Abstract
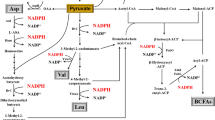
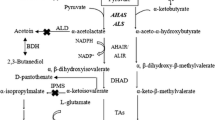
Lichenysin is categorized into the family of lipopeptide biosurfactants and has a variety of applications in the petroleum industry, bioremediation, pharmaceuticals, and the food industry. Currently, large-scale production is limited due to the low yield. This study found that lichenysin production was repressed by supplementation of extracellular amino acids. The global transcriptional factor CodY was hypothesized to prevent lichenysin biosynthesis under an amino acid-rich condition in Bacillus licheniformis. Thus, the codY null strain was constructed, and lichenysin production was increased by 31.0% to 2356 mg/L with the addition of precursor amino acids, and the lichenysin production efficiency was improved by 42.8% to 98.2 mg/L• h. Correspondingly, the transcription levels of the lichenysin synthetase gene lchAA, and its corresponding regulator genes comA, degQ, and degU, were upregulated. Also, the codY deletion enhanced biosynthesis of lichenysin precursor amino acids (Gln, Ile, Leu, and Val) and reduced the formation of byproducts, acetate, acetoin, and 2,3-butanediol. This study firstly reported that lichenysin biosynthesis was negatively regulated by CodY and lichenysin production could be further improved with the precursor amino acid amendment in the codY null strain.





Similar content being viewed by others
References
Belitsky BR (2015) Role of branched-chain amino acid transport in Bacillus subtilis CodY activity. J Bacteriol 197:1330–1338
Belitsky BR, Sonenshein AL (2013) Genome-wide identification of Bacillus subtilis CodY-binding sites at single-nucleotide resolution. Proc Natl Acad Sci 110:7026–7031
Brinsmade SR, Alexander EL, Livny J, Stettner AI, Segrè D, Rhee KY, Sonenshein AL (2014) Hierarchical expression of genes controlled by the Bacillus subtilis global regulatory protein CodY. Proc Natl Acad Sci 111:8227–8232
Dangel V, Westrich L, Smith MCM, Heide L, Gust B (2010) Use of an inducible promoter for antibiotic production in a heterologous host. Appl Microbiol Biotechnol 87:261–269
Duitman EH, Wyczawski D, Boven LG, Venema G, Kuipers OP, Hamoen LW (2007) Novel methods for genetic transformation of natural Bacillus subtilis isolates used to study the regulation of the mycosubtilin and surfactin synthetases. Appl Environ Microbiol 73:3490–3496
Fujita Y, Satomura T, Tojo S, Hirooka K (2014) CcpA-mediated catabolite activation of the Bacillus subtilis ilv-leu operon and its negation by either CodY- or TnrA-mediated negative regulation. J Bacteriol 196:3793–3806
Grangemard I, Wallach J, Maget-Dana R, Peypoux F (2001) Lichenysin: a more efficient cation chelator than surfactin. Appl Biochem Biotechnol 90:199–210
Hashizume H, Igarashi M, Sawa R, Adachi H, Nishimura Y, Akamatsu Y (2008) A new type of tripropeptin with anteiso-branched chain fatty acid from Lysobacter sp. BMK333-48F3. J Antibiot 61:577–582
Jiao S, Li X, Yu HM, Yang H, Li X, Shen ZY (2017) In situ enhancement of surfactin biosynthesis in Bacillus subtilis using novel artificial inducible promoters. Biotechnol Bioeng 114(4):832–842
Kobayashi K (2007) Gradual activation of the response regulator DegU controls serial expression of genes for flagellum formation and biofilm formation in Bacillus subtilis. Mol Microbiol 66:395–409
Liang C, Huo Y, Qi G, Wei X, Wang Q, Chen S (2015) Enhancement of L-valine production in Bacillus licheniformis by blocking three branched pathways. Biotechnol Lett 37:1243–1248
Liu JF, Yang J, Yang SZ, Ye RQ, Mu BZ (2012) Effects of different amino acids in culture media on surfactin variants produced by Bacillus subtilis TD7. Appl Biochem Biotechnol 166:2091–2100
Madslien EH, Ronning HT, Lindback T, Hassel B, Andersson MA, Granum PE (2013) Lichenysin is produced by most Bacillus licheniformis strains. J Appl Microbiol 115:1068–1080
Nerurkar AS (2010) Structural and molecular characteristics of lichenysin and its relationship with surface activity. Biosurfactants 672:304–315
Qi GF, Kang YF, Li L, Xiao AF, Zhang SM, Wen ZY, Xu DH, Chen SW (2014) Deletion of meso-2,3-butanediol dehydrogenase gene budC for enhanced D-2,3-butanediol production in Bacillus licheniformis. Biotechnol Biofuels 7:6–12
Qiu Y, Xiao F, Wei X, Wen Z, Chen S (2014) Improvement of lichenysin production in Bacillus licheniformis by replacement of native promoter of lichenysin biosynthesis operon and medium optimization. Appl Microbiol Biotechnol 98:8895–8903
Roongsawang N, Washio K, Morikawa M (2010) Diversity of nonribosomal peptide synthetases involved in the biosynthesis of lipopeptide biosurfactants. Int J Mol Sci 12:141–172
Santos CC, Libeck BS, Schwan RF (2014) Co-culture fermentation of peanut-soy milk for the development of a novel functional beverage. Int J Food Microbiol 186:32–41
Shivers RP, Dineen SS, Sonenshein AL (2006) Positive regulation of Bacillus subtilis ackA by CodY and CcpA: establishing a potential hierarchy in carbon flow. Mol Microbiol 62:811–822
Sonenshein AL (2005) CodY, a global regulator of stationary phase and virulence in Gram-positive bacteria. Curr Opin Microbiol 8:203–207
Tian G, Fu J, Wei X, Ji Z, Ma X, Qi G, Chen S (2014) Enhanced expression of pgdS gene for high production of poly-γ-glutamic aicd with lower molecular weight in Bacillus licheniformis WX-02. J Chem Technol Biotechnol 89:1825–1832
Wang J, Yuan H, Wei X, Chen J, Chen S (2016) Enhancement of poly-γ-glutamic acid production by alkaline pH stress treatment in Bacillus licheniformis WX-02. J Chem Technol Biotechnol 91(9):2399–2403
Watanabe T, Morita T, Koike H, Yarimizu T, Shinozaki Y, Sameshima-Yamashita Y, Yoshida S, Koitabashi M, Kitamoto H (2016) High-level recombinant protein production by the basidiomycetous yeast Pseudozyma antarctica under a xylose-inducible xylanase promoter. Appl Microbiol Biotechnol 100(7):3207–3217
Wu Q, Zhang R, Peng S, Xu Y (2015) Transcriptional characteristics associated with lichenysin biosynthesis in Bacillus licheniformis from Chinese Maotai-flavor liquor making. J Agr Food Chem 63:888–893
Yakimov MM, Golyshin PN (1997) ComA-dependent transcriptional activation of lichenysin A synthetase promoter in Bacillus subtilis cells. Biotechnol Prog 13:757–761
Yakimov MM, Fredrickson HI, Timmis KN (1996) Effect of heterogeneity of hydrophobic moieties on surface activity of lichenysin A, a lipopeptide biosurfactant from Bacillus licheniformis BAS50. Biotechnol Appl Biochem 23:13–18
Yangtse WM, Zhou YH, Lei Y, Qiu YM, Wei XT, Ji ZX, Qi GF, Yong YC, Chen LL, Chen SW (2012) Genome sequence of Bacillus licheniformis WX-02. J Bacteriol 194:3561–3562
Acknowledgements
This work was funded by the National Science & Technology Pillar Program during the Twelfth Five-Year Plan Period (2013AA102801-52), the National Program on Key Basic Research Project (973 Program, No. 2015CB150505), and the Science and Technology Program of Wuhan (20160201010086).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This paper does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 301 kb)
Rights and permissions
About this article
Cite this article
Zhu, C., Xiao, F., Qiu, Y. et al. Lichenysin production is improved in codY null Bacillus licheniformis by addition of precursor amino acids. Appl Microbiol Biotechnol 101, 6375–6383 (2017). https://doi.org/10.1007/s00253-017-8352-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8352-z