Abstract
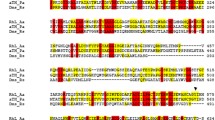
Phenol hydroxylase that catalyzes the conversion of phenol to catechol in Rhodococcus erythropolis UPV-1 was identified as a two-component flavin-dependent monooxygenase. The two proteins are encoded by the genes pheA1 and pheA2, located very closely in the genome. The sequenced pheA1 gene was composed of 1,629 bp encoding a protein of 542 amino acids, whereas the pheA2 gene consisted of 570 bp encoding a protein of 189 amino acids. The deduced amino acid sequences of both genes showed high homology with several two-component aromatic hydroxylases. The genes were cloned separately in cells of Escherichia coli M15 as hexahistidine-tagged proteins, and the recombinant proteins His6PheA1 and His6PheA2 were purified and its catalytic activity characterized. His6PheA1 exists as a homotetramer of four identical subunits of 62 kDa that has no phenol hydroxylase activity on its own. His6PheA2 is a homodimeric flavin reductase, consisting of two identical subunits of 22 kDa, that uses NAD(P)H in order to reduce flavin adenine dinucleotide (FAD), according to a random sequential kinetic mechanism. The reductase activity was strongly inhibited by thiol-blocking reagents. The hydroxylation of phenol in vitro requires the presence of both His6PheA1 and His6PheA2 components, in addition to NADH and FAD, but the physical interaction between the proteins is not necessary for the reaction.






Similar content being viewed by others
References
Abdurachim K, Ellis HR (2006) Detection of protein–protein interactions in the alkanesulfonate monooxygenase system from Escherichia coli. J Bacteriol 188:8153–8159
Annadurai G, Juang RS, Lee DJ (2002) Biodegradation and adsorption of phenol using activated carbon immobilized with Pseudomonas putida. J Environ Sci Health A Tox Hazard Subst Environ Eng 37:1133–1146
Ballou DP, Entsch B, Cole LJ (2005) Dynamics involved in catalysis by single-component and two-component flavin-dependent aromatic hydroxylases. Biochem Biophys Res Commun 338:590–598
Bateman A, Birney E, Cerruti L, Durbin R, Etwiller L, Eddy SR, Griffiths-Jones S, Howe KL, Marshall M, Sonnhammer ELL (2002) The Pfam protein families database. Nucleic Acids Res 30:276–280
Becker D, Schrader T, Andreesen JR (1997) Two-component flavin-dependent pyrrole-2-carboxylate monooxygenase from Rhodococcus sp. Eur J Biochem 249:739–747
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein–dye binding. Anal Biochem 72:248–254
Coves J, Niviere V, Eschenbrenner M, Fontecave M (1993) NADPH-sulfite reductase from Escherichia coli. A flavin reductase participating in the generation of the free radical of ribonucleotide reductase. J Biol Chem 268:18604–18609
Chaiyen P, Suadee C, Wilairat P (2001) A novel two-protein component flavoprotein hydroxylase. Eur J Biochem 268:5550–5561
Duffner FM, Muller R (1998) A novel phenol hydroxylase and catechol 2,3-dioxygenase from the thermophilic Bacillus thermoleovorans strain A2: nucleotide sequence and analysis of the genes. FEMS Microbiol Lett 161:37–45
Duffner FM, Kirchner U, Bauer MP, Muller R (2000) Phenol/cresol degradation by the thermophilic Bacillus thermoglucosidasius A7: cloning and sequence analysis of five genes involved in the pathway. Gene 256:215–221
Dunaway-Mariano D, Babbitt PC (1994) On the origins and functions of the enzymes of the 4-chlorobenzoate to 4-hydroxybenzoate converting pathway. Biodegradation 5:259–276
Eichhorn E, van der Ploeg JR, Leisinger T (1999) Characterization of a two-component alkanesulfonate monooxygenase from Escherichia coli. J Biol Chem 274:26639–26646
Galán B, Díaz E, Prieto MA, García JL (2000) Functional analysis of the small component of the 4-hydroxyphenylacetate 3-monooxygenase of Escherichia coli W: a prototype of a new Flavin:NAD(P)H reductase subfamily. J Bacteriol 182:627–636
Gisi MR, Xun L (2003) Characterization of chlorophenol 4-monooxygenase (TftD) and NADH:flavin adenine dinucleotide oxidoreductase (TftC) of Burkholderia cepacia AC1100. J Bacteriol 185:2786–2792
Harwood CS, Parales RE (1996) The beta-ketoadipate pathway and the biology of self-identity. Annu Rev Microbiol 50:553–590
Hidalgo A, Jaureguibeitia A, Prieto MB, Rodríguez-Fernández C, Serra JL, Llama MJ (2002a) Biological treatment of phenolic industrial wastewaters by Rhodococcus erythropolis UPV-1. Enzyme Microb Technol 31:221–226
Hidalgo A, Lopategi A, Prieto M, Serra JL, Llama MJ (2002b) Formaldehyde removal in synthetic and industrial wastewater by Rhodococcus erythropolis UPV-1. Appl Microbiol Biotechnol 58:260–263
Hubner A, Danganan CE, Xun L, Chakrabarty AM, Hendrickson W (1998) Genes for 2,4,5-trichlorophenoxyacetic acid metabolism in Burkholderia cepacia AC1100: characterization of the tftC and tftD genes and locations of the tft operons on multiple replicons. Appl Environ Microbiol 64:2086–2093
Ingelman M, Ramaswamy S, Niviere V, Fontecave M, Eklund H (1999) Crystal structure of NAD(P)H:flavin oxidoreductase from Escherichia coli. Biochemistry 38:7040–7049
Jaureguibeitia A, Saa L, Llama MJ, Serra JL (2007) Purification, characterization and cloning of aldehyde dehydrogenase from Rhodococcus erythropolis UPV-1. Appl Microbiol Biotechnol 73:1073–1086
Kadiyala V, Spain JC (1998) A two-component monooxygenase catalyzes both the hydroxylation of p-nitrophenol and the oxidative release of nitrite from 4-nitrocatechol in Bacillus sphaericus JS905. Appl Environ Microbiol 64:2479–2484
Kirchner U, Westphal AH, Muller R, van Berkel WJ (2003) Phenol hydroxylase from Bacillus thermoglucosidasius A7, a two-protein component monooxygenase with a dual role for FAD. J Biol Chem 278:47545–47553
Kulkarni M, Chaudhari A (2007) Microbial remediation of nitro-aromatic compounds: an overview. J Environ Manage 85:496–512
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–683
Lee JK, Zhao H (2007) Identification and characterization of the flavin:NADH reductase (PrnF) involved in a novel two-component arylamine oxygenase. J Bacteriol 189:8556–8563
Louie TM, Webster CM, Xun L (2002) Genetic and biochemical characterization of a 2,4,6-trichlorophenol degradation pathway in Ralstonia eutropha JMP134. J Bacteriol 184:3492–3500
Massey V (1994) Activation of molecular oxygen by flavins and flavoproteins. J Biol Chem 269:22459–22462
Moonen MJH, Fraaije MW, Rietjens IMCM, Laane C, van Berkel WJH (2002) Flavoenzyme-catalyzed oxygenations and oxidations of phenolic compounds. Adv Synth Catal 344:1023–1035
Neves LC, Miyamura TT, Moraes DA, Penna TC, Converti A (2006) Biofiltration methods for the removal of phenolic residues. Appl Biochem Biotechnol 129–132:130–152
Ohshiro T, Yamada H, Shimoda T, Matsubara T, Izumi Y (2004) Thermostable flavin reductase that couples with dibenzothiophene monooxygenase, from thermophilic Bacillus sp. DSM411: purification, characterization, and gene cloning. Biosci Biotechnol Biochem 68:1712–1721
Otto K, Hofstetter K, Rothlisberger M, Witholt B, Schmid A (2004) Biochemical characterization of StyAB from Pseudomonas sp. strain VLB120 as a two-component flavin-diffusible monooxygenase. J Bacteriol 186:5292–5302
Prieto MA, García JL (1994) Molecular characterization of 4-hydroxyphenylacetate 3-hydroxylase of Escherichia coli. A two-protein component enzyme. J Biol Chem 269:22823–22829
Prieto MA, Díaz E, García JL (1996) Molecular characterization of the 4-hydroxyphenylacetate catabolic pathway of Escherichia coli W: engineering a mobile aromatic degradative cluster. J Bacteriol 178:111–120
Prieto MB, Hidalgo A, Rodríguez-Fernández C, Serra JL, Llama MJ (2002a) Biodegradation of phenol in synthetic and industrial wastewater by Rhodococcus erythropolis UPV-1 immobilized in an air-stirred reactor with clarifier. Appl Microbiol Biotechnol 58:853–859
Prieto MB, Hidalgo A, Serra JL, Llama MJ (2002b) Degradation of phenol by Rhodococcus erythropolis UPV-1 immobilized on Biolite in a packed-bed reactor. J Biotechnol 97:1–11
Rajkumar D, Palanivelu K (2004) Electrochemical treatment of industrial wastewater. J Hazard Mater 113:123–129
Shevchenko A, Wilm M, Vorm O, Mann M (1996) Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal Chem 68:850–858
Smith MR (1990) The biodegradation of aromatic hydrocarbons by bacteria. Biodegradation 1:191–206
Sucharitakul J, Phongsak T, Entsch B, Svasti J, Chaiyen P, Ballou DP (2007) Kinetics of a two-component p-hydroxyphenylacetate hydroxylase explain how reduced flavin is transferred from the reductase to the oxygenase. Biochemistry 46:8611–8623
Takeo M, Yasukawa T, Abe Y, Niihara S, Maeda Y, Negoro S (2003) Cloning and characterization of a 4-nitrophenol hydroxylase gene cluster from Rhodococcus sp. PN1. J Biosci Bioeng 95:139–145
Thotsaporn K, Sucharitakul J, Wongratana J, Suadee C, Chaiyen P (2004) Cloning and expression of p-hydroxyphenylacetate 3-hydroxylase from Acinetobacter baumannii: evidence of the divergence of enzymes in the class of two-protein component aromatic hydroxylases. Biochim Biophys Acta 1680:60–66
Valton J, Mathevon C, Fontecave M, Niviere V, Ballou DP (2008) Mechanism and regulation of the two-component FMN-dependent monooxygenase ActVA–ActVB from Streptomyces coelicolor. J Biol Chem 283:10287–10296
van Berkel WJ, Kamerbeek NM, Fraaije MW (2006) Flavoprotein monooxygenases, a diverse class of oxidative biocatalysts. J Biotechnol 124:670–689
Vesely M, Knoppova M, Nesvera J, Patek M (2007) Analysis of catRABC operon for catechol degradation from phenol-degrading Rhodococcus erythropolis. Appl Microbiol Biotechnol 76:159–168
Witschel M, Nagel S, Egli T (1997) Identification and characterization of the two-enzyme system catalyzing oxidation of EDTA in the EDTA-degrading bacterial strain DSM 9103. J Bacteriol 179:6937–6943
Xi L, Squires CH, Monticello DJ, Childs JD (1997) A flavin reductase stimulates DszA and DszC proteins of Rhodococcus erythropolis IGTS8 in vitro. Biochem Biophys Res Commun 230:73–75
Xu Y, Mortimer MW, Fisher TS, Kahn ML, Brockman FJ, Xun L (1997) Cloning, sequencing, and analysis of a gene cluster from Chelatobacter heintzii ATCC 29600 encoding nitrilotriacetate monooxygenase and NADH:flavin mononucleotide oxidoreductase. J Bacteriol 179:1112–1116
Xun L (1996) Purification and characterization of chlorophenol 4-monooxygenase from Burkholderia cepacia AC1100. J Bacteriol 178:2645–2649
Xun L, Sandvik ER (2000) Characterization of 4-hydroxyphenylacetate 3-hydroxylase (HpaB) of Escherichia coli as a reduced flavin adenine dinucleotide-utilizing monooxygenase. Appl Environ Microbiol 66:481–486
Acknowledgements
This work was supported by grants from the University of the Basque Country (UPV 42.310-13529/2001 and 42.310-15955/2004). A.J. and L.S. were the recipients of scholarships from the Spanish Ministry of Education and the University of the Basque Country, respectively.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Saa, L., Jaureguibeitia, A., Largo, E. et al. Cloning, purification and characterization of two components of phenol hydroxylase from Rhodococcus erythropolis UPV-1. Appl Microbiol Biotechnol 86, 201–211 (2010). https://doi.org/10.1007/s00253-009-2251-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-009-2251-x