Abstract
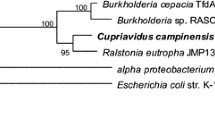
Ferulic acid decarboxylase (FADase) can catalyze the transformation of ferulic acid into 4-vinyl guaiacol via decarboxylation in microorganisms. In this study, a gene encoding FADase was first isolated from the bacterium Enterobacter sp. Px6-4 using degenerate primers and a genome walking technique. The putative encoding gene (fad) of FADase consists of 507-bp nucleotides, coding a polypeptide of 168 amino acid residues. In addition, a putative gene encoding the transcriptional regulator was identified from the upstream of the fad gene. The deduced peptide sequence of the FADase from Enterobacter sp. Px6-4 showed a 51.2–53.3% sequence identity to decarboxylases from other bacteria. The gene fad was successfully expressed in Escherichia coli BL21, and the recombinant FADase was purified as a protein of ca. 23 kDa with an optimal activity at pH 4.0 and 28 °C. The purified FADase could convert ferulic acid to 4-vinyl guaiacol effectively, and its hydrolytic activity could be inhibited by Cu2+ (99%) and Hg2+ (99.5%). A phylogenetic analysis of the FADase protein from bacteria revealed several different clades. Our result provided a basis for further studies of the ferulic acid transformation pathway and for enhanced production of vanillin in the future.







Similar content being viewed by others
References
Akin DE (1988) Biological structure of lignocellulose and its degradation in the rumen. Anim Food Sci Technol 21:295–310
Barthelmebs L, Lecomte B, Divies C, Cavin JF (2000) Inducible metabolism of phenolic acids in Pediococcus pentosaceus is encoded by an autoregulated operon which involves a new class of negative transcriptional regulator. J Bacteriol 182:6724–6731
Barthelmebs L, Diviès C, Cavin JF (2001) Expression in Escherichia coli of native and chimeric phenolic acid decarboxylases with modified enzymatic activities and method for screening recombinant E. coli strains expressing these enzymes. Appl Environ Microbiol 67:1063–1069
Benoit I, Danchin EG, Bleichrodt RJ, de Vries RP (2008) Biotechnological applications and potential of fungal feruloyl esterases based on prevalence, classification and biochemical diversity. Biotechnol Lett 30:387–396
Bradford MM (1976) A rapid and sensitive method for quantitation of microgram quantities of protein utilizing the principle of protein-dye-binding. Anal Biochem 72:248–254
Brezillon C, Kroon PA, Faulds CB, Brett GM, Williamson G (1996) Novel ferulic acid esterases are induced by growth of Aspergillus niger on sugar beet pulp. Appl Microbiol Biotechnol 45:371–376
Brunati M, Marinelli F, Bertolini C, Gandolfi R, Daffonchio D, Molinari F (2004) Biotransformations of cinnamic and ferulic acid with actinomycetes. Enzyme Microb Technol 34:3–9
Cavin JF, Barthelmebs L, Guzzo J, Beeumen JV, Samyn B, Travers JF, Diviès C (1997a) Purification and characterization of an inducible p-coumaric acid decarboxylase from Lactobacillus plantarum. FEMS Microbiol Lett 147:291–295
Cavin JF, Barthelmebs L, Diviès C (1997b) Molecular characterization of an inducible p-coumaric acid decarboxylase from Lactobacillus plantarum: gene cloning, transcriptional analysis, overexpression in Escherichia coli, purification and characterization. Appl Environ Microbiol 63:1939–1944
Cavin JF, Dartois V, Diviès C (1998) Gene cloning, transcriptional analysis, purification and characterization of phenolic acid decarboxylase from Bacillus subtilis. Appl Environ Microbiol 64:1466–1471
Degrassi G, De Laureto PP, Bruschi CV (1995) Purification and characterization of ferulate and p-coumarate decarboxylase from Bacillus pumilus. Appl Environ Microbiol 61:326–332
Donaghy J, McKay AM (1997) Purification and characterization of a feruloyl esterase from the fungus, Penicillium expansum. J Appl Microbiol 83:718–726
Edlin DAN, Narbad A, Gasson MJ, Dickinson JR, Lloyd D (1998) Purification and characterization of hydroxycinnamate decarboxylase from Brettanomyces anomalus. Enzyme Microbiol Technol 22:232–239
Gury J, Barthelmebs L, Tran NP, Divies C, Cavin JF (2004) Cloning, deletion, and characterization of PadR, the transcriptional repressor of the phenolic acid decarboxylase-encoding padA gene of Lactobacillus plantarum. Appl Environ Microbiol 70:2146–2153
Huang ZX, Dostal L, Rosazza JPN (1993) Microbial transformations of ferulic acid by Saccharomyces cerevisiae and Pseudomonas fluorescens. Appl Environ Microbiol 59:2244–2250
Huang Z, Dostal L, Rosazza JPN (1994) Purification and characterization of a ferulic acid decarboxylase from Pseudomonas fluorescens. J Bacteriol 176:5912–5918
Ishii T (1997) Structure and function of feruloylated polysaccharides. Plant Sci 127:111–127
Li XM, Yang JK, Li X, Gu W, Huang JW, Zhang KQ (2008) The metabolism of ferulic acid via 4-vinylguaiacol to vanillin by Enterobacter sp. Px6-4 isolated from Vanilla root. Process Biochem 43:1032–1037
Mathew S, Abraham TE (2004) Ferulic acid: an antioxidant found naturally in plant cell walls and feruloyl esterases involved in its release and their applications. Crit Rev Biotechnol 24:59–83
Mathew S, Abraham TE (2006) Bioconversions of ferulic acid, an hydroxycinnamic acid. Crit Rev Microbiol 32:115–125
Muller-Harvey I, Hartley RD (1986) Linkage of p-coumaroyl and feruloyl groups to cell-wall polysaccharides of barley straw. Carbohydr Res 148:71–85
Oosterveld A, Beldman G, Schols HA, Voragen AGJ (2000) Characterization of arabinose and ferulic acid rich pectic polysaccharides and hemicelluloses from sugar beet pulp. Carbohydr Res 328:185–197
Priefert H, Rabenhorst J, Steinbüchel A (2001) Biotechnological production of vanillin. Appl Microbiol Biotechnol 56:296–314
Prim N, Pastor FI, Diaz P (2003) Biochemical studies on cloned Bacillus sp. BP-7 phenolic acid decarboxylase PadA. Appl Microbiol Biotechnol 63:51–56
Ramachandra RS, Ravishankar GA (2000) Vanilla flavour: production by conventional and biotechnological routes. J Sci Food Agric 80:289–304
Rosazza JPN, Huang Z, Dostal L, Volm T, Rousseau B (1995) Biocatalytic transformations of ferulic acid: an abundant aromatic natural product. J Ind Microbiol 15:457–471
Saulnier L, Thibault JF (1999) Ferulic acid and diferulic acids as components of sugar-beet pectins and maize bran heteroxylans. J Sci Food Agric 79:396–402
Smith MM, Hartley RD (1983) Occurrence and nature of ferulic acid substitution of cell-wall polysaccharides in graminaceous plants. Carbohydr Res 118:65–80
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal_X Windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–4882
Torres JL, Rosazza JPN (2001) Microbial transformation of p-coumaric acid by Bacillus megaterium and Curvularia lunata. J Nat Prod 64:1408–1414
Yang JK, Huang XW, Tian BY, Wang M, Niu QH, Zhang KQ (2005) Isolation and characterization of a serine protease from the nematophagous fungus Lecanicillium psalliotae, displaying nematicidal activity. Biotechnol Lett 27:1123–1128
Zago A, Degrassi G, Bruschi CV (1995) Cloning, sequencing, and expression in Escherichia coli of the Bacillus pumilus gene for ferulic acid decarboxylase. Appl Environ Microbiol 61:4484–4486
Acknowledgements
This work was funded by the National Natural Science Foundation of China (Approval No. 31060012) and the Department of Science and Technology of Yunnan Province (Approval Nos. 2007C170M, 2006GG22, and 2009CI052).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gu, W., Li, X., Huang, J. et al. Cloning, sequencing, and overexpression in Escherichia coli of the Enterobacter sp. Px6-4 gene for ferulic acid decarboxylase. Appl Microbiol Biotechnol 89, 1797–1805 (2011). https://doi.org/10.1007/s00253-010-2978-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-010-2978-4