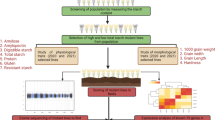
Abstract
The maize genome remains abundant in molecular diversity, and the rich genetic diversity of maize starch-synthesis genes is crucial for controlling various grain traits. To explore the unique mechanism controlling the advantageous waxy trait and characterize the molecular feature of genes relevant to starch composition in two elite waxy inbreds, expression profiling combined with gene organization analysis was performed in them as compared to one normal inbred. Genotype-specific expression patterns were observed for most genes studied. The waxy inbreds were shown to contain mutations in multiple starch-synthesis genes, namely gbssI (wx), gbssIIb and isa2 (potentially isa3 too).The mis-splicing events directly accounted for wx loss of function. Contrarily, disruption of 5′ and 3′ transcript sequence may contribute to the absence of GbssIIb and Isa2 transcripts in waxy inbreds, respectively. Besides, the splicing of Sugary1 transcript was developmentally regulated in the normal inbred, and DNA polymorphisms were detected within SSIIIb-1 gene in waxy inbreds.





Similar content being viewed by others
Abbreviations
- WAP:
-
Weeks after pollination
- GBSS:
-
Granule-bound starch synthase
- SSS:
-
Soluble starch synthase
- SBE:
-
Starch branching enzyme
- DBE:
-
Starch debranching enzyme
References
Brunner S, Fengler K, Morgante M, Tingey S, Rafalski A (2005) Evolution of DNA sequence non-homologies among maize inbreds. Plant Cell 17:343–360
Buckler ES, Stevens NM (2005) Maize origins, domestication, and selection. In: Motley TJ, Zerega N, Cross H (eds) Darwin’s harvest. Columbia University Press, New York, pp 67–90
Buckler ES, Gaut BS, McMullen MD (2006) Molecular and functional diversity of maize. Curr Opin Plant Biol 9:172–176
Delatte T, Trevisan M, Parker ML, Zeeman SC (2005) Arabidopsis mutants Atisa1 and Atisa2 have identical phenotypes and lack the same multimeric isoamylase, which influences the branch point distribution of amylopectin during starch synthesis. Plant J 41:815–830
Denyer K, Johnson P, Zeeman S, Smith AM (2001) The control of amylose synthesis. J Plant Physiol 158:479–487
Doehlert DC, Knutson CA (1991) Two classes of starch debranching enzymes from developing maize kernels. J Plant Physiol 138:566–572
Gonzales JW, Rhodes AM, Dickinson DB (1976) Carbohydrate and enzymic characterization of a high sucrose sugary inbred line of sweet corn. Plant Physiol 58:28–32
Hanashiro I, Itoh K, Kuratomi Y, Yamazaki M, Igarashi T, Matsugasako J, Takeda Y (2008) Granule-bound starch synthase I is responsible for biosynthesis of extra-long unit chains of amylopectin in rice. Plant Cell Physiol 49:925–933
Hennen-Bierwagen TA, Liu F, Marsh RS, Kim S, Gan Q, Tetlow IJ, Emes MJ, James MG, Myers AM (2008) Starch biosynthetic enzymes from developing Zea mays endosperm associate in multisubunit complexes. Plant Physiol 146:1892–1908
Hirose T, Terao T (2004) A comprehensive expression analysis of the starch synthase gene family in rice (Oryza sativa L.). Planta 220:9–16
Isshiki M, Tsumoto A, Shimamoto K (2006) The serine/arginine-rich protein family in rice plays important roles in constitutive and alternative splicing of pre-mRNA. Plant Cell 18:146–158
James MG, Denyer K, Myers AM (2003) Starch synthesis in the cereal endosperm. Curr Opin Plant Biol 6:215–222
Li L, Ilarslan H, James MG, Myers AM, Wurtele ES (2007) Genome wide co-expression among the starch debranching enzyme genes AtISA1, AtISA2, and AtISA3 in Arabidopsis thaliana. J Exp Bot 58:3323–3342
Maddelein ML, Libessart N, Bellanger F, Delrue B, D’Hulst C, Van den Koornhuyse N, Fontaine T, Wieruszeski JM, Decq A, Ball S (1994) Toward an understanding of the biogenesis of the starch granule. Determination of granule-bound and soluble starch synthase functions in amylopectin synthesis. J Biol Chem 269:25150–25157
Nakamura T, Vrinten P, Hayakawa K, Ikeda J (1998) Characterization of a granule-bound starch synthase isoform found in the pericarp of wheat. Plant Physiol 118:451–459
Nelson OE, Rines HW (1962) The enzymatic deficiency in the waxy mutant of maize. Biochem Biophys Res Commun 9:297–300
Rahman A, Wong K, Jane J, Myers AM, James MG (1998) Characterization of SU1 isoamylase, a determinant of storage starch structure in maize. Plant Physiol 117:425–435
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Shure M, Wessler S, Federoff N (1983) Molecular identification and isolation of the waxy locus in maize. Cell 35:225–233
Tetlow IJ (2006) Understanding storage starch biosynthesis in plants: a means to quality improvement. Can J Bot 84:1167–1185
Tracy WF, Whitt SR, Buckler ES (2006) Recurrent mutation and genome evolution: example of Sugary1 and the origin of sweet maize. Crop Sci 46(S1):1–6
Tsai CY (1974) The function of the waxy locus in starch synthesis in maize endosperm. Biochem Genet 11:83–96
Utsumi Y, Nakamura Y (2006) Structural and enzymatic characterization of the isoamylase1 homo-oligomer and the isoamylase1-isoamylase2 hetero-oligomer from rice endosperm. Planta 225:75–87
Varagona MJ, Purugganan M, Wessler SR (1992) Alternative splicing induced by insertion of retrotransposons into the maize waxy gene. Plant Cell 4:811–820
Vrinten PL, Nakamura T (2000) Wheat granule-bound starch synthase I and II are encoded by separate genes that are expressed in different tissues. Plant Physiol 122:255–264
Wang W (2000) Methods for determining plant soluble sugar content. In: Li HS (ed) Theories and methods for plant physiology and biochemistry experiments. Higher Education Press, Beijing, pp 194–197
Wattebled F, Dong Y, Dumez S, Delvallé D, Planchot V, Berbezy P, Vyas D, Colonna P, Chatterjee M, Ball S, D’Hulst C (2005) Mutants of Arabidopsis lacking a chloroplastic isoamylase accumulate phytoglycogen and an abnormal form of amylopectin. Plant Physiol 138:184–190
Wattebled F, Planchot V, Dong Y, Szydlowski N, Pontoire B, Devin A, Ball S, D’Hulst C (2008) Further evidence for the mandatory nature of polysaccharide debranching for the aggregation of semi-crystalline starch and for overlapping functions of debranching enzymes in Arabidopsis leaves. Plant Physiol 148:1309–1323
Wei RH (2006) Studies on genetic diversity of waxy corn germplasm resources. PhD thesis, Jilin Agricultural University, China
Wessler SR, Varagona MJ (1985) Molecular basis of mutations at the waxy locus of maize: correlation with the fine structure genetic map. Proc Natl Acad Sci USA 82:4177–4181
Acknowledgments
We thank Professor Yi-Fa Wang of the Crop Research Institute of SAAS for the kind guidance in the field experiments and Dr. Kevin M. Folta of the University of Florida for critical reading of an earlier version of the manuscript. This work was supported by the Science and Technology Commission of Shanghai Municipality (Major Program, 06DZ19101) and Shanghai Academy of Agricultural Sciences (Key Subject Construction, 2007(12) to K. D.).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by D. Somers.
X.-Z. Ding and B.-G. Wang contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
299_2009_748_MOESM1_ESM.doc
S1 provides gene identities and specific primer sequences for RT-PCR analysis. S2 shows the sequence alignment of Wx (X03935, DNA) with its various transcripts from herein three maize inbreds. S3 is the sequence alignment showing different SSIIIb-1 transcripts from three maize inbreds. S4 is the sequence alignment of maize Sugary1 gene with the alternative transcripts from kernels of different developing stages in 5003 inbred. (DOC 91 kb)
Rights and permissions
About this article
Cite this article
Ding, XZ., Wang, BG., Gao, QH. et al. Molecular diversity and differential expression of starch-synthesis genes in developing kernels of three maize inbreds. Plant Cell Rep 28, 1487–1495 (2009). https://doi.org/10.1007/s00299-009-0748-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-009-0748-5