, Finka A, Goloubinoff P (2012) How do plants feel the heat? Trends Biochem Sci (in press)
Mohnen D (2008) Pectin structure and biosynthesis Curr Op. Plant Biol 11:266–277
Moreira LRS, Filho EXF (2008) An overview of mannan structure and mannan-degrading enzyme systems. Appl Microbiol Biotechnol 79:165–178
Murray MG, Thompson WF (1980) Rapid isolation of high molecular-weight plant DNA. Nucl Acid Res 8:4321–4325
Nieto-Sotelo J, Kannan KB, Segal MC (1999) Characterization of a maize heat-shock protein 101 gene, HSP101, encoding a ClpB/Hsp100 protein homologue. Gene 230:187–195
Nieto-Sotelo J, Martinez LM, Ponce G, Cassab GI, Alagon A, Meeley RB, Ribaut JM, Yang R (2002) Maize HSP101 plays important roles in both induced and basal thermotolerance and primary root growth. Plant Cell 14:1621–1633
Ortiz C, Cardemil L (2001) Heat-shock responses in two leguminous plants. A comparative study. J Exp Bot 52:1711–1719
Ozkaynak E, Finley D, Solomon MJ, Varshvashky A (1987) The yeast ubiquitin genes: a family of natural genes fusions. EMBO J 6:1427–1439
Perales L, Peñarrubia L, Cornejo MJ (2008) Induction of a polyubiquitin gene promoter by dehydration stresses in transformed rice cells. J Plant Physiol 165:159–171
Pierce S, Winter K, Grifftihs H (2002) The role of CAM in high rainfall cloud forest: an in situ comparison of photosynthetic pathways in Bromeliaceae. Plant Cell Environ 25:1181–1189
Quevillon E, Silventoinen V, Pillai S, Harte N, Mulder N, Apweiler R, Lopez R (2005) InterProScan: protein domains identifier. Nucl Acid Res 33 (Web Server issue):W116–W120
Richter K, Haslbeck M, Buchner J (2010) The heat shock response: life on the verge of death. Mol Cell 40:253–266
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Schaff DA, Clayberg CD, Milliken GA (1987) Comparison of TTC and electrical conductivity as heat tolerance screening techniques in Phaseolus. Hort Sci 22:642–645
Schirmer EC, Glover JR, Singer MA, Lindquist S (1996) HSP lO0/Clp proteins: a common mechanism explains diverse functions. Trends Biochem Sci 21:289–296
Sharp PM, Li WH (1987) Ubiquitin genes as a paradigm of concerted evolution of tandem repeats. J Mol Evol 25:58–64
Silva H, Sagardía S, Seguel O, Torres C, Tapia C, Franck N, Cardemil L (2010) Effect of water availability on growth and water use efficiency for biomass and gel production in Aloe Vera (Aloe barbadensis M.). Ind Crops Prod 31:20–27
Sivamani E, Qu R (2006) Expression enhancement of a rice polyubiquitin gene promoter. Plant Mol Biol 60:225–239
Stürzenbaum S, Kille P (2001) Control genes in quantitative molecular biological techniques: the variability of invariante. Comp Bioch Physiol Part B 130:281–289
Sung DY, Vierling E, Guy CL (2001) Comprehensive expression profile analysis of the Arabidopsis hsp70 gene family. Plant Physiol 126:789–800
Vaasen A, Begerow D, Rüdiger H (2006) Phosphoenol pyruvate carboxylase genes in C3, crassulacean acid metabolism (CAM) and C3/CAM intermediate species of the genus Clusia: rapid reversible C3/CAM switches are based on the C3 housekeeping gene. Plant Cell Environ 29:2113–2123
Valluru R, Van den Ende W (2008) Plant fructans in stress environments: emerging concepts and future prospects. J Exp Bot 59:2905–2916
Van den Ende W, De Coninck B, Van Laere A (2004) Plant fructan exohydrolases: a role in signaling and defense? TIBS 9:523–528
Vierling E (1997) Hanbook of molecular chaperones and protein folding catalysts. Oxford University Press, Oxford, pp 253–255
Wang JL, Jiang JD, Oard JH (2000) Structure, expression and promoter activity of two polyubiquitin genes from rice (Oryza sativa L.). Plant Sci 156:201–211
Wang W-J, Li Q–Q, Xu J-D, Cao X–X, Li H-X, Tang F, Chen Q, Yang J-M, Xu Z-D, Liu X-P (2008) Over-expression of ubiquitin carboxy terminal hydrolase-L1 induces apoptosis in breast cancer cells. Int J Oncol 33:1037–1045
Acknowledgments
We thank Professor Claudia Stange for advice on RT-PCR amplification. The technical assistance of Angélica Vega is acknowledged. The research was supported by projects FONDECYT N° 1070899 and N° 7080094 and by Dirección de Investigación, Universidad de Chile, Project N° MULT 05/30-2.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. Puigdomenech.
Rights and permissions
About this article
Cite this article
Huerta, C., Freire, M. & Cardemil, L. Expression of hsp70, hsp100 and ubiquitin in Aloe barbadensis Miller under direct heat stress and under temperature acclimation conditions. Plant Cell Rep 32, 293–307 (2013). https://doi.org/10.1007/s00299-012-1363-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-012-1363-4