Abstract
During the last few decades, whole-cell biosensors have attracted increasing attention for their enormous potential in monitoring bioavailable heavy metal contaminations in the ecosystem. Visual and measurable output signals by employing natural pigments have been demonstrated to offer another potential choice to indicate the existence of bioavailable heavy metals in recent years. The biosynthesis of the blue pigment indigoidine has been achieved in E. coli following heterologous expression of both BpsA (a single-module non-ribosomal peptide synthetase) and PcpS (a PPTase to activate apo-BpsA). Moreover, we demonstrated herein the development of the indigoidine-based whole-cell biosensors to detect bioavailable Hg(II) and Pb(II) in water samples by employing metal-responsive transcriptional regulator MerR and PbrR as the sensory elements, and the indigoidine biosynthesis gene cluster as a reporter element. The resulting indigoidine-based biosensors presented a good selectivity and high sensitivity to target metal ions. High concentration of target metal exposure could be clearly recognized by the naked eye due to the color change by the secretion of indigoidine, and quantified by measuring the absorbance of the culture supernatants at 600 nm. Dose–response relationships existed between the exposure concentrations of target heavy metals and the production of indigoidine. Although fairly good linear relationships were obtained in a relatively limited concentration range of the concentrations of heavy metal ions, these findings suggest that genetically controlled indigoidine biosynthesis triggered by the MerR family transcriptional regulator can enable a sensitive, visual, and qualitative whole-cell biosensor for bioindicating the presence of bioaccessible heavy metal in environmental water samples.
Key points
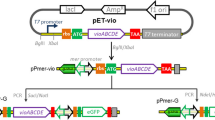
• Biosynthesis pathway of indigoidine reconstructed in a high copy number plasmid in E. coli.
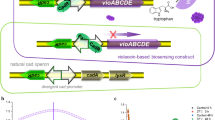
• Visual and colorimetric detection of Hg(II) and Pb(II) by manipulation of indigoidine biosynthesis through MerR family metalloregulator.
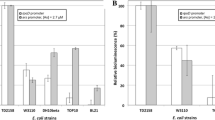
•Enhanced detection sensitivity toward Hg(II) and Pb(II) achieved using novel pigment-based whole-cell biosensors.










Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article (and its Supplementary information files).
References
Bereza-Malcolm LT, Mann G, Franks AE (2015) Environmental sensing of heavy metals through whole cell microbial biosensors: a synthetic biology approach. ACS Synth Biol 4(5):535–546. https://doi.org/10.1021/sb500286r
Bereza-Malcolm L, Aracic S, Franks AE (2016) Development and application of a synthetically-derived lead biosensor construct for use in Gram-negative bacteria. Sensors (Basel) 16(12):2174. https://doi.org/10.3390/s16122174
Brachmann AO, Kirchner F, Kegler C, Kinski SC, Schmitt I, Bode HB (2012) Triggering the production of the cryptic blue pigment indigoidine from Photorhabdus luminescens. J Biotechnol 157(1):96–99. https://doi.org/10.1016/j.jbiotec.2011.10.002
Brocklehurst KR, Megit SJ, Morby AP (2003) Characterisation of CadR from Pseudomonas aeruginosa: a Cd(II)-responsive MerR homologue. Biochem Biophys Res Commun 308(2):234–239. https://doi.org/10.1016/s0006-291x(03)01366-4
Brown NL, Stoyanov JV, Kidd SP, Hobman JL (2003) The MerR family of transcriptional regulators. FEMS Microbiol Rev 27(2-3):145–163. https://doi.org/10.1016/S0168-6445(03)00051-2
Brown AS, Robins KJ, Ackerley DF (2017) A sensitive single-enzyme assay system using the non-ribosomal peptide synthetase BpsA for measurement of L-glutamine in biological samples. Sci Rep 7:41745. https://doi.org/10.1038/srep41745
Cai S, Shen Y, Zou Y, Sun P, Wei W, Zhao J, Zhang C (2018) Engineering highly sensitive whole-cell mercury biosensors based on positive feedback loops from quorum-sensing systems. Analyst 143(3):630–634. https://doi.org/10.1039/c7an00587c
Chen P-H, Lin C, Guo K-H, Yeh Y-C (2017) Development of a pigment-based whole-cell biosensor for the analysis of environmental copper. RSC Adv 7(47):29302–29305. https://doi.org/10.1039/C7RA03778C
Cude WN, Mooney J, Tavanaei AA, Hadden MK, Frank AM, Gulvik CA, May AL, Buchan A (2012) Production of the antimicrobial secondary metabolite indigoidine contributes to competitive surface colonization by the marine roseobacter Phaeobacter sp. strain Y4I. Appl Environ Microbiol 78(14):4771–4780. https://doi.org/10.1128/AEM.00297-12
Cude WN, Prevatte CW, Hadden MK, May AL, Smith RT, Swain CL, Campagna SR, Buchan A (2015) Phaeobacter sp. strain Y4I utilizes two separate cell-to-cell communication systems to regulate production of the antimicrobial indigoidine. Appl Environ Microbiol 81(4):1417–1425. https://doi.org/10.1128/AEM.02551-14
Du R, Guo M, He X, Huang K, Luo Y, Xu W (2019) Feedback regulation mode of gene circuits directly affects the detection range and sensitivity of lead and mercury microbial biosensors. Anal Chim Acta 1084:85–92. https://doi.org/10.1016/j.aca.2019.08.006
Finking R, Solsbacher J, Konz D, Schobert M, Schafer A, Jahn D, Marahiel MA (2002) Characterization of a new type of phosphopantetheinyl transferase for fatty acid and siderophore synthesis in Pseudomonas aeruginosa. J Biol Chem 277(52):50293–50302. https://doi.org/10.1074/jbc.M205042200
Guo M, Du R, Xie Z, He X, Huang K, Luo Y, Xu W (2019a) Using the promoters of MerR family proteins as "rheostats" to engineer whole-cell heavy metal biosensors with adjustable sensitivity. J Biol Eng 13:70. https://doi.org/10.1186/s13036-019-0202-3
Guo Y, Hui CY, Liu L, Zheng HQ, Wu HM (2019b) Improved monitoring of low-level transcription in Escherichia coli by a beta-galactosidase alpha-complementation system. Front Microbiol 10:1454. https://doi.org/10.3389/fmicb.2019.01454
Guo Y, Hui CY, Zhang NX, Liu L, Li H, Zheng HJ (2021) Development of cadmium multiple-signal biosensing and bioadsorption systems based on artificial cad operons. Front Bioeng Biotechnol 9:585617. https://doi.org/10.3389/fbioe.2021.585617
Gupta N, Renugopalakrishnan V, Liepmann D, Paulmurugan R, Malhotra BD (2019) Cell-based biosensors: Recent trends, challenges and future perspectives. Biosens Bioelectron 141:111435. https://doi.org/10.1016/j.bios.2019.111435
Hansen LH, Sorensen SJ (2000) Versatile biosensor vectors for detection and quantification of mercury. FEMS Microbiol Lett 193(1):123–127. https://doi.org/10.1111/j.1574-6968.2000.tb09413.x
Hicks M, Bachmann TT, Wang B (2020) Synthetic biology enables programmable cell-based biosensors. Chemphyschem 21(2):131. https://doi.org/10.1002/cphc.201901191
Hui C, Guo Y, Zhang W, Gao C, Yang X, Chen Y, Li L, Huang X (2018a) Surface display of PbrR on Escherichia coli and evaluation of the bioavailability of lead associated with engineered cells in mice. Sci Rep 8(1):5685. https://doi.org/10.1038/s41598-018-24134-3
Hui CY, Guo Y, Yang XQ, Zhang W, Huang XQ (2018b) Surface display of metal binding domain derived from PbrR on Escherichia coli specifically increases lead(II) adsorption. Biotechnol Lett 40(5):837–845. https://doi.org/10.1007/s10529-018-2533-4
Hui CY, Guo Y, Zhang W, Huang XQ (2018c) Rapid monitoring of the target protein expression with a fluorescent signal based on a dicistronic construct in Escherichia coli. AMB Express 8(1):81. https://doi.org/10.1186/s13568-018-0612-5
Hui CY, Guo Y, Liu L, Zheng HQ, Wu HM, Zhang LZ, Zhang W (2019) Development of a novel bacterial surface display system using truncated OmpT as an anchoring motif. Biotechnol Lett 41(6-7):763–777. https://doi.org/10.1007/s10529-019-02676-4
Hui CY, Guo Y, Liu L, Zhang NX, Gao CX, Yang XQ, Yi J (2020a) Genetic control of violacein biosynthesis to enable a pigment-based whole-cell lead biosensor. RSC Adv 10(47):28106–28113. https://doi.org/10.1039/D0RA04815A
Hui CY, Guo Y, Liu L, Zheng HQ, Gao CX, Zhang W (2020b) Construction of a RFP-lacZalpha bicistronic reporter system and its application in lead biosensing. PLoS One 15(1):e0228456. https://doi.org/10.1371/journal.pone.0228456
Joe MH, Lee KH, Lim SY, Im SH, Song HP, Lee IS, Kim DH (2012) Pigment-based whole-cell biosensor system for cadmium detection using genetically engineered Deinococcus radiodurans. Bioprocess Biosyst Eng 35(1-2):265–272. https://doi.org/10.1007/s00449-011-0610-3
Jung J, Lee SJ (2019) Biochemical and biodiversity insights into heavy metal ion-responsive transcription regulators for synthetic biological heavy metal sensors. J Microbiol Biotechnol 29(10):1522–1542. https://doi.org/10.4014/jmb.1908.08002
Kim HJ, Jeong H, Lee SJ (2018) Synthetic biology for microbial heavy metal biosensors. Anal Bioanal Chem 410(4):1191–1203. https://doi.org/10.1007/s00216-017-0751-6
Kumar S, Verma N, Singh A (2017) Development of cadmium specific recombinant biosensor and its application in milk samples. Sensors Actuators B Chem 240:248–254
Lee SW, Glickmann E, Cooksey DA (2001) Chromosomal locus for cadmium resistance in Pseudomonas putida consisting of a cadmium-transporting ATPase and a MerR family response regulator. Appl Environ Microbiol 67(4):1437–1444. https://doi.org/10.1128/AEM.67.4.1437-1444.2001
Levin-Karp A, Barenholz U, Bareia T, Dayagi M, Zelcbuch L, Antonovsky N, Noor E, Milo R (2013) Quantifying translational coupling in E. coli synthetic operons using RBS modulation and fluorescent reporters. ACS Synth Biol 2(6):327–336. https://doi.org/10.1021/sb400002n
Lim CG, Wong L, Bhan N, Dvora H, Xu P, Venkiteswaran S, Koffas MA (2015) Development of a recombinant Escherichia coli strain for overproduction of the plant pigment anthocyanin. Appl Environ Microbiol 81(18):6276–6284. https://doi.org/10.1128/AEM.01448-15
Lu Y, Liu J, Li J, Bruesehoff PJ, Pavot CM, Brown AK (2003) New highly sensitive and selective catalytic DNA biosensors for metal ions. Biosens Bioelectron 18(5-6):529–540. https://doi.org/10.1016/s0956-5663(03)00013-7
Magrisso S, Erel Y, Belkin S (2008) Microbial reporters of metal bioavailability. Microb Biotechnol 1(4):320–330. https://doi.org/10.1111/j.1751-7915.2008.00022.x
McNerney MP, Michel CL, Kishore K, Standeven J, Styczynski MP (2019) Dynamic and tunable metabolite control for robust minimal-equipment assessment of serum zinc. Nat Commun 10(1):5514. https://doi.org/10.1038/s41467-019-13454-1
Muller M, Auslander S, Auslander D, Kemmer C, Fussenegger M (2012) A novel reporter system for bacterial and mammalian cells based on the non-ribosomal peptide indigoidine. Metab Eng 14(4):325–335. https://doi.org/10.1016/j.ymben.2012.04.002
Qian F, Zhang C, Zhang Y, He W, Gao X, Hu P, Guo Z (2009) Visible light excitable Zn2+ fluorescent sensor derived from an intramolecular charge transfer fluorophore and its in vitro and in vivo application. J Am Chem Soc 131(4):1460–1468. https://doi.org/10.1021/ja806489y
Rasmussen LD, Turner RR, Barkay T (1997) Cell-density-dependent sensitivity of a mer-lux bioassay. Appl Environ Microbiol 63(8):3291–3293. https://doi.org/10.1128/AEM.63.8.3291-3293.1997
Reverchon S, Rouanet C, Expert D, Nasser W (2002) Characterization of indigoidine biosynthetic genes in Erwinia chrysanthemi and role of this blue pigment in pathogenicity. J Bacteriol 184(3):654–665. https://doi.org/10.1128/jb.184.3.654-665.2002
Stoyanov JV, Brown NL (2003) The Escherichia coli copper-responsive copA promoter is activated by gold. J Biol Chem 278(3):1407–1410. https://doi.org/10.1074/jbc.C200580200
Takahashi H, Kumagai T, Kitani K, Mori M, Matoba Y, Sugiyama M (2007) Cloning and characterization of a Streptomyces single module type non-ribosomal peptide synthetase catalyzing a blue pigment synthesis. J Biol Chem 282(12):9073–9081. https://doi.org/10.1074/jbc.M611319200
Tao HC, Peng ZW, Li PS, Yu TA, Su J (2013) Optimizing cadmium and mercury specificity of CadR-based E. coli biosensors by redesign of CadR. Biotechnol Lett 35(8):1253–1258. https://doi.org/10.1007/s10529-013-1216-4
USEPA (1984) National recommended water quality criteria. Washington, DC: Office of water
Vareda JP, Valente AJM, Duraes L (2019) Assessment of heavy metal pollution from anthropogenic activities and remediation strategies: a review. J Environ Manag 246:101–118. https://doi.org/10.1016/j.jenvman.2019.05.126
Wang D, Zheng Y, Fan X, Xu L, Pang T, Liu T, Liang L, Huang S, Xiao Q (2020) Visual detection of Hg2+ by manipulation of pyocyanin biosynthesis through the Hg2+-dependent transcriptional activator MerR in microbial cells. J Biosci Bioeng 129(2):223–228. https://doi.org/10.1016/j.jbiosc.2019.08.005
Watstein DM, Styczynski MP (2018) Development of a pigment-based whole-cell zinc biosensor for human serum. ACS Synth Biol 7(1):267–275. https://doi.org/10.1021/acssynbio.7b00292
WHO (2011) Guidelines for drinking water quality (4th ed.). Water quality standards. Geneva: World health organization
Xie Z, Zhang Z, Cao Z, Chen M, Li P, Liu W, Qin H, Zhao X, Tao Y, Chen Y (2017) An external substrate-free blue/white screening system in Escherichia coli. Appl Microbiol Biotechnol 101(9):3811–3820. https://doi.org/10.1007/s00253-017-8252-2
Xu F, Gage D, Zhan J (2015) Efficient production of indigoidine in Escherichia coli. J Ind Microbiol Biotechnol 42(8):1149–1155. https://doi.org/10.1007/s10295-015-1642-5
Yoon Y, Kim S, Chae Y, Kim SW, Kang Y, An G, Jeong SW, An YJ (2016) Simultaneous detection of bioavailable arsenic and cadmium in contaminated soils using dual-sensing bioreporters. Appl Microbiol Biotechnol 100(8):3713–3722. https://doi.org/10.1007/s00253-016-7338-6
Yu D, Xu F, Valiente J, Wang S, Zhan J (2013) An indigoidine biosynthetic gene cluster from Streptomyces chromofuscus ATCC 49982 contains an unusual IndB homologue. J Ind Microbiol Biotechnol 40(1):159–168. https://doi.org/10.1007/s10295-012-1207-9
Zhang W, Hu X, Wang L, Wang X (2014) Reconstruction of the carotenoid biosynthetic pathway of Cronobacter sakazakii BAA894 in Escherichia coli. PLoS One 9(1):e86739. https://doi.org/10.1371/journal.pone.0086739
Zhang NX, Guo Y, Li H, Yang XQ, Gao CX, Hui CY (2021) Versatile artificial mer operons in Escherichia coli towards whole cell biosensing and adsorption of mercury. PLoS One 16(5):e0252190. https://doi.org/10.1371/journal.pone.0252190
Zhu G, Zhang CY (2014) Functional nucleic acid-based sensors for heavy metal ion assays. Analyst 139(24):6326–6342. https://doi.org/10.1039/c4an01069h
Funding
This work was supported by the National Natural Science Foundation of China (82073517); the Natural Science Foundation of Guangdong Province (2019A1515011989; 2021A1515012472); the Science and Technology Program of Shenzhen (JCYJ20180306170237563; JCYJ20190808175205480); Shenzhen Fund for Guangdong Provincial High-level Clinical Key Specialties (SZGSP015); and Shenzhen Key Medical Discipline Construction Fund (SZXK068).
Author information
Authors and Affiliations
Contributions
C-Y. H. and N-X. Z. designed the study; Y. G., L-M. L., Y-T. C., and J. Y. performed the experiments. C-Y. H. and L. L. analyzed the experimental data; C-Y. H. and L. L. wrote and revised the manuscript. All the authors reviewed the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(PDF 594 kb)
Rights and permissions
About this article
Cite this article
Hui, Cy., Guo, Y., Li, Lm. et al. Indigoidine biosynthesis triggered by the heavy metal-responsive transcription regulator: a visual whole-cell biosensor. Appl Microbiol Biotechnol 105, 6087–6102 (2021). https://doi.org/10.1007/s00253-021-11441-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11441-5