Abstract
Members of marine Actinobacteria have been highly regarded as potentially important sources of antimicrobial compounds. Here, we isolated a strain of Actinobacteria, SZJ61, and showed that it inhibits the in vitro growth of fungi pathogenic to plants. This new isolate was identified as Streptomyces luteoverticillatus by morphological, biochemical and genetic analyses. Antifungal compounds were isolated from S. luteoverticillatus strain SZJ61 and characterized as carbazomycin B by nuclear magnetic resonance spectra. We then sequenced the genome of the S. luteoverticillatus SZJ61 strain, which consists of only one 7,367,863 bp linear chromosome that has a G+C content of 72.05%. Thirty-five putative biosynthetic gene clusters for secondary metabolites, including a variety of bioactive products, were found. Mining of the genome sequence information revealed the putative biosynthetic gene cluster of carbazomycin B. This genomic information is valuable for interpreting the biosynthetic mechanisms of diverse bioactive compounds that have potential applications in the pharmaceutical industry.


Similar content being viewed by others
References
Hwang KS et al (2014) Systems biology and biotechnology of streptomyces species for the production of secondary metabolites. Biotechnol Adv 32(2):255–268
Baltz RH (2008) Renaissance in antibacterial discovery from actinomycetes. Curr Opin Pharmacol 8(5):557–563
Dhakal D et al (2018) Complete genome sequence of, streptomyces peucetius, atcc 27952, the producer of anticancer anthracyclines and diverse secondary metabolites. J Biotechnol 267:50–54
Yu Y et al (2018) Identification of the streptothricin and tunicamycin biosynthetic gene clusters by genome mining in streptomyces sp. strain fd1-xmd. Appl Microbiol Biotechnol 102(6):2621–2633
Kaneda M et al (1990) Biosynthesis of carbazomycin B. II. Origin of the whole carbon skeleton. J Antibiot 43(12):1623–1626
Yamasaki K et al (1983) New antibiotics, carbazomycins A and B III. Taxonomy and biosynthesis. J Antibiot 36(5):552–558
Knolker HJ, Frohner W (1989) Improved total syntheses of the antibiotic alkaloids carbazomycin A and B. Neuroscience 28(3):735–744
Markad SB, Argade NP (2014) Diversity oriented convergent access for collective total synthesis of bioactive multifunctional carbazole alkaloids: synthesis of carbazomycin A, carbazomycin B, hyellazole, chlorohyellazole, and clausenaline D. Org Lett 16(20):5470–5473
Kato S et al (1993) In vitro and ex vivo free radical scavenging activities of carazostatin, carbazomycin B and their derivatives. J Antibiot 46(12):1859–1865
Managamuri U et al (2017) Isolation, identification, optimization, and metabolite profiling of streptomyces sparsus vsm-30. 3 Biotech 7(3):217
Meng X et al (2017) Characterization and complete genome sequence of a novel siphoviridae bacteriophage BS5. Curr Microbiol 74(7):815–820
Grady EN et al (2019) Characterization and complete genome analysis of the surfactin-producing, plant-protecting bacterium bacillus velezensis 9d-6. BMC Microbiol 19(1):5
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Krishnan P et al (2004) Characterisation of germinating and non-germinating wheat seeds by nuclear magnetic resonance (nmr) spectroscopy. Eur Biophys J 33(1):76–82
Koren S et al (2012) Hybrid error correction and de novo assembly of single-molecule sequencing reads. Nat Biotechnol 30(7):693–700
Chin CS et al (2013) Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Methods 10(6):563–569
McKenna A et al (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20(9):1297–1303
Rhoads A, Au KF (2015) PacBio Sequencing and Its Applications. Genom Proteom Bioinform 13(5):278–289
Weber T et al (2015) antiSMASH 3.0-a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res 43:W237–W243
Sakano KI, Nakamura S (1980) New antibiotics, carbazomycins A and B II. Structural elucidation. J Antibiot 33(9):961–966
Orihara N, Furihata K, Seto H (1997) Studies on the biosynthesis of terpenoidal compounds produced by actinomycetes. 2. Biosynthesis of carquinostatin B via the non-mevalonate pathway in Streptomyces exfoliatus. J Antibiot 50(11):979–981
Huang S et al (2015) Biosynthesis of neocarazostatin A reveals the sequential carbazole prenylation and hydroxylation in the tailoring steps. Chem Biol 22(12):1633–1642
Acknowledgments
This work was financially supported by the National Key R&D Program of China (Grant No. 2017YFD0501800), the Innovation Team Project for Modern Agricultural Industrious Technology System of Shandong Province (Grant No. SDAIT-11-10), Major Scientific and Technological Innovation Project of Shandong Province (Grant No. 2017GGH5129), and Yantai Science and Technology Project (Grant No. 2017NC049).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Research Involving Human and Animals Participants
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
284_2019_1711_MOESM1_ESM.tif
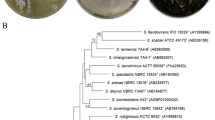
Supplementary material 1—Antagonistic effect of S. luteoverticillatus SZJ61on the growth of plant pathogenic fungi. Aspergillus niger (A) and Fusarium oxysporum (B). (TIFF 226 kb)
Rights and permissions
About this article
Cite this article
Feng, Z., Chen, G., Zhang, J. et al. Characterization and Complete Genome Analysis of the Carbazomycin B-Producing Strain Streptomyces luteoverticillatus SZJ61. Curr Microbiol 76, 982–987 (2019). https://doi.org/10.1007/s00284-019-01711-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01711-x