Abstract
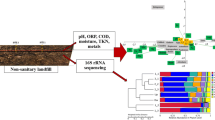
Little is known about the bacterial diversity of landfills and how environmental factors impact the diversity. In this study, PCR-based 454 pyrosequencing was used to investigate the bacterial communities of ten landfill leachate samples from five landfill sites in China. A total of 137 K useable sequences from the V3-V6 regions of the 16S rRNA gene were retrieved from 205 K reads. These sequences revealed the presence of a large number of operational taxonomic units (OTUs) in the landfills (709–1599 OTUs per sample). The most predominant bacterial representatives in the landfills investigated, regardless of geographic area, included Gammaproteobacteria, Firmicutes, and Bacteroidetes. The phyla Fusobacteria and Tenericutes were also found for the first time to be predominant in the landfills. The phylum Fusobacteria predominated (51.5 and 48.8 %) in two semi-arid landfills, and the phylum Tenericutes dominated (30.6 %) at one humid, subtropical landfill. Further, a large number of Pseudomonas was detected in most samples, comprising the dominant group and accounting for 40.9 to 92.4 % of the total abundance. Principal component analysis (PCA) and cluster analysis based on OTU abundance showed that the abundant taxa separated the bacterial community. Canonical correlation analysis (CCA) suggested that precipitation and landfilling age significantly impact on the bacterial community structure. The bacterial community function (e.g., cellulolytic bacteria, sulfate-reducing bacteria (SRB), sulfate-oxidizing bacteria, and xenobiotic organic compound (XOC)-degrading bacteria) was also diverse, but the pattern is unclear.



Similar content being viewed by others
References
Bareither CA, Wolfe GL, McMahon KD, Benson CH (2013) Microbial diversity and dynamics during methane production from municipal solid waste. Waste Manag 33(10):1982–1992. doi:10.1016/j.wasman.2012.12.013
Barlaz MA (1997) Microbial studies of landfills and anaerobic refuse decomposition. Manual of environmental microbiology. ASM Press, Washington DC, pp 541–557
Barlaz MA, Schaefer DM, Ham RK (1989) Bacterial population development and chemical characteristics of refuse decomposition in a simulated sanitary landfill. Appl Environ Microb 55:55–65
Bolhuis H, Stal LJ (2011) Analysis of bacterial and archaeal diversity in coastal microbial mats using massive parallel 16S rRNA gene tag sequencing. ISME J 5:1701–1712. doi:10.1038/ismej.2011.52
Braak CJ, FtaŠ P (2002) CANOCO Reference manual and CanoDraw for Windows User’s Guide: software for Canocical Community Ordination. Version 4.5
Brad T, van Breukelen BM, Braster M, van Straalen NM, Roling WFM (2008) Spatial heterogeneity in sediment-associated bacterial and eukaryotic communities in a landfill leachate-contaminated aquifer. FEMS Microbiol Ecol 65:534–543. doi:10.1111/j.1574-6941.2008.00533.x
Chen ACIH, Sekiguchi Y, Ohashi A, Harada H (2003) Archaeal community compositions at different depths (up to 30 m) of a municipal solid waste landfill in Taiwan as revealed by 16S rDNA cloning analyses. Biotechnol Lett 25:6. doi:10.1023/A:1023458631699
Czepiel PM, Shorter JH, Mosher B, Allwine E, McManus JB, Harriss RC, Kolb CE, Lamb BK (2003) The influence of atmospheric pressure on landfill methane emissions. Waste Manag 23:593–598. doi:10.1016/S0956-053X(03)00103-X
Daly K, Sharp RJ, McCarthy AJ (2000) Development of oligonucleotide probes and PCR primers for detecting phylogenetic subgroups of sulfate-reducing bacteria. Microbiology 146:1693–1705
Devereux R, Kane MD, Winfrey J, Stahl DA (1992) Genus- and group-specific hybridization probes for determinative and environmental studies of sulfate-reducing bacteria. Syst Appl Microbiol 15:601–609. doi:10.1016/S0723-2020(11)80122-0
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. doi:10.1093/bioinformatics/btr381
Finegold SM, Vaisanen M-L, Molitoris DR, Tomzynski TJ, Song Y, Liu C, Collins MD, Lawson PA (2003) Cetobacterium somerae sp. nov. from human feces and emended description of the genus Cetobacterium. Syst Appl Microbiol 26:177–181. doi:10.1078/072320203322346010
Guo HL, Chen C, Lee DJ, Wang AJ, Ren NQ (2013) Sulfur-nitrogen-carbon removal of Pseudomonas sp C27 under sulfide stress. Enzyme Microb Tech 53:6–12. doi:10.1016/j.enzmictec.2013.04.002
Huang LNZH, Chen YQ, Luo S, Lan CY, Qu LH (2002) Diversity and structure of the archaeal community in the leachate of a full-scale recirculating landfill as examined by direct 16S rRNA gene sequence retrieval. FEMS Microbiol Lett 214:6. doi:10.1111/j.1574-6968.2002.tb11353.x
Huang L-N, Chen Y-Q, Zhou H, Luo S, Lan C-Y, Qu L-H (2003) Characterization of methanogenic Archaea in the leachate of a closed municipal solid waste landfill. FEMS Microbiol Ecol 46:171–177. doi:10.1016/s0168-6496(03)00218-6
Huang L-N, Zhou H, Zhu S, Qu L-H (2004) Phylogenetic diversity of bacteria in the leachate of a full-scale recirculating landfill. FEMS Microbiol Ecol 50:175–183. doi:10.1016/j.femsec.2004.06.008
Jami E, Israel A, Kotser A, Mizrahi I (2013) Exploring the bovine rumen bacterial community from birth to adulthood. ISME J 7:1069–1079. doi:10.1038/ismej.2013.2
Jia S, Zhang X, Zhang G, Yin A, Zhang S, Li F, Wang L, Zhao D, Yun Q, Wang J (2013) Seasonally variable intestinal metagenomes of the red palm weevil (Rhynchophorus ferrugineus). Environ Microbiol 15:3020–3029. doi:10.1111/1462-2920.12262
Jones JG, Simon BM (1984) The presence and activity of Desulfotomaculum spp in sulphate-limited freshwater sediments. FEMS Microbiol Lett 21:47–50. doi:10.1111/j.1574-6968.1984.tb00184.x
Jørgensen BB (1982) Mineralization of organic matter in the sea bed—the role of sulphate reduction. Nature 296:643–645. doi:10.1038/296643a0
Kirchman DL (2002) The ecology of Cytophaga–Flavobacteria in aquatic environments. FEMS Microbiol Ecol 39:91–100. doi:10.1111/j.1574-6941.2002.tb00910.x
Kjeldsen P, Barlaz MA, Rooker AP, Baun A, Ledin A, Christensen TH (2002) Present and long-term composition of MSW landfill leachate: a review. Crit Rev Env Sci Tec 32:297–336. doi:10.1080/10643380290813462
Krishnamurthi S, Chakrabarti T (2013) Diversity of bacteria and archaea from a landfill in Chandigarh, India as revealed by culture-dependent and culture-independent molecular approaches. Syst Appl Microbiol 36:56–68. doi:10.1016/j.syapm.2012.08.009
Laloui-Carpentier W, Li T, Vigneron V, Mazéas L, Bouchez T (2006) Methanogenic diversity and activity in municipal solid waste landfill leachates. Anton Leeuw 89:423–434. doi:10.1007/s10482-005-9051-9
Lalucat J, Bennasar A, Bosch R, García-Valdés E, Palleroni NJ (2006) Biology of Pseudomonas stutzeri. Microbiol Mol Biol R 70:510–547. doi:10.1128/MMBR.00047-05
Lynd LR, Weimer PJ, van Zyl WH, Pretorius IS (2002) Microbial cellulose utilization: fundamentals and biotechnology. Microbiol Mol Biol R 66:506–577. doi:10.1128/mmbr.66.3.506-577.2002
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209
Marchesi JR, Sato T, Weightman AJ, Martin TA, Fry JC, Hiom SJ, Wade WG (1998) Design and evaluation of useful Bacterium-specific PCR primers that amplify genes coding for bacterial 16S rRNA. Appl Environ Microb 64:795–799
McDonald JE, Lockhart RJ, Cox MJ, Allison HE, McCarthy AJ (2008) Detection of novel Fibrobacter populations in landfill sites and determination of their relative abundance via quantitative PCR. Environ Microbiol 10:1310–1319. doi:10.1111/j.1462-2920.2007.01544.x
McDonald JE, Allison HE, McCarthy AJ (2010) Composition of the landfill microbial community as determined by application of domain- and group-specific 16S and 18S rRNA-targeted oligonucleotide probes. Appl Environ Microb 76:1301–1306. doi:10.1128/aem.01783-09
Nevin KP, Holmes DE, Woodard TL, Hinlein ES, Ostendorf DW, Lovley DR (2005) Geobacter bemidjiensis sp. nov. and Geobacter psychrophilus sp. nov., two novel Fe (III)-reducing subsurface isolates. Int J Syst Evol Micro 55:1667–1674. doi:10.1099/ijs.0.63417-0
Pourcher A-M, Sutra L, Hébé I, Moguedet G, Bollet C, Simoneau P, Gardan L (2001) Enumeration and characterization of cellulolytic bacteria from refuse of a landfill. FEMS Microbiol Ecol 34:229–241. doi:10.1111/j.1574-6941.2001.tb00774.x
Qiu G, Y-h S, Zeng P, Duan L, Xiao S (2013) Characterization of bacterial communities in hybrid upflow anaerobic sludge blanket (UASB)–membrane bioreactor (MBR) process for berberine antibiotic wastewater treatment. Bioresource Technol 142:52–62. doi:10.1016/j.biortech.2013.04.077
Riviere D, Desvignes V, Pelletier E, Chaussonnerie S, Guermazi S, Weissenbach J, Li T, Camacho P, Sghir A (2009) Towards the definition of a core of microorganisms involved in anaerobic digestion of sludge. ISME J 3:700–714. doi:10.1038/ismej.2009.2
Roeselers G, Mittge EK, Stephens WZ, Parichy DM, Cavanaugh CM, Guillemin K, Rawls JF (2011) Evidence for a core gut microbiota in the zebrafish. ISME J 5:1595–1608. doi:10.1038/ismej.2011.38
Sawamura H, Yamada M, Endo K, Soda S, Ishigaki T, Ike M (2010) Characterization of microorganisms at different landfill depths using carbon-utilization patterns and 16S rRNA gene based T-RFLP. J Biosci Bioeng 109:130–137. doi:10.1016/j.jbiosc.2009.07.020
Shah HN, Olsen I, Bernard K, Finegold SM, Gharbia S, Gupta RS (2009) Approaches to the study of the systematics of anaerobic, Gram-negative, non-sporeforming rods: current status and perspectives. Anaerobe 15:179–194. doi:10.1016/j.anaerobe.2009.08.003
Sievert SM, Scott KM, Klotz MG, Chain PSG, Hauser LJ, Hemp J, Hügler M, Land M, Lapidus A, Larimer FW, Lucas S, Malfatti SA, Meyer F, Paulsen IT, Ren Q, Simon J (2008) Genome of the Epsilonproteobacterial Chemolithoautotroph Sulfurimonas denitrificans. Appl Environ Microb 74:1145–1156. doi:10.1128/aem.01844-07
Song L-Y, Wang Y-Q (2015). Investigation of microbial community structure of a shallow lake after one season copper sulfate algaecide treatment. Microbiol Res 170:105–113. doi:10.1016/j.micres.2014.08.008
Song L, Shi L, Zhao Y, Li H (2011) Novel engineering controls to increase leachate contaminant degradation by refuse: from lab test to in situ engineering application. Ecol Eng 37:1914–1919. doi:10.1016/j.ecoleng.2011.06.023
Song L, Wang Y, Tang W, Lei Y (2015) Archaeal community diversity in municipal waste landfill sites. Appl Microbiol Biotechnol: 1–13 doi:10.1007/s00253-015-6493-5
Staley BF, de los Reyes FL, Barlaz MA (2012) Comparison of Bacteria and Archaea communities in municipal solid waste, individual refuse components, and leachate. FEMS Microbiol Ecol 79:465–473. doi:10.1111/j.1574-6941.2011.01239.x
Townsend TG, Miller WL, Lee HJ, Earle JFK (1996) Acceleration of landfill stabilization using leachate recycle. J Environ Eng ASCE 122:263–268. doi:10.1061/(asce)0733-9372(1996)122:4(263)
Tsuchiya C, Sakata T, Sugita H (2008) Novel ecological niche of Cetobacterium somerae, an anaerobic bacterium in the intestinal tracts of freshwater fish. Lett Appl Microbiol 46:43–48. doi:10.1111/j.1472-765X.2007.02258.x
Urios L, Agogué H, Lesongeur F, Stackebrandt E, Lebaron P (2006) Balneola vulgaris gen. nov., sp. nov., a member of the phylum Bacteroidetes from the north-western Mediterranean Sea. Int J Syst Evol Micro 56:1883–1887. doi:10.1099/ijs.0.64285-0
Urios L, Intertaglia L, Lesongeur F, Lebaron P (2008) Balneola alkaliphila sp. nov., a marine bacterium isolated from the Mediterranean Sea. Int J Syst Evol Micro 58:1288–1291. doi:10.1099/ijs.0.65555-0
Uroz S, Oger P, Morin E, Frey-Klett P (2012) Distinct ectomycorrhizospheres share similar bacterial communities as revealed by pyrosequencing-based analysis of 16S rRNA Genes. Appl Environ Microb 78:3020–3024. doi:10.1128/aem.06742-11
Van Dyke MI, McCarthy AJ (2002) Molecular biological detection and characterization of Clostridium populations in municipal landfill sites. Appl Environ Microb 68:2049–2053. doi:10.1128/aem.68.4.2049-2053.2002
van Kessel MA, Dutilh BE, Neveling K, Kwint MP, Veltman JA, Flik G, Jetten MS, Klaren PH, den Camp HJO (2011) Pyrosequencing of 16S rRNA gene amplicons to study the microbiota in the gastrointestinal tract of carp (Cyprinus carpio L.). AMB Expr 1:1–9. doi:10.1186/2191-0855-1-41
Wan R, Zhang S, Xie S (2012) Microbial community changes in aquifer sediment microcosm for anaerobic anthracene biodegradation under methanogenic condition. J Environ Sci 24(8):1498–1503. doi:10.1016/S1001-0742(11)60959-5
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microb 73:5261–5267. doi:10.1128/aem.00062-07
Wang Z, Yang Y, Dai Y, Xie S (2015) Anaerobic biodegradation of nonylphenol in river sediment under nitrate-or sulfate-reducing conditions and associated bacterial community. J Hazard Mater 286:306–314. doi:10.1016/j.jhazmat.2014.12.057
Westlake K, Archer DB, Boone DR (1995) Diversity of cellulolytic bacteria in landfill. J Appl Microbiol 79:73–78. doi:10.1111/j.1365-2672.1995.tb03126.x
Xie B, Xiong S, Liang S, Hu C, Zhang X, Lu J (2012) Performance and bacterial compositions of aged refuse reactors treating mature landfill leachate. Bioresource Technol 103:71–77. doi:10.1016/j.biortech.2011.09.114
Yang Y, Wang Z, He T, Dai Y, Xie S (2014) Sediment bacterial communities associated with anaerobic biodegradation of bisphenol a. Microb Ecol 12:1–8. doi:10.1007/s00248-014-0551-x
Zhang T, Shao M-F, Ye L (2012) 454 Pyrosequencing reveals bacterial diversity of activated sludge from 14 sewage treatment plants. ISME J 6:1137–1147. doi:10.1038/ismej.2011.188
Zhao Y, Song L, Huang R, Song L, Li X (2007) Recycling of aged refuse from a closed landfill. Waste Manag Res 25:130–138. doi:10.1177/0734242x07074053
Acknowledgments
The authors gratefully acknowledge the financial support of Chinese Academy of Science, China, (Contract No: KZCX2-XB3-14) and Natural Science Foundation of Chongqing (Contract No: cstc2014jcyjA20006). We thank the anonymous reviewers for offering critical suggestions that greatly improved the manuscript.
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1396 kb)
Rights and permissions
About this article
Cite this article
Song, L., Wang, Y., Tang, W. et al. Bacterial community diversity in municipal waste landfill sites. Appl Microbiol Biotechnol 99, 7745–7756 (2015). https://doi.org/10.1007/s00253-015-6633-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6633-y