Abstract
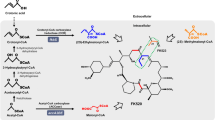
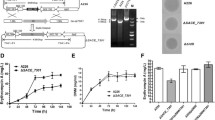
Ascomycin (FK520), a macrocyclic polyketide natural antibiotic, displays high anti-fungal and immunosuppressive activity. In this study, the LysR family transcriptional regulator FkbR1 was characterized, and its role in ascomycin biosynthesis was explored by gene deletion, complementation, and overexpression. Inactivation of fkbR1 led to 67.5% reduction of ascomycin production, which was restored by complementation of fkbR1. Overexpression of fkbR1 resulted in a 33.5% increase in ascomycin production compared with the parent strain FS35. These findings indicated that FkbR1 was a positive regulator for ascomycin production. Quantitative RT-PCR analysis revealed that the expressions of fkbE, fkbF, fkbS, and fkbU were downregulated in the fkbR1 deletion strain and upregulated in the fkbR1 overexpression strain. Electrophoretic mobility shift assays (EMSAs) in vitro and chromatin immunoprecipitation (ChIP)-qPCR assays in vivo indicated that FkbR1 bound to the intergenic region of fkbR1-fkbE. To investigate the roles of the target genes fkbE and fkbF in ascomycin production, the deletion and overexpressions of fkbE and fkbF were implemented, respectively. Overexpression of fkbE resulted in a 45.6% increase in ascomycin production, but little change was observed in fkbF overexpression strain. To further enhance ascomycin production, the fkbR1 and fkbE combinatorial overexpression strain OfkbRE was constructed with the ascomycin yield increased by 69.9% to 536.7 mg/L compared with that of the parent strain. Our research provided a helpful strategy to increase ascomycin production via engineering FkbR1 and its target gene.






Similar content being viewed by others
References
Alanazi AM, Neidle EL, Momany C (2013) The DNA-binding domain of BenM reveals the structural basis for the recognition of a T-N11—a sequence motif by LysR-type transcriptional regulators. Acta Crystallogr 69:1995–2007. doi:10.1107/S0907444913017320
Allen JP, Neely MN (2011) The Streptococcus iniae transcriptional regulator CpsY is required for protection from neutrophil-mediated killing and proper growth in vitro. Infect Immun 79:4638–4648. doi:10.1128/IAI.05567-11
An JH, Yu SK (1998) A gene cluster encoding malonyl-CoA decarboxylase (MatA), malonyl-CoA synthetase (MatB) and a putative dicarboxylate carrier protein (MatC) in Rhizobium trifolii. Eur J Biochem 257:395–402. doi:10.1046/j.1432-1327.1998.2570395.x
Andexer JN, Kendrew SG, Nur-e-Alam M, Lazos O, Foster TA, Zimmermann A-S, Warneck TD, Suthar D, Coates NJ, Koehn FE (2011) Biosynthesis of the immunosuppressants FK506, FK520, and rapamycin involves a previously undescribed family of enzymes acting on chorismate. Proc Natl Acad Sci U S A 108(12):4776–4781. doi:10.1073/pnas.1015773108
Barreiro C, Martinez-Castro M (2014) Trends in the biosynthesis and production of the immunosuppressant tacrolimus (FK506). Appl Microbiol Biotechnol 98:497–507. doi:10.1007/s00253-013-5362-3
Bibb MJ (2005) Regulation of secondary metabolism in streptomycetes. Curr Opin Microbiol 8:208–215. doi:10.1016/j.mib.2005.02.016
Chen Y, Wendt-Pienkowski E, Shen B (2008) Identification and utility of FdmR1 as a Streptomyces antibiotic regulatory protein activator for fredericamycin production in Streptomyces griseus ATCC 49344 and heterologous hosts. J Bacteriol 190:5587–5596. doi:10.1128/JB.00592-08
Gatto GJ, McLoughlin SM, Kelleher NL, Walsh CT (2005) Elucidating the substrate specificity and condensation domain activity of FkbP, the FK520 pipecolate-incorporating enzyme. Biochemist 44:5993–6002. doi:10.1021/bi050230w
Goranovic D, Blazic M, Magdevska V, Horvat J, Kuscer E, Polak T, Santos-Aberturas J, Martinez-Castro M, Barreiro C, Mrak P, Kopitar G, Kosec G, Fujs S, Martin JF, Petkovic H (2012) FK506 biosynthesis is regulated by two positive regulatory elements in Streptomyces tsukubaensis. BMC Microbiol 12:238. doi:10.1186/1471-2180-12-238
Guo J, Zhao J, Li L, Chen Z, Wen Y, Li J (2010) The pathway-specific regulator AveR from Streptomyces avermitilis positively regulates avermectin production while it negatively affects oligomycin biosynthesis. Mol Gen Genomics 283:123–133. doi:10.1007/s00438-009-0502-2
Huang D, Li S, Xia M, Wen J, Jia X (2013) Genome-scale metabolic network guided engineering of Streptomyces tsukubaensis for FK506 production improvement. Microb Cell Factories 12:52. doi:10.1186/1475-2859-12-52
Kieser T, Bibb M, Buttner M, Chater K (2000) Practical streptomyces genetics. John Innes Foundation, Norwich
Kim J, Senadheera DB, Levesque CM, Cvitkovitch DG (2012) TcyR regulates L-cystine uptake via the TcyABC transporter in Streptococcus mutans. FEMS Microbiol Lett 328:114–121. doi:10.1111/j.1574-6968.2011.02492.x
Kovacikova G, Lin W, Skorupski K (2010) The LysR-type virulence activator AphB regulates the expression of genes in Vibrio cholerae in response to low pH and anaerobiosis. J Bacteriol 192:4181–4191. doi:10.1128/JB.00193-10
Kumar P, Sharma SK, Malviya HK, Balasubramanian R, Dalal A (2007) An improved fermentation process for preparing ascomycin. WO 2007/029082
Li Y, He ZG (2012) The mycobacterial LysR-type regulator OxyS responds to oxidative stress and negatively regulates expression of the catalase-peroxidase gene. PLoS One 7:e30186. doi:10.1371/journal.pone.0030186
Liu SP, Yu P, Yuan PH, Zhou ZX, Bu QT, Mao XM, Li YQ (2015a) Sigma factor WhiGch positively regulates natamycin production in Streptomyces chattanoogensis L10. Appl Microbiol Biotechnol 99:2715–2726. doi:10.1007/s00253-014-6307-1
Liu WS, Zhang QL, Guo J, Chen Z, Li JL, Wen Y (2015b) Increasing avermectin production in Streptomyces avermitilis by manipulating the expression of a novel TetR-family regulator and its target gene product. Appl Environ Microbiol 81:5157–5173. doi:10.1128/AEM.00868-15
Lu ZL, Takeuchi M, Sato T (2007) The LysR-type transcriptional regulator YofA controls cell division through the regulation of expression of ftsW in Bacillus subtilis. J Bacteriol 189:5642–5651. doi:10.1128/JB.00467-07
Maddocks SE, Oyston PC (2008) Structure and function of the LysR-type transcriptional regulator (LTTR) family proteins. Microbiology 154:3609–3623. doi:10.1099/mic.0.2008/022772-0
Mao XM, Sun ZH, Liang BR, Wang ZB, Feng WH, Huang FL, Li YQ (2013) Positive feedback regulation of stgR expression for secondary metabolism in Streptomyces coelicolor. J Bacteriol 195:2072–2078. doi:10.1128/JB.00040-13
Martin JF, Liras P (2010) Engineering of regulatory cascades and networks controlling antibiotic biosynthesis in Streptomyces. Curr Opin Microbiol 13:263–273. doi:10.1016/j.mib.2010.02.008
Mo S, Yoo YJ, Ban YH, Lee SK, Kim E, Suh JW, Yoon YJ (2012) Roles of fkbN in positive regulation and tcs7 in negative regulation of FK506 biosynthesis in Streptomyces sp. strain KCTC 11604BP. Appl Environ Microbiol 78:2249–2255. doi:10.1128/aem.06766-11
Monaghan P, Fardis M, Revill WP, Bell A (2005) Antimalarial effects of macrolactones related to FK520 (ascomycin) are independent of the immunosuppressive properties of the compounds. J Infect Dis 191:1342–1349. doi:10.1086/428454
Myronovskyi M, Welle E, Fedorenko V, Luzhetskyy A (2011) Beta-glucuronidase as a sensitive and versatile reporter in actinomycetes. Appl Environ Microbiol 77:5370–5383. doi:10.1128/AEM.00434-11
Oliver P, Peralta-Gil M, Tabche ML, Merino E (2016) Molecular and structural considerations of TF-DNA binding for the generation of biologically meaningful and accurate phylogenetic footprinting analysis: the LysR-type transcriptional regulator family as a study model. BMC Genomics 17:686. doi:10.1186/s12864-016-3025-3
Paradkar AS, Aidoo KA, Jensen SE (1998) A pathway-specific transcriptional activator regulates late steps of clavulanic acid biosynthesis in Streptomyces clavuligerus. Mol Microbiol 27:831–843. doi:10.1046/j.1365-2958.1998.00731.x
Picossi S, Belitsky BR, Sonenshein AL (2007) Molecular mechanism of the regulation of Bacillus subtilis gltAB expression by GltC. J Mol Biol 365:1298–1313. doi:10.1016/j.jmb.2006.10.100
Qi HS, Xin X, Li SS, Wen JP, Chen Y, Jia XQ (2012) Higher-level production of ascomycin (FK520) by Streptomyces hygroscopicus var. ascomyceticus irradiated by femtosecond laser. Biotechnol Bioproc E 17:770–779. doi:10.1007/s12257-012-0114-2
Qi HS, Zhao SM, Fu H, Wen JP, Jia XQ (2014a) Enhancement of ascomycin production in Streptomyces hygroscopicus var. ascomyceticus by combining resin HP20 addition and metabolic profiling analysis. J Ind Microbiol Biotechnol 41:1365–1374. doi:10.1007/s10295-014-1473-9
Qi HS, Zhao SM, Wen JP, Chen Y, Jia XQ (2014b) Analysis of ascomycin production enhanced by shikimic acid resistance and addition in Streptomyces hygroscopicus var. ascomyceticus. Biochem Eng J 82:124–133. doi:10.1016/j.bej.2013.11.006
Qi HS, Lv MM, Song KJ, Wen JP (2016) Integration of parallel 13C-labeling experiments and in silico pathway analysis for enhanced production of ascomycin. Biotechnol Bioeng. doi:10.1002/bit.26223
Rashid MM, Ikawa Y, Tsuge S (2016) GamR, the LysR-type galactose metabolism regulator, regulates hrp gene expression via transcriptional activation of two key hrp regulators, HrpG and HrpX, in Xanthomonas oryzae pv. oryzae. Appl Environ Microbiol 82:3947–3958. doi:10.1128/AEM.00513-16
Revill WP, Voda J, Reeves CR, Chung L, Schirmer A, Ashley G, Carney JR, Fardis M, Carreras CW, Zhou Y (2002) Genetically engineered analogs of ascomycin for nerve regeneration. J Pharmacol Exp Ther 302(3):1278–1285. doi:10.1124/jpet.102.034264
Rhee KY, Opel M, Ito E, Hung S-p, Arfin SM, Hatfield GW (1999) Transcriptional coupling between the divergent promoters of a prototypic LysR-type regulatory system, the ilvYC operon of Escherichia coli. Proc Natl Acad Sci U S A 96:14294–14299. doi:10.1073/pnas.96.25.14294
Rodriguez M, Nunez LE, Brana AF, Mendez C, Salas JA, Blanco G (2008) Identification of transcriptional activators for thienamycin and cephamycin C biosynthetic genes within the thienamycin gene cluster from Streptomyces cattleya. Mol Microbiol 69:633–645. doi:10.1111/j.1365-2958.2008.06312.x
Sierra-Paredes G, Sierra-Marcuño G (2008) Ascomycin and FK506: pharmacology and therapeutic potential as anticonvulsants and neuroprotectants. CNS Neurosci Ther 14:36–46. doi:10.1111/j.1527-3458.2008.00036.x
Wang WS, Ji JJ, Li X, Wang J, Li SS, Pan GH, Fan KQ, Yang KQ (2014) Angucyclines as signals modulate the behaviors of Streptomyces coelicolor. Proc Natl Acad Sci U S A 111(15):5688–5693. doi:10.1073/pnas.1324253111
Wu K, Chung L, Revill WP, Katz L, Reeves CD (2000) The FK520 gene cluster of Streptomyces hygroscopicus var. ascomyceticus (ATCC 14891) contains genes for biosynthesis of unusual polyketide extender units. Gene 251:81–90. doi:10.1016/S0378-1119(00)00171-2
Yang SJ, Rice KC, Brown RJ, Patton TG, Liou LE, Park YH, Bayles KW (2005) A LysR-type regulator, CidR, is required for induction of the Staphylococcus aureus cidABC operon. J Bacteriol 187:5893–5900. doi:10.1128/JB.187.17.5893-5900.2005
Zhang XS, Luo HD, Tao Y, Wang YY, Jiang XH, Jiang H, Li YQ (2016) FkbN and Tcs7 are pathway-specific regulators of the FK506 biosynthetic gene cluster in Streptomyces tsukubaensis L19. J Ind Microbiol Biotechnol 43:1693–1703. doi:10.1007/s10295-016-1849-0
Zhu ZH, Han L, Yu P, Guo YY, Shuai L, Chen ZB, Mao XM, Guan WJ, Li YQ (2017) SlnR is a positive pathway-specific regulator for salinomycin biosynthesis in Streptomyces albus. Appl Microbiol Biotechnol:1547–1557. doi:10.1007/s00253-016-7918-5
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (No. 21676189), the Key Technologies R&D Program of Tianjin (No. 16YFZCSY00780), the National 973 Project of China (No. 2013CB733600), and the Key Program of National Natural Science Foundation of China (No. 21236005).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 827 kb)
Rights and permissions
About this article
Cite this article
Song, K., Wei, L., Liu, J. et al. Engineering of the LysR family transcriptional regulator FkbR1 and its target gene to improve ascomycin production. Appl Microbiol Biotechnol 101, 4581–4592 (2017). https://doi.org/10.1007/s00253-017-8242-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8242-4