Abstract
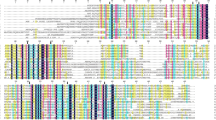
In the cyanobacterium Spirulina platensis, the desaturation process is carried out by three desaturases: the Δ9, Δ12 and Δ6 desaturases, encoded by desC, desA and desD, respectively. The Δ6 desaturase is responsible for the catalysis of linoleic acid, yielding γ-linolenic acid (18:3Δ9,12,6), the end-product of the process. In this study, the desD gene was expressed in Escherichia coli using a pTrcHisA expression system. In order to identify the amino acid residues involved in the enzymatic activity, a sequence comparison was performed using various organisms. The alignment revealed three conserved histidine clusters, a number of conserved residues among all listed organisms and a few conserved residues among cyanobacterial species possibly involved in the desaturation activity. A series of site-directed mutations were generated in the desD gene to evaluate the role of these residues vis-à-vis the enzyme function. This approach revealed that: (1) H313 is involved in the regioselectivity of the enzyme, (2) the three histidine clusters together with H313, H315, D138 and E140 are required for enzymatic activity, most likely as providers of the catalytic Fe center and (3) W294 is also essential for the activity of Δ6 desaturase, possibly by forming part of the substrate-binding pocket.





Similar content being viewed by others
References
Avelange-Macherel M, Macherel D, Wada H, Murata N (1991) Site-directed mutagenesis of histidine residues in the delta 12 acyl-lipid desaturase of Synechocystis. FEBS 361:111–114
Bloomfield DK, Bloch K (1960) Formation of unsaturated fatty acids. J Biol Chem 235:337–345
Broadwater JA, Whittle E, Shanklin J (2002) Desaturation and hydroxylation; residue 148 and 324 of Arabidopsis FAD2, in addition to substrate chain length, exert a major influence in partitioning of catalytic specificity. Biol Chem 277:15613–15620
Cahoon EB, Mills LA, Shanklin J (1996) Modification of the fatty acid composition of Escherichia coli by coexpression of a plant acyl–acyl carrier protein desaturase and ferredoxin. J Bacteriol 178:936–939
Cahoon EB, Coughlan SJ, Shanklin J (1997) Characterization of a structurally and functionally diverged acyl–acyl carrier protein desaturase from milkweed seed. Plant Mol Biol 33:1105–1110
Cahoon EB, Lindqvist Y, Schneider G, Shanklin J (1997) Redesign of soluble fatty acid desaturases from plants for altered substrate specificity and double bond position. Proc Natl Acad Sci USA 94:4872–4877
Cahoon EB, Shah S, Shanklin J, Browse J (1998) A determinant of substrate specificity predicted from the acyl–acyl carrier protein desaturase of developing cat’s claw seed. Plant Physiol 117:593–598
Creighton TE (1993) Proteins: structures and molecular properties. Freeman, New York
Deng WP, Nickoloff JA (1992) Site directed mutagenesis of virtually any plasmid by eliminating a unique site. Anal Biochem 200:81–88
Deshnium P, Paithoonrangsarid K, Suphratrakul A, Meesapyodsuk D, Tanticharoen M, Cheevadhanarak S (2000) Temperature-independent and dependent expression of desaturase genes in filamentous cyanobacterium Spirulina platensis C1 (Arthrospira sp. PCC 9438). FEMS Microbiol Lett 184:207–213
Diaz AR, Mansilla MC, Vila AJ, Mendoza D (2002) Membrane topology of the acyl-lipid desaturase from Bacillus subtilis. Biol Chem 277:48099–48106
Esfahani M, Barnes EMJ, Wakil S (1969) Control of fatty acid composition in phospholipids of Escherichia coli: response to fatty acid supplements in a fatty acid auxotroph. Proc Natl Acad Sci USA 64:1057–1064
Fay L, Richli U (1991) Location of double bonds in polyunsaturated fatty acids by gas chromatography-mass spectrometry after 4,4-dimethyloxazoline derivatization. J Chromatogr 541:89–98
Gravel P, Golaz O (1996) Protein blotting by the semi-dry method. In: Walker JM (ed) The protein protocols handbook. Humana, New Jersey, pp 249–260
Holloway PW (1983) Fatty acid desaturation. In Boyer PD (ed) The enzymes, vol 16. Academic, Orlando, pp 63–83
Hongsthong A, Deshnium P, Paithoonrangsarid K, Cheevadhanarak S, Tanticharoen M (2003) Differential responses of the three acyl-lipid desaturases to immediate temperature reduction occurred in two lipid membranes of Spirulina platensis strain C1. J Biosci Bioeng 96:519–524
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193:269–275
Mcguire KA, Siggaard-Andersen M, Bangera MG, Olsen JG, Wettstein-Knowles P (2001) β-Ketoacyl-(acyl carrier protein) synthase I of Escherichia coli: aspects of the condensation mechanism revealed by analyses of mutations in the active site pocket. Biochemistry 40:9836–9845
Murata N, Nishida I (1987) Lipids of blue-green algae. In: Stumpf PK (ed) The biochemistry of plants, vol 9. Academic, Orlando, pp 315–347
Murata N, Wada H (1995) Acyl-lipid desaturases and their importance in the tolerance and acclimation to cold of cyanobacteria. Biochem J 308:1–8
Murata N, Deshnium P, Tasaka Y (1996) Biosynthesis of γ-linolenic acid in the cyanobacterium Spirulina platensis. In: Huang YS, Mills DE (eds) γ-Linolenic acid metabolism and its roles in nutrition and medicine. AOCS, Urbana-Champaign, pp 22–32
Mustady L, Los DA, Gombos Z, Murata N (1996) Immunocytochemical location of acyl-lipid desaturases in cyanobacterial cells: evidence that both thylakoid membranes and cytoplasmic membranes are the sites for lipid desaturation. Proc Natl Acad Sci USA 93:10524–10527
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
Schmidt H, Heinz E (1993) Direct desaturation of intact galactolipids by a desaturase solubilized from spinach (Spinacia oleracea) chloroplast envelopes. Biochem J 289:777–782
Schmidt H, Sperling P, Heinz E (1993) Biochemistry and molecular biology of membrane and storage lipids of plants. American Society for Plant Physiologists, Rockville
Schultz DJ, Chung Suh M, Ohlrogge JB (2000) Stearoyl–acyl carrier protein and unusual acyl–acyl carrier protein desaturase activities are differentially influences by ferredoxin. Plant Physiol 124:681–692
Shanklin J, Cahoon EB (1998) Desaturation and related modifications of fatty acids. Annu Rev Plant Physiol Plant Mol Biol 49:611–641
Shanklin J, Whittle E, Fox BG (1994) Eight histidine residues are catalytically essential in a membrane-associated iron enzyme, steroyl-CoA desaturase and are conserved in alkene hydroxylase and xylene monooxygenase. Biochemistry 33:12787–12794
Sperling P, Ternes P, Zank TK, Heinz E (2003) The evolution of desaturases. Prostaglandins Leukot Essent Fatty Acids 68:73–95
Sundberg RJ, Martin RB (1974) Interactions of histidine and other imidazole derivatives with transition metal ions in chemical and biological systems. Chem Rev 74:471–517
Ta TD, Vickery LE (1992) Cloning, sequencing and overexpression of a [2Fe–2S] ferredoxin gene from Escherichia coli. J Biol Chem 267:11120–11125
Tocher DR, Leaver MJ, Hodgson PA (1998) Recent advances in the biochemistry and molecular biology of fatty acyl desaturases. Prog Lipid Res 37:73–117
Wada H, Avelange-Macherel M, Murata N (1993) The desA gene of the cyanobacterium Synechocystis sp. strain PCC 6803 is the structural gene for Δ12 desaturase. J Bacteriol 175:6056–6058
Whittle E, Shanklin J (2001) Engineering delta9-16:0-acyl carrier protein (ACP) desaturase specificity based on combinatorial saturation mutagenesis and logical redesign of the castor delta9-18:0-ACP desaturase. Biol Chem 276:21500–21505
Zámocky M, Gunther R, Jakopitsch C, Obinger C (2001) The molecular peculiarities of catalase–peroxidase. FEBS 492:177–182
Acknowledgements
This research was funded by a grant from the National Center for Genetic Engineering and Biotechnology (BIOTEC), Bangkok, Thailand. The experiments comply with the current laws of Thailand.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hongsthong, A., Subudhi, S., Sirijuntarat, M. et al. Mutation study of conserved amino acid residues of Spirulina Δ6-acyl-lipid desaturase showing involvement of histidine 313 in the regioselectivity of the enzyme. Appl Microbiol Biotechnol 66, 74–84 (2004). https://doi.org/10.1007/s00253-004-1655-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-004-1655-x