Abstract
Background
Over 200 genetic loci have been associated with multiple sclerosis (MS) explaining ~ 50% of its heritability, suggesting that additional mechanisms may account for the “missing heritability” phenomenon.
Objective
To analyze a large cohort of Italian individuals to identify markers associated with MS with potential functional impact in the disease.
Methods
We studied 2571 MS and 3234 healthy controls (HC) of continental Italian origin. Discovery phase included a genome wide association study (1727 MS, 2258 HC), with SNPs selected according to their association in the Italian cohort only or in a meta-analysis of signals with a cohort of European ancestry (4088 MS, 7144 HC). Top associated loci were then tested in two Italian cohorts through array-based genotyping (903 MS, 884 HC) and pool-based target sequencing (588 MS, 408 HC). Finally, functional prioritization through conditional eQTL and mQTL has been performed.
Results
Top associated signals overlap with already known MS loci on chromosomes 3 and 17. Three SNPs (rs4267364, rs8070463, rs67919208), all involved in the regulation of TBKBP1, were prioritized to be functionally relevant.
Conclusions
No evidence of novel signal of association with MS specific for the Italian continental population has been found; nevertheless, two MS loci seems to play a relevant role, raising the interest to further investigations for TBKBP1 gene.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Multiple Sclerosis (MS) is a complex autoimmune disease caused by the interplay of multiple genetic and environmental factors [1]. Large genome wide association studies (GWAS) provided important clues for the identification of genetic causes with more than 233 loci being discovered [2,3,4]. Although they are robustly associated, the identification of causative variants and the functional mechanism influencing MS pathogenesis has been identified only in few cases [5, 6]. Moreover, these studies rely on the inclusion of different populations to reach very large cohorts necessary to detect small effects like those observed for MS; however, different frequencies of disease variants across the different populations may mask the existence of additional associations. This was clearly evidenced in a GWAS in Sardinians, a genetically isolated population, which led to the identification of a novel MS variant in the TNFSF13B gene, whose allelic frequency allows its identification only in Sardinians and Southern European populations [6].
In the present project, we aim to analyse a large cohort of more than 5500 Italian individuals to identify markers associated with MS with potential functional impact in the disease. To boost the power of the analysis, we meta-analysed association results from different cohorts and leveraged quantitative molecular traits on adequately large resources for functional SNPs prioritization.
Materials and methods
Subjects
A total of 2571 Italian MS patients and 3234 unrelated HC have been recruited across Italian centres by three consortia (PROGRESSO, PROGEMUS and HYPERGENES). All participants were of Caucasian ancestry.
Based on the genotyping platform used for the genotyping, this entire set of samples was divided in four cohorts:
-
a)
ITAGWAS: it was composed of 776 MS and 1315 HC. MS samples were previously genotyped in the context of the International Multiple Sclerosis Consortium (IMSGC) at genome-wide level at the Wellcome Trust Sanger Institute using the Illumina® Human660-Quad chip [2]. Healthy individuals (mean (± SD) age of 53.4 ± 9.9 years and no abnormal findings on physical and neurological examination) have been collected within the HYPERGENES project (European Network for Genetic-Epidemiological Studies; www.hypergenes.eu) [7] in Italy and genotyped at the University of Milan, using the Illumina® Infinium 1 M-duo BeadChips.
-
b)
ITAiChip: it was composed of 964 MS patients and 1054 matched controls non-overlapping with the ITAGWAS cohort. Samples were previously genotyped in the context of the IMSGC at the Wellcome Trust Sanger Institute using the Illumina® Immunochip covering 196,524 SNPs and 184 non-HLA genomic regions with potential immunological function [3].
-
c)
ITAOA: it was composed of 917 MS patients and 924 controls. Samples were genotyping in the present study for selected SNPs using custom and pre-designed TaqMan assays (Thermo Fisher Scientific, Waltham, USA) assembled on two OpenArray chips run on the TaqMan Open Array Genotyping System (Thermo Fisher Scientific, Waltham, USA) or, in the context of the IMSGC, with the Illumina MS replication chip (Illumina, San Diego, USA), a custom array covering more than 300,000 SNPs localized in MS associated loci [4]
-
d)
ITANGS: subjects were chosen starting from the ITAGWAS and ITAiChip cohorts based on their genetic risk burden [8], estimated as a weighted sum of risk alleles in non-HLA known MS-associated loci [9,10,11]; 600 MS patients with a higher genetic risk (based on the distribution of the entire Italian cohort) were selected together with age- and sex-matched 408 HC, according to sex and Principal Components (PCs) to account for population stratification. ITANGS cohort underwent pooled-sequencing as previously described [12] using the Agilent SureSelect target enrichment method (Agilent Technologies, Santa Clara, USA) according to the manufacturer’s protocol.
The four cohorts are partially overlapping, as shown in Online Resource 1 fig. S1.
Demographic information for the mentioned cohorts is listed in Table 1, while details on genetic data generation are described in the Online Resource 1.
Additionally to the Italian cohorts, a total of 4088 MS patients and 7,144 HC of European descent (“NonIT-EUR” cohort) have been included in the analyses for the “meta-analysis approach” (see below). This represents a combination of 6 individual sample collections previously described [13].
All patients were diagnosed according to Poser and McDonald criteria and further revisions [14].
Design of the study
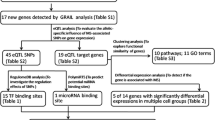
Figure 1 shows the workflow of the study. Briefly, we followed a multi-step approach including a discovery step at genome-wide level followed by replication and fine-mapping through next-generation sequencing of the top-associated loci. Finally, the effect of the identified variants on gene expression and DNA methylation was evaluated to prioritize markers with potential functional role.
Workflow of the study. The overall workflow of the study is shown. The discovery phase includes two Italian cohorts (ITAGWAS and ITAiChip) from which a set of top MS associated SNPs were identified through a genome-wide approach (in the ITAGWAS cohort only) or a meta-analytic approach including also the NonIT-EUR dataset. The identified top SNPs (P < 5 × 10– 7) were further tested for replication in a third independent Italian cohort (ITAOA) leading to the confirmation of two associated loci. Those loci were sequenced (ITANGS cohort) and the associated SNPs underwent prioritization through conditional cis eQTL and mQTL analyses
Discovery
We initially investigated the presence of association signals specific for the Italian population by performing a GWAS in the ITAGWAS cohort alone (here called the “genome-wide Italian approach”), this led to the selection of SNPs (P < 5 × 10– 7) to further replicate in the replication step.
An alternative approach (here called the “meta-analysis approach”) was designed to increase the sample size of the discovery phase and detect association signals with a lower impact on MS susceptibility not detectable when studying only the ITAGWAS cohort. Specifically, two additional cohorts were included in the study design: the ITAiChip, composed by Italian individuals genotyped at relevant immunological genomic regions, and the NonIT-EUR dataset, which represents a large cohort of non-Italian individuals for which genome-wide genotyping data are available. According to this approach, we selected SNPs with suggestive evidence of association (P < 0.005) in the ITAGWAS cohort alone (to detect signals at genome-wide level), as well as performing a meta-analysis between ITAGWAS and ITAiChip cohorts. A meta-analysis with the NonIT-EUR dataset was then performed for the selected signals and significant SNPs (P < 5 × 10– 7) were considered for replication.
Replication
We defined loci for replication as described in the Online Resource 1. Due to the known strong associations within the MHC region with MS [4], SNPs located in the HLA locus were excluded.
Genotyping was performed using TaqMan assays and the TaqMan Open Array Genotyping System (Thermo Fisher Scientific, Waltham, USA) or the MS Chip [4] (Illumina, San Diego, USA). Single-SNP logistic regression using sex as covariate was conducted under additive model of inheritance. Meta-analyses between the discovery and replication cohorts were performed assuming a fixed-effect model.
Fine mapping with next generation sequencing (NGS)
We sequenced the Linkage Disequilibrium (LD) blocks containing the replicated associated signals in the ITANGS cohort using the Agilent SureSelect kit (Agilent Technologies, Santa Clara, USA) and a pool-based approach on the GaIIx platform (Illumina, San Diego, USA). Bioinformatics pipeline and variant calling were performed as described [12]. Fisher exact test comparing MS and HC was performed on allelic counts, no correction based on sex or ancestry was included since HC were individually matched with MS patients, as described in the “Subjects” paragraph. For methodological details see Online Resource 1.
Prioritization
Identified associated variants were prioritized according to their cis effect on expression or CpG methylation by performing conditional cis-eQTL and conditional cis-mQTL analyses. Cis-eQTL analysis was performed taking advantage of the GTEx Portal v6p [15] dataset (release phs000424.v6.p1) for 7 MS-related tissues (whole blood, Brain Frontal_Cortex (BA9), Brain Cortex, Cerebellar Hemisphere, Cerebellum, Anterior cingulate cortex (BA24), Hypothalamus). Cis-mQTL analysis was performed on CD4+ T-cells collected from 156 MS patients within the Comprehensive Longitudinal Investigation of Multiple Sclerosis at the Brigham and Women's Hospital (CLIMB study [16, 17]). Both analyses were performed using QTLtools [18]. For details see Online Resource 1.
Results
Discovery
As described in the method section, the “genome-wide Italian approach” includes the ITAGWAS cohort as discovery dataset. After quality control (QC), the association analysis was performed in a final dataset of 737 MS and 1291 HC individuals and ~ 6.6 million SNPs. Seventy-one variants reached a p value of association < 5 × 10–7, within 26 distinct loci (Online Resource 2, Online Resource 1 Fig. S2).
The “meta-analysis approach” includes both the ITAGWAS and ITAiChip cohorts as discovery datasets, with signals with suggestive evidence of association (P < 0.005) selected to be meta-analysed with the NonIT-EUR. Regarding the ITAGWAS cohort alone, 32,309 SNPs (P < 0.005) were meta-analysed with the NonIT-EUR leading to a list of 49 SNPs with a p-value of association < 5 × 10– 7 (Online Resource 3A). The meta-analysis of the two Italian discovery datasets led to the identification of 1373 SNPs suggestive of association (P < 0.005). Meta-analysis of these SNPs with the NonIT-EUR dataset resulted in 21 SNPs with a p value < 5 × 10– 7 (Online Resource 3B) within 5 loci. Overall, and given the fact that some signals overlapped between the different analyses, 127 signals across 35 loci were selected (Online Resource 2 and Online Resource 3).
Replication
Starting from the 127 signals identified through the discovery analyses (Online Resource 2 and Online Resource 3), we selected 60 SNPs for replication as representative of the identified genomic loci (see Online Resource 1 for details and Online Resource 2 and Online Resource 3 for the final list). After genotyping and QC (see Online Resource 1 for details), 48 SNPs within 30 loci were tested for association in the ITAOA dataset (Table 2).
No SNPs identified through the “genome-wide Italian approach” showed evidence of association, while according to the “meta-analysis approach” (thus starting from the two ITAGWAS and ITAiChip cohorts), the locus 1 on chromosome 3 and locus 8 on chromosome 17 confirmed their association, with rs338603 (OR = 1.27, P = 8.01 × 10− 4) and rs669607 (OR = 1.23, P = 4.47 × 10− 3) and rs8070463 (OR = 1.24, P = 3.43 × 10− 3) as the top associated SNPs. These associations were confirmed also performing a meta-analysis excluding the NonIT-EUR cohort and including only dataset of Italian origin (P = 3.73 × 10– 7, 6.01 × 10– 7 and 1.72 × 10− 5 respectively for rs338603, rs669607 and rs8070463) (Table 2, Online Resource 1 Fig. S3).
Fine mapping
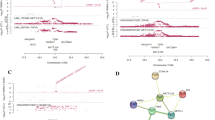
The replicated locus on chromosome 3 spans from position 28024996 to 28124996 (hg19 assembly). This is an intergenic region including the non-coding genes LINC01980 (~ 123 kb) and LINC01981 (~ 150 kb) and the coding genes EOMES (~ 160 Kb) and CMC1 (~ 150 kb) as the closest downstream genes. A total of 608 variants were identified through the sequencing approach. Comparing to the post-QC imputed ITAGWAS dataset, the inclusion of a sequencing step allowed to investigate additional 74 common variants (MAF ≥ 0.01), as well as 636 rare variants (Minor allele Frequency (MAF) < 0.01), while 109 were excluded and not tested in the ITANGS cohort due to the stringent QC applied. Fisher’s exact test on sequenced data revealed the association of 12 variants (Bonferroni-adjustment alpha = 5.87 × 10– 5). The most associated SNPs were rs669607 and rs427221 (P = 5.02 × 10− 13) (Fig. 2a, Online Resource 4).
Regional association plots of replicated loci in ITANGS cohort. Regional association plots for locus 1 on chromosome 3 (a) and locus 8 on chromosome 17 (b) designed with LocusZoom (http://locuszoom.sph.umich.edu) are shown. For each locus, the negative log-transformed P values (left y-axis) and the recombination rate (right y-axis) according to the 1000 Genomes Project (Nov 2014, Europeans only) are plotted over the genomic position (hg19). Each symbol represents one SNP, with the most associated SNP marked in purple and shading of the other points based on the linkage disequilibrium with the top SNP. Positions of genes, if present, are shown below the plot
The replicated locus on chromosome 17 spans a large LD block of ~ 500 kb (chr17:45309693-45824486, hg19 assembly) which includes 8 genes: ITGB3, EFCAB13 (C17orf57), NPEPPS, KPNB1, TBKBP1, TBX21, MRPL45P2 (pseudogene) and LOC102724508 (a non-coding RNA, with alias THCAT158). In this region, a total of 2,428 variants were identified through the sequencing approach. Comparing to the post-QC imputed ITAGWAS dataset, the inclusion of the fine mapping step allowed to investigate additional 446 common variants and 2,395 rare variants, while 341 SNPs were not tested in the ITANGS cohort for the sequencing QC. Applying a pipeline which allows an accurate allele frequency estimation from NGS pooled data [12] to perform the association analysis, we found that 202 SNPs were significantly associated after Bonferroni-adjustment (alpha = 1.57 × 10− 5) (Fig. 2b, Online Resource 4). The strongest association signal was rs4239162 (P = 1.04 × 10− 20), intronic to KPNB1.
Locus on chromosome 3
The locus on chromosome 3 is a small intergenic genomic region, with apparently no evidence of chromatin annotation (Online Resource 1 Fig. S4), except for the presence of sporadic weak enhancers. Cis eQTL analysis did not reveal any association between the 12 SNPs identified in the genetic analyses and the expression of nearby genes.
Locus on chromosome 17: functional SNPs prioritization
We sought to characterize the identified MS risk variants prioritizing those with a putative functional impact. We then performed a conditional cis-eQTL analysis in 7 tissues relevant for MS according to data generated within the GTEx project [15]. As shown in Online Resource 5A, we found 7 SNP-gene pairs associated in the tested tissues. Among them, rs4267364 showed a consistent association with MS in all the steps of the study (Table 3). This SNP is intronic to TBKBP1 and the MS associated allele (rs4267364A) is associated to a higher expression of this gene in blood (Fig. 3a). According to the chromatin state prediction in 14 epigenomes of immune-related cells profiled by the Roadmap Epigenomics [19] and ENCODE [20] projects, this SNP is localized in a putative enhancer, supporting its regulatory function (Online Resource 1 Fig. S5).
Prioritized eQTL and mQTL effects. eQTL and mQTL effects are depicted in the figures. a TBKBP1 normalized expression values in whole blood are plotted against the genotype of rs4267364; b cg08452456 M values are plotted against the genotype of rs8070463; c cg12183861 M values are plotted against the genotype of rs67919208. Representation is in box-plot format (Tukey method) with each dot representing one individual
To explore additional mechanisms of regulation, we performed a conditional cis-mQTL in CD4+ cells collected from MS patients. The Online Resource 5B lists the significant SNP-CpG pairs resulted from this analysis. Among the identified mQTL associations, one SNP associated to MS according to the discovery strategy and at least one of the replication steps, rs8070463, was found to be associated also with the methylation status of cg08452456 (Table 3, Fig. 3b), exonic to TBKBP1. An additional SNP, rs67919208, associated with the methylation status of cg12183861 (Table 3, Fig. 3c), localized in the 5’UTR region of TBKBP1, was also found to be associated with MS (discovery step). Both rs8070463 and rs67919208 had the MS risk variant associated with a higher CpG methylation (Table 3).
All the three prioritized variants suggest an involvement of TBKBP1 gene, thus we then evaluated the expression of TBKBP1 across immune cells according to the DICE dataset [21]. As shown in Online Resource 1 fig. S6, TBKBP1 is expressed across several immune cell types, with the highest levels in activated T cells (CD4+ and CD8+) and Natural Killer cells.
Discussion
Large scale international studies showed a highly polygenic architecture for MS genetic susceptibility [4]; nonetheless, the identified variants explain ~ 50% of the genetic predisposition to MS, suggesting that additional signals and disease mechanisms exist. This “missing heritability” could be explained, at least partially, by the presence of population-specific associations that cannot be fully captured by array-based international studies, as already shown studying the Sardinian [6] and German populations [22].
To investigate the presence of strong association signals enriched in the Italian population, we initially performed a genome-wide association study (referred along the paper as the “genome-wide Italian approach”) in a cohort of Italian individuals that led us to the identification of several loci potentially associated with MS, most of them never reported in previous studies (Online Resource 2). However, replication did not confirm any of these associations, supporting the idea that no strong signals exist in the continental Italian population. Nevertheless, we can’t exclude that other events occurred, including population stratification, genetic heterogeneity, or an insufficient statistical power [23].
We then followed an alternative approach, exploring also signals with suggestive evidence of association and meta-analysing them with a cohort of European descent with larger adequate sample size and statistical power. The genome-wide Italian approach would have the potential to identify signals with a strong association with MS specific for the Italian population, while the second approach combined the advantage of using an ancestrally homogeneous cohort as the driving cohort of loci selection with the statistical power gained using a larger cohort. Specifically, this second approach, referred as “meta-analysis approach”, highlighted the presence of two loci representing the main results of this effort. Both lie in loci already known to be associated with MS [2,3,4]: an intergenic genomic region on chromosome 3 (~ 100 Kb) and a large locus of ~ 500 kb on chromosome 17.
To better characterize the two selected loci and to prioritize putative causal variants, we then performed a deep target sequencing followed by functional prioritization, allowing to investigate additional associated SNPs, not covered by arrays or imputation. In the case of the locus on chromosome 3, this analysis led to the identification of 12 associated SNPs, with rs669607 and rs427221 as the top associated ones [2]. No clear function is suggested, since this is an intergenic region with no annotation suggestive of specific regulatory function [4]. Nevertheless, very recently, a regulatory effect on the upstream gene EOMES in T cells has been highlighted [24], in line with previous studies supporting a role of this gene in MS [25].
On the contrary, the locus on chromosome 17 is more complex, with several genes and potentially regulatory regions. Sequencing of this region led to the identification of 202 signals of associations with the top associated signals localized in the downstream part of the region, as suggested also by the discovery analysis. This high number of associated signals could be explained by the strong linkage disequilibrium characterizing this locus (Fig. 2b), which further complicates the identification of the causative signal of association. We therefore resorted to exploit regulatory data for prioritization of the identified SNPs. Several studies reported an enrichment of regulatory regions within MS and other complex diseases GWAS loci [24, 26,27,28,29,30], thus we performed a conditional cis eQTL analysis to either overcome the presence of the large LD block and to combine the results with the obtained genetic association. This analysis highlighted rs4267364, which is an intronic polymorphism to TBKBP1, localized in a putative enhancer. This SNP is associated with the expression of TBKBP1 in blood, but not in brain-related tissues, with the MS risk allele rs4267364A associated with a higher expression of TBKBP1 (P = 1.71 × 10– 23). We also investigated additional mechanisms of regulation performing a conditional cis mQTL analysis to identify SNPs associated with the methylation state of CpG in CD4+ lymphocytes of MS patients. Several associations were found (Online Resource 5B), with two SNPs, rs8070463 and rs67919208, which were also selected by the genetic approach. These two SNPs were found to be associated with the methylation status of respectively cg08452456, exonic to TBKBP1, and cg12183861, localized in the 5’UTR of TBKBP1. Interestingly enough, rs8070463 SNP has been previously associated with ankylosing spondylitis, a rare autoimmune condition [31]. It is well established that methylation of regulatory regions is an important mechanism [32]: DNA methylation in promoter regions is known to induce gene silencing [33], while it has been reported that DNA methylation in transcribed regions seems to correlate with an induced gene expression [34]. Given their genomic position, we could hypothesize that the influence on the methylation status of the mentioned CpGs could in turn lead to an upregulation of TBKBP1, as suggested by eQTL analysis on GTEx dataset, with the MS risk allele being associated with an upregulation of TBKBP1 in whole blood (P = 1.03 × 10– 20 and 2.17 × 10– 20 respectively for rs8070463 and rs67919208), although we could not exclude that this reflects a LD with the eQTL leading SNP rs4267364 (r2 = 0.66 for both the SNPs). Functional experiments testing methylation and expression profiles in same samples are necessary to clarify the direction of the signal.
All of the three prioritized SNPs seems to be involved in the regulation of the expression of TBKBP1, suggesting that an upregulation of this gene could be relevant in MS pathogenesis. TBKBP1 codes for the TBK1 binding protein 1, an adaptor protein able to bind TBK1 and involved in the TNFα/NFkB pathway [35]. Contrary to TBKBP1, for which few reports are available regarding its function, TBK1 is involved in several immune processes, including T-cell homeostasis and migration cells from lymph nodes to the CNS in the MS murine model [36]. Although the direct involvement of TBKBP1 in TBK1-mediated signaling in migration of T cells has never been investigated, we could not exclude that an alteration of the expression of this gene could also influence this mechanism. Moreover, TBKBP1 was recently found to be an important regulator of development and survival of natural killer (NK) cells [37]. The role of NK cells in MS is still debated, however there is increasing evidence that certain subtypes of NK cells are able to modulate other immune cells and could be involved in autoimmune processes [38, 39]. Supporting these hypotheses, TBKBP1 is highly expressed in activated CD4+ and CD8+ T cells, as well as NK cells (Online Resource 1 Fig. S6). Further tailored experiments are needed to elucidate the involvement of TBKBP1 in MS.
Concluding, our results suggest that no novel signal associated with MS with a strong effect and specific for the continental Italian population exists. Nevertheless, we prioritized two loci, already identified to be associated with MS, that may have ethnic specificity in the Italian population, one of them pointing to the TBKBP1 gene as a candidate to be investigated further.
Change history
16 September 2022
Missing Open Access funding information has been added in the Funding Note.
25 June 2022
A Correction to this paper has been published: https://doi.org/10.1007/s00415-022-11216-6
References
Compston A, Coles A (2008) Multiple sclerosis. Lancet 372:1502–1517
Sawcer S, International Multiple Sclerosis Genetics Consortium, Wellcome Trust Case Control Consortium 2 et al (2011) Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 476:214–219
Beecham AH, Patsopoulos NA, International Multiple Sclerosis Genetics Consortium (IMSGC) et al (2013) Analysis of immune-related loci identifies 48 new susceptibility variants for multiple sclerosis. Nat Genet 45:1353–1360. https://doi.org/10.1038/ng.2770
International Multiple Sclerosis Genetics Consortium (IMSGC) (2019) Multiple sclerosis genomic map implicates peripheral immune cells and microglia in susceptibility. Science 365:eaav7188. https://doi.org/10.1126/science.aav7188
Gregory AP, Dendrou CA, Attfield KE et al (2012) TNF receptor 1 genetic risk mirrors outcome of anti-TNF therapy in multiple sclerosis. Nature 488:508–511. https://doi.org/10.1038/nature11307
Steri M, Orrù V, Idda ML et al (2017) Overexpression of the cytokine BAFF and autoimmunity risk. N Engl J Med 376:1615–1626. https://doi.org/10.1056/NEJMoa1610528
Salvi E, Kutalik Z, Glorioso N et al (2012) Genomewide association study using a high-density single nucleotide polymorphism array and case-control design identifies a novel essential hypertension susceptibility locus in the promoter region of endothelial NO synthase. Hypertension 59:248–255. https://doi.org/10.1161/HYPERTENSIONAHA.111.181990
De Jager PL, Chibnik LB, Cui J et al (2009) Integration of genetic risk factors into a clinical algorithm for multiple sclerosis susceptibility: a weighted genetic risk score. Lancet Neurol 8:1111–1119. https://doi.org/10.1016/S1474-4422(09)70275-3
Sorosina M, Brambilla P, Clarelli F et al (2014) Genetic burden of common variants in progressive and bout-onset multiple sclerosis. Mult Scler J 20:802–811. https://doi.org/10.1177/1352458513512707
Sorosina M, Esposito F, Guaschino C et al (2015) Inverse correlation of genetic risk score with age at onset in bout-onset and progressiveonset multiple sclerosis. Mult Scler J 21:1463–1467. https://doi.org/10.1177/1352458514561910
Barizzone N, Zara I, Sorosina M et al (2015) The burden of multiple sclerosis variants in continental Italians and Sardinians. Mult Scler 21:1385–1395. https://doi.org/10.1177/1352458515596599
Anand S, Mangano E, Barizzone N et al (2016) Next generation sequencing of pooled samples: guideline for variants’ filtering. Sci Rep 6:33735. https://doi.org/10.1038/srep33735
Patsopoulos NA, Genetics Working Group, Steering Committees of Studies Evaluating IFNβ-1b and a CCR1-Antagonist et al (2011) Genome-wide meta-analysis identifies novel multiple sclerosis susceptibility loci. Ann Neurol 70:897–912. https://doi.org/10.1002/ana.22609
Polman CH, Reingold SC, Banwell B et al (2011) Diagnostic criteria for multiple sclerosis: 2010 Revisions to the McDonald criteria. Ann Neurol 69:292–302. https://doi.org/10.1002/ana.22366
GTEx Consortium (2013) The genotype-tissue expression (GTEx) project. Nat Genet 45:580–585. https://doi.org/10.1038/ng.2653
Gauthier SA, Glanz BI, Mandel M, Weiner HL (2006) A model for the comprehensive investigation of a chronic autoimmune disease: the multiple sclerosis CLIMB study. Autoimmun Rev 5:532–536
Roostaei T, Klein H-U, Felsky D, et al (2020) Impact of genetic susceptibility to multiple sclerosis on the T cell epigenome: proximal and distal effects. bioRxiv 2020.07.11.198721. https://doi.org/10.1101/2020.07.11.198721
Delaneau O, Ongen H, Brown AA et al (2017) A complete tool set for molecular QTL discovery and analysis. Nat Commun 8:15452. https://doi.org/10.1038/ncomms15452
Roadmap Epigenomics Consortium, Kundaje A, Meuleman W et al (2015) Integrative analysis of 111 reference human epigenomes. Nature 518:317–329. https://doi.org/10.1038/nature14248
ENCODE Project Consortium (2012) An integrated encyclopedia of DNA elements in the human genome. Nature 489:57–74. https://doi.org/10.1038/nature11247
Schmiedel BJ, Singh D, Madrigal A et al (2018) Impact of genetic polymorphisms on human immune cell gene expression. Cell 175:1701-1715.e16. https://doi.org/10.1016/j.cell.2018.10.022
Andlauer TFM, Buck D, Antony G et al (2016) Novel multiple sclerosis susceptibility loci implicated in epigenetic regulation. Sci Adv 2:e1501678. https://doi.org/10.1126/sciadv.1501678
Greene CS, Penrod NM, Williams SM, Moore JH (2009) Failure to replicate a genetic association may provide important clues about genetic architecture. PLoS ONE 4:e5639. https://doi.org/10.1371/journal.pone.0005639
International Multiple Sclerosis Genetics Consortium (2019) A systems biology approach uncovers cell-specific gene regulatory effects of genetic associations in multiple sclerosis. Nat Commun 10:2236. https://doi.org/10.1038/s41467-019-09773-y
Parnell GP, Gatt PN, Krupa M et al (2014) The autoimmune disease-associated transcription factors EOMES and TBX21 are dysregulated in multiple sclerosis and define a molecular subtype of disease. Clin Immunol 151:16–24. https://doi.org/10.1016/j.clim.2014.01.003
Disanto G, Sandve GK, Berlanga-Taylor AJ et al (2012) Genomic regions associated with multiple sclerosis are active in B cells. PLoS ONE 7:e32281. https://doi.org/10.1371/journal.pone.0032281
Maurano MT, Humbert R, Rynes E et al (2012) Systematic localization of common disease-associated variation in regulatory DNA. Science 337:1190–1195. https://doi.org/10.1126/science.1222794
Corradin O, Saiakhova A, Akhtar-Zaidi B et al (2014) Combinatorial effects of multiple enhancer variants in linkage disequilibrium dictate levels of gene expression to confer susceptibility to common traits. Genome Res 24:1–13. https://doi.org/10.1101/gr.164079.113
Farh KKH, Marson A, Zhu J et al (2015) Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature 518:337–343. https://doi.org/10.1038/nature13835
Cotsapas C, Mitrovic M (2018) Genome-wide association studies of multiple sclerosis. Clin Trans Immunol 7:e1018. https://doi.org/10.1002/cti2.1018
Evans DM, Spencer CCA, Pointon JJ et al (2011) Interaction between ERAP1 and HLA-B27 in ankylosing spondylitis implicates peptide handling in the mechanism for HLA-B27 in disease susceptibility. Nat Genet 43:761–767. https://doi.org/10.1038/ng.873
Smith ZD, Meissner A (2013) DNA methylation: roles in mammalian development. Nat Rev Genet 14:204–220
Schultz MD, He Y, Whitaker JW et al (2015) Human body epigenome maps reveal noncanonical DNA methylation variation. Nature 523:212–216. https://doi.org/10.1038/nature14465
Yang X, Han H, DeCarvalho DD et al (2014) Gene body methylation can alter gene expression and is a therapeutic target in cancer. Cancer Cell 26:577–590. https://doi.org/10.1016/j.ccr.2014.07.028
Ryzhakov G, Randow F (2007) SINTBAD, a novel component of innate antiviral immunity, shares a TBK1-binding domain with NAP1 and TANK. EMBO J 26:3180–3190. https://doi.org/10.1038/sj.emboj.7601743
Yu J, Zhou X, Chang M et al (2015) Regulation of T-cell activation and migration by the kinase TBK1 during neuroinflammation. Nat Commun 6:6074. https://doi.org/10.1038/ncomms7074
Zhu L, Xie X, Zhang L et al (2018) TBK-binding protein 1 regulates IL-15-induced autophagy and NKT cell survival. Nat Commun 9:2812. https://doi.org/10.1038/s41467-018-05097-5
Gross CC, Schulte-Mecklenbeck A, Wiendl H et al (2016) Regulatory functions of natural killer cells in multiple sclerosis. Front Immunol 7:606. https://doi.org/10.3389/fimmu.2016.00606
Nagy N (2017) Establishment of EBV-infected lymphoblastoid cell lines. Methods in Molecular Biology. Frontiers Media S.A,Humana Press, New York, pp 57–64
Acknowledgements
The authors thank the International MS Genetics Consortium for providing part of the genotype data. They also thank individuals with MS and their families for making this study possible and nurses at the MS center at San Raffaele Hospital for help in collection of samples. Members of the PROGEMUS consortium: Lucia Florioa, Paolo Ragoneseb, Alberto Gajofattoc, Elio Scarpinid, Domenico Caputoe, Claudio Gasperinif, Franco Granellag, Paola Cavallah, Roberto Bergamaschii, Giovanni Ristorij, Claudio Solarok, Filippo Martinelli Boneschil, Francesco Passantinom, Maura Pugliattin, Antonio Galloo, Laura Brambillap, Marinella Clericoq, Fioravante Caponer, Maria Trojanos, aIRCCS Casa Sollievo della Sofferenza, San Giovanni Rotondo, Italy, bPoliclinico Paolo Giaccone, Palermo, Italy, cPoliclinico Borgo Roma, Verona, Italy, dFoundation IRCCS Cà Granda Ospedale Maggiore Policlinico, Milan, Italy, eFoundation Don Carlo Gnocchi, Milan, Italy, fAzienda Ospedaliera S. Camillo Forlanini, Rome, Italy, gAzienda Ospedaliera-Universitaria di Parma, Parma, Italy, hAzienda Ospedaliera-Universitaria S. Giovanni Battista, Molinette (TO), Italy, iFondazione Istituto Neurologico Nazionale Casimiro Mondino, Pavia, Italy, jSant’Andrea Hospital, Rome, Italy, kC.R.R.F. “Mons. Luigi Novarese”, Moncrivello (VC), Italy, lIRCCS Policlinico San Donato Milanese (MI), Italy, mOspedale SS Antonio e Biagio, Alessandria, Italy, nUO Clinica Neurologica, University of Ferrara, Ferrara, Italy, oI Division of Neurology, II university of Naples, Naples, Italy, pFondazione IRCCS Istituto Neurologico “Carlo Besta”, Milan, Italy, qAzienda Ospedaliero-Universitaria San Luigi Gonzaga, Orbassano (TO), Italy, rPoliclinico Universitario Campus Biomedico, Rome, Italy, sUniversity of Bari”Aldo Moro”, Bari, Italy.
Funding
Open access funding provided by Università degli Studi di Milano within the CRUI-CARE Agreement. This work was supported by the Italian Foundation of Multiple Sclerosis [grant numbers: 2011/R/14, 2013/R/13, 2015/R/10, 2019/R-Multi/033, 2019/S/4]; the Italian Ministry of Health [grant number RF-2016-02361294]; the AGING Project for Department of Excellence at the Department of Translational Medicine (DIMET) at Università del Piemonte Orientale (Novara, Italy); and HYPERGENES [grant number HEALTH-F4-2007-201550]; Miriam Zuccalà is supported by Consorzio Interuniversitario di Biotecnologie (CIB) and AGING Project; Melissa Sorosina and Nadia Barizzone are partially supported by the Horizon 2020 European Grant [MultipleMS project]; Nikolaos A. Patsopoulos is supported by the National Multiple Sclerosis Society [grant numbers RG-1707-28657, JF-1808-32223 Harry Weaver Award].
Author information
Authors and Affiliations
Consortia
Contributions
Study was concepted and designed by FMB, SDA, FE, MS, NB and FC. Material preparation, data collection and analysis were performed by MS, NB, FC, SA, SL, ES, EM, RB, TR, EM, MZ, DV, PC, SS, LF, AZ, CB, DC, VM, GC, ML, MF, NAP, PLDeJ, GDeB. The first draft of the manuscript was written by MS and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
M. Sorosina, N. Barizzone, F. Clarelli, S. Anand, S. Lupoli, E. Salvi, E. Mangano, R. Bordoni, T. Roostaei, E. Mascia, M. Zuccalà, D. Vecchio, S. Santoro, L. Ferrè, M. Pozzato, C. Barlassina, D. Cusi, A. Patsopoulous, P.L. De Jager, G. De Bellis, S. D’Alfonso, F. Martinelli Boneschi have no competing interests to declare that are relevant to the content of this article. P. Cavalla received honoraria for speaking and/or for consultancy and support for participation to scientific congresses from Almirall, Biogen, Merck-Serono, Novartis, Sanofi-Genzyme, Roche and Teva. V. Martinelli received honoraria for speaking and/or for consultancy and support for travel expenses and participation in Congresses from Almirall, Biogen, Merck-Serono, Novartis, Genzyme and Teva. G. Comi has received consulting and speaking fees from Novartis, Sanofi Genzyme, Genzyme Corporation, Merck KGgA, Merck Serono SpA, Celgene Group, F. Hoffman-La Roche, Almirall SpA, Janssen-Cilag GmbH, Janssen Global Services LLC, Magenta Therapeutics Inc. Almirall and Merck Serono SpA outside the submitted work. Dr. Leone reports grants from Italian Ministry of Health (RF-2016-02361294 grant), grants from Italian Ministry of Health- Ricerca corrente 2017—RC1701NE08, during the conduct of the study. M. Filippi received compensation for consulting services and/or speaking activities from Almiral, Alexion, Bayer, Biogen, Celgene, Eli Lilly, Genzyme, Merck-Serono, Novartis, Roche, Sanofi, Takeda, and Teva Pharmaceutical Industries and receives research support from Biogen Idec, Merck-Serono, Novartis, Roche, Teva Pharmaceutical Industries. F. Esposito received consulting and speaking fees from Novartis, Sanofi Genzyme.
Ethics approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by each study site from the local ethics committee or institutional research board. Written informed consent was obtained for all individuals at every site by the principal investigator.
Additional information
The original online version of this article was revised: The table 2 caption has been updated.
Membership of the consortia are listed in the acknowledgement.
The original online version of this article was revised: The table 2 caption has been updated.
Supplementary Information
Below is the link to the electronic supplementary material.
415_2022_11109_MOESM1_ESM.docx
Supplementary file1 Online Resource 1: supplementary methods and figures. Online Resource 2: association results according to the “genome-wide Italian approach.” Online Resource 3: association results according to the “meta-analysis approach.” Online Resource 4: association results of target sequencing.Online Resource 5: independent eQTL and mQTL associations in MS-related tissues (DOCX 11035 kb)
415_2022_11109_MOESM2_ESM.xlsx
Supplementary file1 Online Resource 1: supplementary methods and figures. Online Resource 2: association results according to the “genome-wide Italian approach.” Online Resource 3: association results according to the “meta-analysis approach.” Online Resource 4: association results of target sequencing.Online Resource 5: independent eQTL and mQTL associations in MS-related tissues (DOCX 27 kb)
415_2022_11109_MOESM3_ESM.xlsx
Supplementary file1 Online Resource 1: supplementary methods and figures. Online Resource 2: association results according to the “genome-wide Italian approach.” Online Resource 3: association results according to the “meta-analysis approach.” Online Resource 4: association results of target sequencing.Online Resource 5: independent eQTL and mQTL associations in MS-related tissues (DOCX 34 kb)
415_2022_11109_MOESM4_ESM.xlsx
Supplementary file1 Online Resource 1: supplementary methods and figures. Online Resource 2: association results according to the “genome-wide Italian approach.” Online Resource 3: association results according to the “meta-analysis approach.” Online Resource 4: association results of target sequencing.Online Resource 5: independent eQTL and mQTL associations in MS-related tissues (DOCX 44 kb)
415_2022_11109_MOESM5_ESM.xlsx
Supplementary file1 Online Resource 1: supplementary methods and figures. Online Resource 2: association results according to the “genome-wide Italian approach.” Online Resource 3: association results according to the “meta-analysis approach.” Online Resource 4: association results of target sequencing.Online Resource 5: independent eQTL and mQTL associations in MS-related tissues (DOCX 29 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sorosina, M., Barizzone, N., Clarelli, F. et al. A multi-step genomic approach prioritized TBKBP1 gene as relevant for multiple sclerosis susceptibility. J Neurol 269, 4510–4522 (2022). https://doi.org/10.1007/s00415-022-11109-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00415-022-11109-8