Abstract
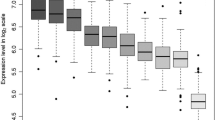
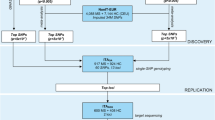
Multiple sclerosis (MS) is an inflammatory and demyelinating disease of central nervous system. Many genetic variants associated with MS have been identified by genome-wide association studies, but functional mechanism underlying the associations is largely unclear. Utilizing the publically available datasets, we carried out gene relationships among implicated loci (GRAIL) analyses to search for MS-associated SNPs/genes. Expression quantitative trait loci (eQTLs) analyses were conducted to identify eQTL SNPs/target genes. Further, functional prediction for SNP, differential gene expression, and functional annotation clustering analyses for gene were conducted to explore their functional relevance to MS. Among the 284 identified MS-associated SNPs (P < 10−4), eQTL analysis showed that 45 SNPs act as cis-effect regulators on 19 MS-associated genes. Among the 19 eQTL target genes, 14 showed significantly differential expressions in MS-related cells. Among the 45 SNPs, 15 were predicted most likely located in transcription factor (TF) binding sites, and five predicted SNPs (rs3095329 of TUBB, rs9469220/rs2647046 of HLA-DQB1, rs11154801 of AHI1, and rs1062158 of NDFIP1) have corresponding target genes with significantly differential expressions in multiple cell groups, while rs7194 of HLA-DRA was predicted in the has-miR-6507-3p binding site. The functional evidence, taken together, highlighted the functional relevance of the six SNPs to MS. The present findings provide novel insights into the functional mechanisms underlying the MS-associated genetic variants, which improve our understanding of the genetic association for MS.

Similar content being viewed by others
References
Annibali V et al (2011) CD161(high)CD8 + T cells bear pathogenetic potential in multiple sclerosis. Brain: J Neurol 134:542–554
Aulchenko YS et al (2008) Genetic variation in the KIF1B locus influences susceptibility to multiple sclerosis. Nat Genet 40:1402–1403
Australia, New Zealand Multiple Sclerosis Genetics C (2009) Genome-wide association study identifies new multiple sclerosis susceptibility loci on chromosomes 12 and 20. Nat Genet 41:824–828. doi:10.1038/ng.396
Bao L, Zhou M, Wu L, Lu L, Goldowitz D, Williams RW, Cui Y (2007) PolymiRTS database: linking polymorphisms in microRNA target sites with complex traits. Nucleic Acids Res 35:D51–D54. doi:10.1093/nar/gkl797
Baranzini SE (2011) Revealing the genetic basis of multiple sclerosis: are we there yet? Curr Opin Genet Dev 21:317–324. doi:10.1016/j.gde.2010.12.006
Baranzini SE et al (2009) Genome-wide association analysis of susceptibility and clinical phenotype in multiple sclerosis. Hum Mol Genet 18:767–778. doi:10.1093/hmg/ddn388
Baranzini SE et al (2010) Genetic variation influences glutamate concentrations in brains of patients with multiple sclerosis. Brain: J Neurol 133:2603–2611. doi:10.1093/brain/awq192
Basciano PA, Bussel J, Hafeez Z, Christos PJ, Giannakakou P (2013) The beta 1 tubulin R307H single nucleotide polymorphism is associated with treatment failures in immune thrombocytopenia (ITP). Br J Haematol 160:237–243
Boyle AP et al (2008) High-resolution mapping and characterization of open chromatin across the genome. Cell 132:311–322. doi:10.1016/j.cell.2007.12.014
Boyle AP et al (2012) Annotation of functional variation in personal genomes using RegulomeDB. Genome Res 22:1790–1797. doi:10.1101/gr.137323.112
Comabella M et al (2008) Identification of a novel risk locus for multiple sclerosis at 13q31.3 by a pooled genome-wide scan of 500,000 single nucleotide polymorphisms. PLoS One 3:e3490
Consoli L, Lefevre A, Zivy M, de Vienne D, Damerval C (2002) QTL analysis of proteome and transcriptome variations for dissecting the genetic architecture of complex traits in maize. Plant Mol Biol 48:575–581
Coppieters F et al (2010) Genetic screening of LCA in Belgium: predominance of CEP290 and identification of potential modifier alleles in AHI1 of CEP290-related phenotypes. Hum Mutat 31:E1709–E1766
da Huang W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57. doi:10.1038/nprot.2008.211
De Jager PL et al (2009) Meta-analysis of genome scans and replication identify CD6, IRF8 and TNFRSF1A as new multiple sclerosis susceptibility loci. Nat Genet 41:776–782
Dimas AS et al (2009) Common regulatory variation impacts gene expression in a cell type-dependent manner. Science 325:1246–1250. doi:10.1126/science.1174148
Fernando MM et al (2008) Defining the role of the MHC in autoimmunity: a review and pooled analysis. PLoS Genet 4:e1000024. doi:10.1371/journal.pgen.1000024
Gaffney DJ et al (2012) Dissecting the regulatory architecture of gene expression QTLs. Genome Biol 13:R7. doi:10.1186/gb-2012-13-1-r7
Hochberg Y (1988) A sharper Bonferroni procedure for multiple tests of significance. Biometrika 75:800–802
Huang L, Teng D, Wang H, Sheng G, Liu T (2012) Association of copy number variation in the AHI1 gene with risk of obesity in the Chinese population. Eur J Endocrinol / Eur Fed Endocr Soc 166:727–734
International Multiple Sclerosis Genetics C (2011) Genome-wide association study of severity in multiple sclerosis. Genes Immun 12:615–625. doi:10.1038/gene.2011.34
International Multiple Sclerosis Genetics C et al (2007) Risk alleles for multiple sclerosis identified by a genomewide study. N Engl J Med 357:851–862. doi:10.1056/NEJMoa073493
International Multiple Sclerosis Genetics C et al (2011) Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 476:214–219. doi:10.1038/nature10251
Jakkula E et al (2010) Genome-wide association study in a high-risk isolate for multiple sclerosis reveals associated variants in STAT3 gene. Am J Hum Genet 86:285–291
Jernas M, Malmestrom C, Axelsson M, Nookaew I, Wadenvik H, Lycke J, Olsson B (2013) MicroRNA regulate immune pathways in T-cells in multiple sclerosis (MS). BMC Immunol 14:32. doi:10.1186/1471-2172-14-32
Karchin R et al (2005) LS-SNP: large-scale annotation of coding non-synonymous SNPs based on multiple information sources. Bioinformatics 21:2814–2820. doi:10.1093/bioinformatics/bti442
Kemppinen AK, Kaprio J, Palotie A, Saarela J (2011) Systematic review of genome-wide expression studies in multiple sclerosis. BMJ Open 1:e000053. doi:10.1136/bmjopen-2011-000053
Kraft P, Zeggini E, Ioannidis JP (2009) Replication in genome-wide association studies. Stat Sci Rev J Inst Math Stat 24:561–573. doi:10.1214/09-STS290
Litvinov IV, Kupper TS, Sasseville D (2012) The role of AHI1 and CDKN1C in cutaneous T-cell lymphoma progression. Exp Dermatol 21:964–966
Lutterotti A et al (2012) No proinflammatory signature in CD34+ hematopoietic progenitor cells in multiple sclerosis patients. Mult Scler 18:1188–1192
Mangravite LM et al (2013) A statin-dependent QTL for GATM expression is associated with statin-induced myopathy. Nature 502:377–380. doi:10.1038/nature12508
Martinelli-Boneschi F et al (2012) A genome-wide association study in progressive multiple sclerosis. Mult Scler 18:1384–1394. doi:10.1177/1352458512439118
Michaelson JJ, Loguercio S, Beyer A (2009) Detection and interpretation of expression quantitative trait loci (eQTL). Methods 48:265–276. doi:10.1016/j.ymeth.2009.03.004
Montgomery SB et al (2010) Transcriptome genetics using second generation sequencing in a Caucasian population. Nature 464:773–777. doi:10.1038/nature08903
Nakahara J, Maeda M, Aiso S, Suzuki N (2012) Current concepts in multiple sclerosis: autoimmunity versus oligodendrogliopathy. Clin Rev Allergy Immunol 42:26–34. doi:10.1007/s12016-011-8287-6
Nischwitz S et al (2010) Evidence for VAV2 and ZNF433 as susceptibility genes for multiple sclerosis. J Neuroimmunol 227:162–166
Olivier M (2004) From SNPs to function: the effect of sequence variation on gene expression. Focus on “a survey of genetic and epigenetic variation affecting human gene expression”. Physiol Genomics 16:182–183
Pastinen T, Ge B, Hudson TJ (2006) Influence of human genome polymorphism on gene expression. Hum Mol Genet 15 Spec No 1:R9-16 doi:10.1093/hmg/ddl044
Patsopoulos NA et al (2011) Genome-wide meta-analysis identifies novel multiple sclerosis susceptibility. loci Annals Neurol 70:897–912. doi:10.1002/ana.22609
Pickrell JK et al (2010) Understanding mechanisms underlying human gene expression variation with RNA sequencing. Nature 464:768–772. doi:10.1038/nature08872
Ramon HE, Riling CR, Bradfield J, Yang B, Hakonarson H, Oliver PM (2011) The ubiquitin ligase adaptor Ndfip1 regulates T cell-mediated gastrointestinal inflammation and inflammatory bowel disease susceptibility. Mucosal Immunol 4:314–324
Ramos EM et al (2014) Phenotype-Genotype Integrator (PheGenI): synthesizing genome-wide association study (GWAS) data with existing genomic resources. Eur J Hum Genet: EJHG 22:144–147
Raychaudhuri S et al (2009) Identifying relationships among genomic disease regions: predicting genes at pathogenic SNP associations and rare deletions. PLoS Genet 5:e1000534. doi:10.1371/journal.pgen.1000534
Sale S et al (2002) Conservation of the class I beta-tubulin gene in human populations and lack of mutations in lung cancers and paclitaxel-resistant ovarian cancers. Mol Cancer Ther 1:215–225
Sang Q et al (2006) Nedd4-WW domain-binding protein 5 (Ndfip1) is associated with neuronal survival after acute cortical brain injury. J Neurosci: Off J Soc Neurosci 26:7234–7244
Sanna S et al (2010) Variants within the immunoregulatory CBLB gene are associated with multiple sclerosis. Nat Genet 42:495–497. doi:10.1038/ng.584
Schaub MA, Boyle AP, Kundaje A, Batzoglou S, Snyder M (2012) Linking disease associations with regulatory information in the human genome. Genome Res 22:1748–1759. doi:10.1101/gr.136127.111
Stranger BE et al (2007) Population genomics of human gene expression. Nat Genet 39:1217–1224. doi:10.1038/ng2142
Trynka G, Sandor C, Han B, Xu H, Stranger BE, Liu XS, Raychaudhuri S (2013) Chromatin marks identify critical cell types for fine mapping complex trait variants. Nat Genet 45:124–130. doi:10.1038/ng.2504
Veyrieras JB, Kudaravalli S, Kim SY, Dermitzakis ET, Gilad Y, Stephens M, Pritchard JK (2008) High-resolution mapping of expression-QTLs yields insight into human gene regulation. PLoS Genet 4:e1000214. doi:10.1371/journal.pgen.1000214
Wang JH et al (2011) Modeling the cumulative genetic risk for multiple sclerosis from genome-wide association data. Genome Med 3:3. doi:10.1186/gm217
Weber F et al (2012) Single-nucleotide polymorphisms in HLA- and non-HLA genes associated with the development of antibodies to interferon-beta therapy in multiple sclerosis patients. Pharmacogenomics J 12:238–245. doi:10.1038/tpj.2011.14
Westra HJ, Franke L (2014) From genome to function by studying eQTLs Biochimica et Biophysica acta doi:10.1016/j.bbadis.2014.04.024
Yates CM, Sternberg MJ (2013) The effects of non-synonymous single nucleotide polymorphisms (nsSNPs) on protein-protein interactions. J Mol Biol 425:3949–3963. doi:10.1016/j.jmb.2013.07.012
Zeller T et al (2010) Genetics and beyond—the transcriptome of human monocytes and disease susceptibility. PLoS One 5:e10693
Acknowledgments
The study was supported by Natural Science Foundation of China (81473046, 81401343, 31401079, 31271336, 31071097, and 81373010), the Natural Science Foundation of Jiangsu Province (BK20130300), the Startup Fund from Soochow University (Q413900112, Q413900712), the Scientific Research Foundation for the Returned Overseas Chinese Scholars, State Education Ministry, and a Project of the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(XLSX 63 kb)
Rights and permissions
About this article
Cite this article
Lin, X., Deng, FY., Mo, XB. et al. Functional relevance for multiple sclerosis-associated genetic variants. Immunogenetics 67, 7–14 (2015). https://doi.org/10.1007/s00251-014-0803-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-014-0803-4