Abstract
Recombinant polyesterase (Est119) from Thermobifida alba AHK119 was purified by two chromatography steps. The final protein was observed as a single band in SDS–PAGE, and the specific activity of Est119 for p-nitrophenyl butyrate was 2.30 u/mg. Purified Est119 was active with aliphatic and aliphatic-co-aromatic polyesters. Kinetic data indicated that p-nitrophenyl butyrate (pNPB) or hexanoate was the best substrate for Est119 among p-nitrophenyl acyl esters. Calcium was required for full activity and thermostability of Est119, which was stable at 50 °C for 16 h. Three-dimensional modeling and biochemical characterization showed that Est119 is a typical cutinase-type enzyme that has the compact ternary structure of an α/β-hydrolase. Random and site-directed mutagenesis of wild-type Est119 resulted in improved activity with increased hydrophobic interaction between the antiparallel first and second β-sheets (A68V had the greatest effect). Introduction of a proline residue (S219P) in a predicted substrate-docking loop increased the thermostability. The specific activity of the A68V/S219P mutant on pNPB was increased by more than 50-fold over the wild type. The mutant was further activated by 2.6-fold (299 u/mg) with 300 mM Ca2+ and was stable up to 60 °C with 150 mM Ca2+. Another identical gene was located in tandem in the upstream of est119.






Similar content being viewed by others
References
Amada K, Kwon HJ, Haruki M, Morikawa M, Kanaya S (2001) Ca2+-induced folding of a family 1.3 lipase with repetitive Ca2+ binding motifs at the C-terminus. FEBS Lett 509:17–21
Angkawidjaja C, You DJ, Matsumura H, Kuwahara K, Koga Y, Takano K, Kanaya S (2007) Crystal structure of a family I.3 lipase from Pseudomonas sp. MIS38 in a closed conformation. FEBS Lett 581:5060–5064
Arpigny JL, Jäger K (1999) Bacterial lipolytic enzymes: classification and properties. Biochem J 343:177–183
Bellia G, Tosin M, Floridi G, Degli-Innocenti F (1999) Activated vermiculite, a solid bed for testing biodegradability under composting conditions. Polym Degrad Stab 66:65–79
Bornscheuer UT (2002) Microbial carboxyl esterases: classification, properties and application in biocatalysis. FEMS Microbiol Rev 26:73–81
Calabia BP, Tokiwa Y (2004) Microbial degradation of poly(D-3-hydroxybutyrate) by a new thermophilic Streptomyces isolate. Biotechnol Lett 26:15–19
Carvalho CML, Aires-Barros MR, Cabral JMS (1999) Cutinase: from molecular level to bioprocess development. Biotechnol Bioeng 66:17–34
Chen S, Tong X, Woodard RW, Du G, Wu J, Chen J (2008) Identification and characterization of bacterial cutinase. J Biol Chem 283:25854–25862
Dresler K, Heuvel J, Müller RJ, Deckwer WD (2006) Production of a recombinant polyester-cleaving hydrolase from Thermobifida fusca in Escherichia coli. Bioprocess Biosyst Eng 29:169–183
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In: Walker JM (ed) The proteomics protocols handbook. Humana, Totowa, pp 571–607
Hoang KC, Tseng M, Shu WJ (2007) Degradation of polyethylene succinate (PES) by a new thermophilic Microbispora strain. Biodegradation 18:333–342
Hofer P, Fringeli US, Hopff WH (1984) Activation of acetylcholinesterase by monovalent (Na+, K+) and divalent (Ca2+, Mg2+) cations. Biochemistry 23:2730–2734
Hu X, Osaki S, Hayashi M, Kaku M, Katuen S, Kobayashi H, Kawai F (2008) Degradation of a terephthalate-containing polyester by thermophilic actinomycetes and Bacillus species derived from composts. J Polymer Environ 16:103–108
Hu X, Thumarat U, Zhang X, Tang M, Kawai F (2010) Diversity of polyester-degrading bacteria in compost and molecular analysis of a thermoactive esterase from Thermobifida alba AHK119. Appl Microbiol Biotechnol 87:771–779
Kawai F (2010) The biochemistry and molecular biology of xenobiotic polymer degradation by microorganisms. Biosci Biotechnol Biochem 74:1743–1759
Kawai F, Nakadai K, Nishioka E, Nakajima H, Ohara H, Masaki K, Iefuji H (2011) Different enantioselectivity of two types of poly(lactic acid) depolymerases toward poly(l-lactic acid) and poly(d-lactic acid). Polym Degrad Stab 96:1342–1348
Kim MH, Kim HK, Lee JK, Park SY, Oh TK (2000) Thermostable lipase of Bacillus stearothermophilus: high-level production, purification, and calcium-dependent thermostability. Biosci Biotechnol Biochem 64:280–286
Kleeberg I, Hetz C, Kroppenstedt RM, Müller RJ, Deckwer WD (1998) Biodegradation of aliphatic-aromatic copolyester-by Thermomonospora fusca and other thermophillic compost isolates. Appl Environ Microbiol 64:1731–1735
Kleeberg I, Welzel K, VandenHeuvel J, Müller RJ, Deckwer WD (2005) Characterization of a new extracellular hydrolase from Thermobifida fusca degrading aliphatic-aromatic copolyesters. Biomacromolecules 6:262–270
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Longieras A, Copinet A, Bureau G, Tighzert L (2004) An inert solid medium for simulation of material in compost carbon balance achievement. Polym Degrad Stab 83:187–194
Lykidis A, Mavromatis K, Ivanova I, Anderson I, Land M, DiBartolo G, Martinez M, Lapidus A, Lucas S, Copeland A, Richardson P, Wilson DB, Kyrpides N (2007) Genome sequence and analysis of the soil cellulolytic actinomycete Thermobifida fusca YX. J Bacteriol 189:2477–2486
Maeda H, Yamagata Y, Abe K, Hasegawa F, Machida M, Ishioka R, Gomi K, Nakajima T (2005) Purification and characterization of a biodegradable plastic-degrading enzyme from Aspergillus oryzae. Appl Microbiol Biotechnol 67:778–788
Ma X, Wang ZS, Li SX, Shen Q, Yuan QS (2006) Effect of CaCl2 as activity stabilizer on purification of heparinase I from Flavobacterium heparinum. J Chromatogr B 843:209–215
Nardini M, Dijkstra BW (1999) α/β Hydrolase fold enzymes: the family keeps growing. Curr Opin Struct Biol 9:732–737
Niu WN, Li ZP, Zhang DW, Yu MR, Tan TW (2006) Improved thermostability and the optimum temperature of Rhizopus arrhizus lipase by directed evolution. J Mol Catal B: Enzym 43:33–39
Ollis DL, Cheahm E, Cygler M, Dijkstra B, Frolow F, Franken SM, Harel M, Remington SJ, Silman I, Schrag J, Sussman JL, Verschueren KHG, Goldma A (1992) The α/β hydrolase fold. Protein Eng 5:197–211
Pio TF, Macedo GA (2009) Cutinases: properties and industrial applications. Adv Appl Microbiol 66:77–95
Phithakrotchanakoon C, Rudeekit Y, Tanapongpipat S, Leejakpai T, Aiba S, Noda I, Champreda V (2009) Microbial degradation and physico-chemical alteration of polyhydroxyalkanoates by a thermophilic Streptomyces sp. Biologia 64:246–251
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor
Simons JFA, Van Kampen MD, Ubarretxena-Belandia I, Cox RC, Alves dos Santos CM, Egmond MR, Verheij HM (1999) Identification of a calcium binding site in Staphylococcus hyicus lipase: generation of calcium-independent variants. Biochemistry 38:2–10
Sinsereekul N, Wangkam T, Thamchaipenet A, Srikhirin T, Eurwilaichitr L, Champreda V (2010) Recombinant expression of BTA hydrolase in Streptomyces rimosus and catalytic analysis on polyesters by surface plasmone resonance. Appl Microbiol Biotechnol 86:1775–1784
Suzuki Y, Hatagaki K, Oda H (1991) A hyperthermostable pullulanase produced by an extreme thermophile, Bacillus flavocaldarius KP 1228, and evidence for the proline theory of increasing protein thermostability. Appl Microbiol Biotechnol 34:707–714
Tsutsumi M, Otaki JM (2011) Parallel and antiparallel β-strands differ in amino acid composition and availability of short constituent sequences. J Chem Inf Model Article ASAP (DOI: 10.1021/ci200027d. Publication Date (Web): April 26, 2011)
Vieille C, Zeikus GJ (2001) Hyperthermophilic enzymes: sources, uses, and molecular mechanisms for thermostability. Microbiol Mol Biol Rev 65:1–43
Wei Y, Swenson L, Castro C, Derewenda U, Minor W, Arai H, Aoki J, Inoue K, Servin-Gonzalez L, Derewenda ZS (1988) Structure of a microbial homologue of mammalian platelet-activating factor acetylhydrolases: Streptomyces exfoliatus lipase at 1.9°A resolution. Structure 6:511–519
Yang Y, Malten M, Grote A, Jahn D, Deckwer W-D (2007) Codon optimized Thermobifida fusca hydrolase secreted by Bacillus megaterium. Biotechnol Bioeng 96:780–794
Acknowledgements
The study was supported by funds from the Institute for Fermentation, Osaka (Japan) to F. K. We appreciate Prof. Y. Yamamoto, Institute for Plant Resources, Okayama University, for her kind help in atomic absorption spectrometry measurements. We are grateful to Dr. T. Nakajima-Kambe, University of Tsukuba, for his kind supply of PBSA (Bionolle™EM-301). This work is linked with the Asia Core Program supported by JSPS (Japan) and NRCT (Thailand).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
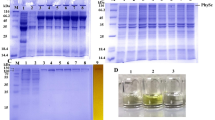
Fig. 1
SDS-PAGE of Est119 purified by one-step and two-step chromatographies. 1, protein standards; 2, crude cell lysate from E. coli Rosetta-gami B (DE3) harboring pQE80L-est119 induced with 0.1 mM IPTG; 3, Purified Est119 by one-step chromatography on a Ni-Sepharose 6 Fast Flow column; 4, Purified Est119 by the second step chromatography on a Mono Q anion exchange column. (DOC 300 kb)
Rights and permissions
About this article
Cite this article
Thumarat, U., Nakamura, R., Kawabata, T. et al. Biochemical and genetic analysis of a cutinase-type polyesterase from a thermophilic Thermobifida alba AHK119. Appl Microbiol Biotechnol 95, 419–430 (2012). https://doi.org/10.1007/s00253-011-3781-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-011-3781-6