Abstract
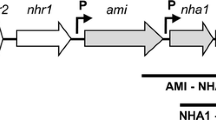
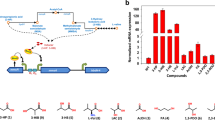
Rhodococcus erythropolis CCM2595 is able to efficiently utilize phenol and other aromatic compounds. We cloned and sequenced its complete gene cluster — catA, catB, catC, catR, pheR, pheA2, pheA1 — involved in the ortho-cleavage pathway of phenol. The activity of the key enzyme of the phenol degradation pathway, two-component phenol hydroxylase, was found to be induced by phenol. When both phenol and succinate were present in the medium, phenol hydroxylase activity decreased substantially. To analyze the regulation of phenol degradation at the transcriptional level, the transcriptional fusions of the divergently oriented promoters PpheA2 and PpheR with the gfpuv reporter gene were constructed. The promoters driving expression of the genes of the pheR–pheA2pheA1 cluster were localized by determining the respective transcriptional start points. Measurements of GFP fluorescence as well as quantitative RT-PCR revealed that expression of the phe genes is induced by phenol at the transcriptional level. The transcription of pheA2A1 and pheR was repressed by succinate, whereas no repression by glucose or glycerol was observed. Activation of the R. erythropolis CCM2595 pheA2 promoter by PheR, an AraC-type transcriptional regulator, was demonstrated by overexpression of the pheR gene. Analysis of the transcriptional regulation of two similar phe clusters from R. jostii RHA1 by various substrates showed that the type of carbon catabolite repression and the temporal transcriptional pattern during cultivation are different in each of the three phe clusters analyzed.







Similar content being viewed by others
References
Bleichrodt FS, Fischer R, Gerischer UC (2010) The β-ketoadipate pathway of Acinetobacter baylyi undergoes carbon catabolite repression, cross-regulation and vertical regulation, and is affected by Crc. Microbiology 156:1313–1322
Bradford MA (1972) A rapid and sensitive method for the quantitation of microgram quantities of protein using the principle of dye binding. Anal Biochem 72:248–252
Cases I, de Lorenzo V (2005) Promoters in the environment: transcriptional regulation in its natural context. Nat Rev Microbiol 3:105–118
Choi KY, Zylstra GJ, Kim E (2007) Benzoate catabolite repression of the phthalate degradation pathway in Rhodococcus sp. strain DK17. Appl Environ Microbiol 73:1370–1374
Duffner FM, Müller R (1998) A novel phenol hydroxylase and catechol 2,3-dioxygenase from the thermophilic Bacillus thermoleovorans strain A2: nucleotide sequence and analysis of the genes. FEMS Microbiol Lett 161:37–45
Dusch N, Pühler A, Kalinowski J (1999) Expression of the Corynebacterium glutamicum panD gene encoding L-aspartate-α-decarboxylase leads to pantothenate overproduction in Escherichia coli. Appl Environ Microbiol 65:1530–1539
González-Pérez MM, Ramos JL, Gallegos MT, Marqués S (1999) Critical nucleotides in the upstream region of the XylS-dependent TOL meta-cleavage pathway operon promoter as deduced from analysis of mutants. J Biol Chem 274:2286–2290
Hanahan E (1985) Techniques for transformation of E. coli. In: Glover DM (ed) DNA cloning. A practical approach, vol 1. IRL, Oxford, pp 109–135
Hernandez MA, Mohn WW, Martinez E, Rost E, Alvarez AF, Alvarez HM (2008) Biosynthesis of storage compounds by Rhodococcus jostii RHA1 and global identification of genes involved in their metabolism. BMC Genomics 9:600
Holátko J, Elišáková V, Prouza M, Sobotka M, Nešvera J, Pátek M (2009) Metabolic engineering of the L-valine biosynthesis pathway in Corynebacterium glutamicum using promoter activity modulation. J Biotechnol 139:203–210
Kim IC, Oriel PJ (1995) Characterization of the Bacillus stearothermophilus BR219 phenol hydroxylase gene. Appl Environ Microbiol 61:1252–1256
Kirchner O, Tauch A (2003) Tools for genetic engineering in the amino acid-producing bacterium Corynebacterium glutamicum. J Biotechnol 104:287–299
Kirchner U, Westphal AH, Müller R, van Berkel WJ (2003) Phenol hydroxylase from Bacillus thermoglucosidasius A7, a two-protein component monooxygenase with a dual role for FAD. J Biol Chem 278:47545–47553
Knoppová M, Phensaijai M, Veselý M, Zemanová M, Nešvera J, Pátek M (2007) Plasmid vectors for testing in vivo promoter activities in Corynebacterium glutamicum and Rhodococcus erythropolis. Curr Microbiol 55:234–239
Larkin MJ, Kulakov LA, Allen CC (2005) Biodegradation and Rhodococcus—masters of catabolic versatility. Curr Opin Biotechnol 16:282–290
Martin RW (1949) Rapid colorimetric estimation of phenol. Nature 21:1419–1420
Martínková L, Uhnáková B, Pátek M, Nešvera J, Křen V (2009) Biodegradation potential of the genus Rhodococcus. Environ Int 35:162–177
McLeod MP, Warren RL, Hsiao WW, Araki N, Myhre M, Fernandes C, Miyazawa D, Wong W, Lillquist AL, Wang D, Dosanjh M, Hara H, Petrescu A, Morin RD, Yang G, Stott JM, Schein JE, Shin H, Smailus D, Siddiqui AS, Marra MA, Jones SJ, Holt R, Brinkman FS, Miyauchi K, Fukuda M, Davies JE, Mohn WW, Eltis LD (2006) The complete genome of Rhodococcus sp. RHA1 provides insights into a catabolic powerhouse. Proc Natl Acad Sci U S A 103:15582–15587
Müller C, Petruschka L, Cuypers H, Burchhardt G, Herrmann H (1996) Carbon catabolite repression of phenol degradation in Pseudomonas putida is mediated by the inhibition of the activator protein PhlR. J Bacteriol 178:2030–2036
Neujahr HY, Gaal A (1973) Phenol hydroxylase from yeast. Purification and properties of the enzyme from Trichosporon cutaneum. Eur J Biochem 35:386–400
Nichols NN, Harwood CS (1995) Repression of 4-hydroxybenzoate transport and degradation by benzoate: a new layer of regulatory control in the Pseudomonas putida β-ketoadipate pathway. J Bacteriol 177:7033–7040
Omokoko B, Jäntges UK, Zimmermann M, Reiss M, Hartmeier W (2008) Isolation of the phe-operon from G. stearothermophilus comprising the phenol degradative meta-pathway genes and a novel transcriptional regulator. BMC Microbiol 8:197
Paisio CE, Talano MA, Gonzalez PS, Busto VD, Talou JR, Agostini E (2012) Isolation and characterization of a Rhodococcus strain with phenol-degrading ability and its potential use for tannery effluent biotreatment. Environ Sci Pollut Res Int 19:3430–3439
Pátek M, Muth G, Wohlleben W (2003) Function of Corynebacterium glutamicum promoters in Escherichia coli, Streptomyces lividans, and Bacillus subtilis. J Biotechnol 104:325–334
Prieto MB, Hidalgo A, Rodriguez-Fernandez C, Serra JL, Llama MJ (2002) Biodegradation of phenol in synthetic and industrial wastewater by Rhodococcus erythropolis UPV-1 immobilized in an air-stirred reactor with clarifier. Appl Microbiol Biotechnol 58:853–859
Putrinš M, Tover A, Tegova R, Saks Ü, Kivisaar M (2007) Study of factors which negatively affect expression of the phenol degradation operon pheBA in Pseudomonas putida. Microbiology 153:1860–1871
Rojo F (2010) Carbon catabolite repression in Pseudomonas: optimizing metabolic versatility and interactions with the environment. FEMS Microbiol Rev 34:658–684
Saa L, Jaureguibeitia A, Largo E, Llama MJ, Serra JL (2009) Cloning, purification and characterization of two components of phenol hydroxylase from Rhodococcus erythropolis UPV-1. Appl Microbiol Biotechnol 86:201–211
Sambrook J, Russel DV (2001) Molecular cloning. A laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Schleif R (2010) AraC protein, regulation of the L-arabinose operon in Escherichia coli, and the light switch mechanism of AraC action. FEMS Microbiol Rev 34:779–796
Shingler V (2003) Integrated regulation in response to aromatic compounds: from signal sensing to attractive behaviour. Environ Microbiol 5:1226–1241
Shingler V, Bartilson M, Moore T (1993) Cloning and nucleotide sequence of the gene encoding the positive regulator (DmpR) of the phenol catabolic pathway encoded by pVI150 and identification of DmpR as a member of the NtrC family of transcriptional activators. J Bacteriol 175:1596–1604
Takeo M, Murakami M, Niihara S, Yamamoto K, Nishimura M, Kato D, Negoro S (2008) Mechanism of 4-nitrophenol oxidation in Rhodococcus sp. Strain PN1: characterization of the two-component 4-nitrophenol hydroxylase and regulation of its expression. J Bacteriol 190:7367–7374
Teramoto M, Harayama S, Watanabe K (2001) PhcS represses gratuitous expression of phenol-metabolizing enzymes in Comamonas testosteroni R5. J Bacteriol 183:4227–4234
Tomás-Gallardo L, Santero E, Floriano B (2012) Involvement of a putative cyclic AMP receptor protein (CRP)-like binding sequence and a CRP-like protein in glucose-mediated catabolite repression of thn genes in Rhodococcus sp. strain TFB. Appl Environ Microbiol 78:5460–5462
van der Geize R, Dijkhuizen L (2004) Harnessing the catabolic diversity of rhodococci for environmental and biotechnological applications. Curr Opin Microbiol 7:255–261
Veselý M, Pátek M, Nešvera J, Čejková A, Masák J, Jirků V (2003) Host–vector system for phenol-degrading Rhodococcus erythropolis based on Corynebacterium plasmids. Appl Microbiol Biotechnol 61:523–527
Veselý M, Knoppová M, Nešvera J, Pátek M (2007) Analysis of catRABC operon for catechol degradation from phenol-degrading Rhodococcus erythropolis. Appl Microbiol Biotechnol 76:159–168
Yu H, Peng Z, Zhan Y, Wang J, Yan Y, Chen M, Lu W, Ping S, Zhang W, Zhao Z, Li S, Takeo M, Lin M (2011) Novel regulator MphX represses activation of phenol hydroxylase genes caused by a XylR/DmpR-type regulator MphR in Acinetobacter calcoaceticus. PLoS ONE 6:e17350
Zídková L, Szököl J, Rucká l, Pátek M, Nešvera J (2013) Biodegradation of phenol using recombinant plasmid-carrying Rhodococcus erythropolis strains. Int Biodeter Biodegradation 84:179–184
Acknowledgments
This work was supported by a grant (2B08062 AROMAGEN) from the Czech Ministry of Education, Youth and Sports, grant P504/11/0394 from Czech Science Foundation and internal project RVO61388971 (Institute of Microbiology).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Szőköl, J., Rucká, L., Šimčíková, M. et al. Induction and carbon catabolite repression of phenol degradation genes in Rhodococcus erythropolis and Rhodococcus jostii . Appl Microbiol Biotechnol 98, 8267–8279 (2014). https://doi.org/10.1007/s00253-014-5881-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-5881-6