Abstract
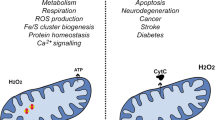
Cytochrome c delicately tilts the balance between cell life (respiration) and cell death (apoptosis). Whereas cell life is governed by transient electron transfer interactions of cytochrome c inside the mitochondria, the cytoplasmic adducts of cytochrome c that lead to cell death are amazingly stable. Interestingly, the contacts of cytochrome c with its counterparts shift from the area surrounding the heme crevice for the redox complexes to the opposite molecule side when the electron flow is not necessary. The cytochrome c signalosome shows a higher level of regulation by post-translational modifications—nitration and phosphorylation—of the hemeprotein. Understanding protein interfaces, as well as protein modifications, would puzzle the mitochondrial cytochrome c-controlled pathways out and enable the design of novel drugs to silence the action of pro-survival and pro-apoptotic partners of cytochrome c.



Similar content being viewed by others
Abbreviations
- Adx:
-
Adrenodoxin
- AdxR:
-
NADPH-dependent adrenodoxin reductase
- bc 1 :
-
Cytochrome bc 1 complex
- CB:
-
Cytochrome binding
- Cb 5 :
-
Cytochrome b 5
- Cb 5R:
-
NADH-dependent cytochrome b 5 reductase
- Cc :
-
Cytochrome c
- Cc 552 :
-
Cytochrome c 552
- CcO:
-
Cytochrome c oxidase
- CcP:
-
Cytochrome c peroxidase
- CH1:
-
Collagen homologous 1
- CH2:
-
Collagen homologous 2
- CL:
-
CardioLipin
- ET:
-
Electron transfer
- GALDH:
-
l-GAlactono-1,4-Lactone DeHydrogenase
- IMM:
-
Inner mitochondrial membrane
- IMS:
-
Intermembrane mitochondrial space
- n-Cc :
-
Nitrated cytochrome c
- NMR:
-
Nuclear magnetic resonance
- OMM:
-
Outer mitochondrial membrane
- p-Cc :
-
Phosphorylated cytochrome c
- PCD:
-
Programmed cell death
- PKCβ:
-
Protein kinase C β
- PKCδ:
-
Protein kinase C δ
- PRE:
-
Paramagnetic relaxation enhancement
- PTB:
-
PhosphoTyrosine binding
- R(N)OS:
-
Reactive (nitrogen)oxygen species
- Sco:
-
Synthesis of cytochrome c oxidase
- SH2:
-
Src homology 2
- WT:
-
Wild-type
References
Acehan D, Jiang X, Morgan DG, Heuser JE, Wang X, Akey CW (2002) Three-dimensional structure of the apoptosome: implications for assembly, procaspase-9 binding, and activation. Mol Cell 9:423–432
Acin-Perez R, Hoyos B, Gong J, Vinogradov V, Fischman DA, Leitges M, Borhan B, Starkov A, Manfredi G, Hammerling U (2010) Regulation of intermediary metabolism by the PKCδ signalosome in mitochondria. FASEB J 24:5033–5042
Banci I, Bertini I, Felli IC, Krippahl L, Kubicek K, Moura JJG, Rosato A (2003) A further investigation of the cytochrome b 5–cytochrome c complex. J Biol Inorg Chem 8:777–786
Banci L, Bertini I, Cavallaro G, Ciofi-Baffoni S (2011a) Seeking the determinants of the elusive functions of Sco proteins. FEBS J 278:2244–2262
Banci L, Bertini I, Ciofi-Baffoni S, Kozyreva T, Mori M, Wang S (2011b) Sco proteins are involved in electron transfer processes. J Biol Inorg Chem 16:391–403
Bashir Q, Volkov AN, Ullmann GM, Ubbink M (2010) Visualization of the encounter ensemble of the transient electron transfer complex of cytochrome c and cytochrome c peroxidase. J Am Chem Soc 132:241–247
Bashir Q, Scanu S, Ubbink M (2011) Dynamics in electron transfer protein complexes. FEBS J 278:1391–1400
Batthyány C, Souza JM, Durán R, Cassina A, Cerveñansky C, Radi R (2005) Time course and site(s) of cytochrome c tyrosine nitration by peroxynitrite. Biochemistry 44:8038–8046
Bayir H, Fadeel B, Palladino MJ, Witasp E, Kurnikov IV, Tyurina YY, Tyurin VA, Amoscato AA, Jiang J, Kochanek PM, DeKosky ST, Greenberger JS, Shvedova AA, Kagan VE (2006) Apoptotic interactions of cytochrome c: redox flirting with anionic phospholipids within and outside of mitochondria. Biochim Biophys Acta 1757:648–659
Belikova NA, Vladimirov YA, Osipov AN, Kapralov AA, Tyurin VA, Potapovich MV, Basova LV, Peterson J, Kurnikov IV, Kagan VE (2006) Peroxidase activity and structural transitions of cytochrome c bound to cardiolipin-containing membranes. Biochemistry 45:4998–5009
Bernardi P, Azzone GF (1981) Cytochrome c as an electron shuttle between the outer and inner mitochondrial membrane. J Biol Chem 256:7187–7192
Berry EA, Guergova-Kuras M, Huang LS, Crofts AR (2000) Structure and function of cytochrome bc complexes. Annu Rev Biochem 69:1005–1075
Bertini I, Cavallaro G, Rosato R (2005) A structural model for the adduct between cytochrome c and cytochrome c oxidase. J Biol Inorg Chem 10:613–624
Bertini I, Chevance S, Del Conte R, Lalli D, Turano P (2011a) The anti-apoptotic Bcl-xL protein, a new piece in the puzzle of cytochrome c interactome. PLos One. doi:10.1371/journal.pone.0018329
Bertini I, Cavallaro G, Rosato A (2011b) Principles and patterns in the interaction between mono-heme cytochrome c and its partners in electron transfer processes. Metallomics. doi:10.1039/c0mt00108b
Bihlmaier K, Mesecke N, Terzyiska N, Bien M, Hell K, Herrmann JM (2007) The disulfide relay system of mitochondria is connected to the respiratory chain. J Cell Biol 179:389–395
Cai J, Yang J, Jones DP (1998) Mitochondrial control of apoptosis: the role of cytochrome c. Biochim Biophys Acta 1366:139–149
Cassina AM, Hodara R, Souza JM, Thomson L, Castro L, Ischiropoulos H, Freeman BA, Radi R (2000) Cytochrome c nitration by peroxynitrite. J Biol Chem 275:21409–21415
Chance B, Sies H, Boveris A (1979) Hydroperoxide metabolism in mammalian organs. Physiol Rev 59:527–605
Chen Q, Vazquez EJ, Moghaddas S, Hoppel CL, Lesnefsky EJ (2003) Production of reactive oxygen species by mitochondria: central role of complex III. J Biol Chem 278:36027–36031
Chen YR, Chen CL, Chen W, Zweier JL, Augusto O, Radi R, Mason RP (2004) Formation of protein tyrosine ortho-semiquinone radical and nitrotyrosine from cytochrome c-derived tyrosyl radical. J Biol Chem 279:18054–18062
Cortese J, Voglino AL, Hackenbrock CR (1995) Persistence of cytochrome c binding to membranes at physiological mitochondrial intermembrane space ionic strength. Biochim Biophys Acta 1228:216–228
Dabir DV, Leverich EP, Kim S-K, Tsai FD, Hirasawa M, Knaff DB, Koehler CM (2007) A role for cytochrome c and cytochrome c peroxidase in electron shuttling from Erv1. EMBO J 26:4801–4811
Daum G (1985) Lipids of mitochondria. Biochim Biophys Acta 822:1–42
Deep S, Im S-C, Zuiderweg ERP, Waskell L (2005) Characterization and calculation of a cytochrome c–cytochrome b 5 complex using NMR data. Biochemistry 44:10654–10668
Demel RA, Jordi W, Lambrechts H, van Damme H, Hovius R, de Kruijff B (1989) Differential interactions of apo- and holocytochrome c with acidic membrane lipids in model systems and the implications for their import into mitochondria. J Biol Chem 264:3988–3997
Díaz-Moreno I, De La Rosa MA (2011a) Transient interactions between biomolecules. Eur Biophys J. doi:10.1007/s00249-011-0728-x
Díaz-Moreno I, De La Rosa MA (2011b) Transient interactions in metalloproteins. FEBS J. doi:10.1111/j.1742-4658.2011.08065.x
Díaz-Moreno I, García-Heredia JM, Díaz-Quintana A, Teixeira M, De La Rosa MA (2011c) Nitration of tyrosines 46 and 48 induces the specific degradation of cytochrome c upon change of the heme iron state to high-spin. Biochem Biophys Acta Bioenerg. doi:10.1016/j.bbabio.2011.09.012
Dudkina NV, Kudryashev M, Stahlberg H, Boekema EJ (2011) Interaction of complexes I, III, and IV within the bovine respirasome by single particle cryoelectron tomography. Proc Natl Acad Sci USA 108:15196–15200
Durham B, Fairris JL, McLean M, Millett F, Scott JR, Sligar SG, Willie A (1995) Electron transfer from cytochrome b 5 to cytochrome c. J Bioenerg Biomembr 27:331–340
Eble KS, Coleman WB, Hantgan RR, Cunningham CC (1990) Tightly associated cardiolipin in the bovine heart mitochondrial ATP synthase as analyzed by 31P nuclear magnetic resonance spectroscopy. J Biol Chem 265:19434–19440
Engstrom G, Rajagukguk R, Saunders AJ, Patel CN, Rajagukguk S, Merbitz-Zahradnik T, Xiao K, Pielak GJ, Trumpower B, Yu C-A, Yu L, Durham B, Millet F (2003) Design of a ruthenium-labeled cytochrome c derivative to study electron transfer with the cytochrome bc 1 complex. Biochemistry 42:2816–2824
Fry M, Green DE (1981) Cardiolipin requirement for electron transfer in complex I and III of the mitochondrial respiratory chain. J Biol Chem 256:1874–1880
García-Heredia JM, Díaz-Moreno I, Nieto PM, Orzáez M, Kocanis S, Teixeira M, Pérez-Payá E, Díaz-Quintana A, De la Rosa MA (2010) Nitration of tyrosine 74 prevents human cytochrome c to play a key role in apoptosis signaling by blocking caspase-9 activation. Biochem Biophys Acta Bioenerg 1797:981–993
García-Heredia JM, Díaz-Quintana A, Salzano M, Orzáez M, Pérez-Payá E, Teixeira M, De la Rosa MA, Díaz-Moreno I (2011) Tyrosine phosphorylation turns alkaline transition into a biologically relevant process and makes human cytochrome c behave as an anti-apoptotic switch. J Biol Inorg Chem. doi:10.1007/s00775-011-0804-9
Giles SS, Perfect JR, Cox GM (2005) Cytochrome c peroxidase contributes to the antioxidant defense of Cryptococcus neoformans. Fungal Genet Biol 42:20–29
Giorgio M, Migliaccio E, Orsini F, Paolucci D, Moroni M, Contursi C, Pelliccia G, Luzi L, Minucci S, Marcaccio M, Pinton P, Rizzuto R, Bernardi P, Paolucci F, Pelicci PG (2005) Electron transfer between cytochrome c and p66Shc generates reactive oxygen species that trigger mitochondrial apoptosis. Cell 122:221–233
Gomez B Jr, Robinson NC (1999) Phospholipase digestion of bound cardiolipin reversibly inactivates bovine cytochrome bc 1. Biochemistry 38:9031–9038
Haines TH, Dencher NA (2002) Cardiolipin: a proton trap for oxidative phosphorylation. FEBS Lett 528:35–39
Hannemann F, Guyot A, Zöllner A, Müller JJ, Heinemann U, Bernhardt R (2009) The dipole moment of the electron carrier adrenodoxin is not critical for redox partner interaction and electron transfer. J Inorg Biochem 103:997–1004
Heinemeyer J, Braun H-P, Boekema EJ, Kouřil R (2007) A structural model of the cytochrome c reductase/oxidase supercomplex from yeast mitochondria. J Biol Chem 282:12240–12248
Hüttemann M, Pecina P, Rainbolt M, Sanderson TH, Kagan VE, Samavati L, Doan JW, Lee I (2011a) The multiple functions of cytochrome c and their regulation in life and death decisions of the mammalian cell: from respiration to apoptosis. Mitochondrion 11:369–381
Hüttemann M, Helling S, Sanderson TH, Sinkler C, Samavati L, Mahapatra G, Varughese A, Lu G, Liu J, Ramzan R, Vogt S, Grossman LI, Doan JW, Marcus K, Lee I (2011b) Regulation of mitochondrial respiration and apoptosis through cell signaling: cytochrome c oxidase and cytochrome c in ischemia/reperfusion injury and inflammation. Biochim Biophys Acta. doi:10.1016/j.bbabio.2011.07.001
Jeng WY, Chen CY, Chang HC, Chuang WJ (2002) Expression and characterization of recombinant human cytochrome c in E. coli. J Bioenerg Biomembr 34:423–431
Jiang H, English AM (2006) Phenotypic analysis of the ccp1Δ and ccp1Δ-ccp1 W191F mutant strains of Saccharomyces cerevisiae indicates that cytochrome c peroxidase functions in oxidative-stress signaling. J Inorg Biochem 100:1996–2008
Kagan VE, Borisenko GG, Tyurina YY, Tyurin VA, Jiang J, Potapovich AI, Kini V, Amoscato AA, Fujii Y (2004) Oxidative lipidomics of apoptosis: redox catalytic interactions of cytochrome c with cardiolipin and phosphatidylserine. Free Rad Biol Med 37:1963–1985
Kagan VE, Tyurina YY, Batir H, Chu CT, Kapralov AA, Vlasova II, Belikova NA, Tyurin VA, Amoscato A, Epperly M, Greenberger J, DeKosky S, Shvedova AA, Jiang J (2006) The “pro-apoptotic genies” get out of mitochondria: oxidative lipidomics and redox activity of cytochrome c/cardiolipin complexes. Chem Biochem Interact 163:15–28
Kalanxhi E, Wallace CJ (2007) Cytochrome c impaled: investigation of the extended lipid anchorage of a soluble protein to mitochondrial membrane models. Biochem J 407:179–187
Kapralov AA, Yanamala N, Tyurina YY, Castro L, Samhan-Arias AK, Vladimirov YA, Maeda A, Mylnikov D, Demicheli V, Tortora V, Klein-Seetharaman J, Radi R, Kagan VE (2011) Topography of tyrosine residues and their involvement in peroxidation of polyunsaturated cardiolipin in cytochrome c/cardiolipin peroxidase complexes. Biochim Biophys Acta 1808:2147–2155
Kong SK, Yim MB, Stadtman ER, Chock PB (1996) Peroxynitrite disables the tyrosine phosphorylation regulatory mechanism: lymphocyte-specific tyrosine kinase fails to phosphorylate nitrated cdc2(6–20)NH2 peptide. Proc Natl Acad Sci USA 93:33777–33782
Lambeth JD, Kamin H (1979) Adrenodoxin reductase: adrenodoxin complex flavin to iron-sulfur transfer as the rate-limiting step in the NADPH-cytochrome c reductase reaction. J Biol Chem 254:2766–2774
Lange C, Hunte C (2002) Crystal structure of the yeast cytochrome bc 1 complex with its bound substrate cytochrome c. Proc Natl Acad Sci USA 99:2800–2805
Lee I, Salomon AR, Yu K, Doan JW, Grossman LI, Hüttemann M (2006) New prospects for an old enzyme: mammalian cytochrome c is tyrosine-phosphorylated in vivo. Biochemistry 47:9121–9128
Leesch VW, Bujous J, Mauk AG, Hoffman BM (2000) Cytochrome c peroxidase-cytochrome c complex: locating the second binding domain on cytochrome c peroxidase with site-directed mutagenesis. Biochemistry 39:10132–10139
Leferink NGH, van der Berg WAM, van Berkel WJH (2008) l-Galactono-γ-lactone dehydrogenase from Arabidopsis thaliana, a flavoprotein involved in vitamin C biosynthesis. FEBS J 275:713–726
Leferink NGH, Fraaije MW, Joosten H-J, Schaap PJ, Mattevi A, van Berkel WJH (2009) Identification of a gatekeeper residue that prevents dehydrogenases from acting as oxidases. J Biol Chem 284:4392–4397
Louie GV, Brayer GD (1990) High-resolution refinement of yeast iso-1-cytochrome c and comparisons with other eukaryotic cytochromes c. J Mol Biol 214:527–555
Matlib MA, O’Brien PJ (1976) Properties of rat liver mitochondria with intermembrane cytochrome c. Arch Biochem Biophys 173:27–33
Mei H, Geren L, Miller MA, Durham B, Millett F (2002) Role of the low-affinity binding site in electron transfer from cytochrome c to cytochrome c peroxidase. Biochemistry 41:3968–3976
Migliaccio E, Mele S, Salcini AE, Pelicci G, Lai K-MV, Superti-Furga G, Pawson T, Di Fiore PP, Lanfrancone L, Pelicci PG (1997) Opposite effects of the p52Shc/p46Shc and p66Shc splicing isoforms on the EGF receptor-MAP kinase-fos signalling pathway. EMBO J 16:706–716
Migliaccio E, Giorgio M, Mele S, Pelicci G, Reboldi P, Pandolfi PP, Lanfrancone L, Pelicci PG (1999) The p66Shc adaptor protein controls oxidative stress response and life span in mammals. Nature 402:309–313
Millett F, Miller MA, Geren L, Durham B (1995) Electron transfer between cytochrome c and cytochrome c peroxidase. J Bioenerg Biomembr 27:341–351
Moore GR, Pettigrew GW (1990) Cytochromes c. Evolutionary, structural, and physicochemical aspects. Springer, New York
Mustonen P, Virtanen JA, Somerharju PJ, Kinnunen PKJ (1987) Binding of cytochrome c to liposomes as revealed by the quenching of fluorescence from pyrene-labeled phospholipids. Biochemistry 26:2991–2997
Nyola A, Hunte C (2008) A structural analysis of the transient interaction between the cytochrome bc 1 complex and its substrate cytochrome c. Biochem Soc Trans 36:981–985
Orrenius S (2007) Reactive oxygen species in mitochondria-mediated cell death. Drugs Metab Res 39:443–455
Orsini F, Migliaccio E, Moroni M, Contursi C, Raker VA, Piccini D, Martin-Padura I, Pelliccia G, Trinei M, Bono M, Puri C, Tacchetti C, Ferrini M, Mannucci R, Nicoletti I, Lanfrancone L, Giorgio M, Pelicci PG (2004) The life span determinant p66Shc localizes to mitochondria where it associates with mitochondrial heat shock protein 70 and regulates trans-membrane potential. J Biol Chem 279:25689–25695
Ott M, Robertson J, Gogvadze V, Zhivotovsky B, Orrenius S (2002) Cytochrome c release from mitochondria proceeds by a two-step process. Proc Natl Acad Sci USA 99:1259–1263
Oursler MJ, Bradley EW, Elfering SL, Giulivi C (2005) Native, not nitrated, cytochrome c and mitochondria-derived hydrogen peroxide drive osteoclast apoptosis. Am J Physiol Cell Physiol 288:156–168
Papa S, Capitanio N, Capitanio G (2004) A cooperative model for proton pumping in cytochrome c oxidase. Biochem Biophys Acta 1655:353–364
Pearl NM, Jacobson T, Arisa M, Vitello LB, Erman JE (2007) Effect of single-site charge-reversal mutations on the catalytic properties of yeast cytochrome c peroxidase: mutations near the high-affinity cytochrome c binding site. Biochemistry 46:8263–8272
Pearl NM, Jacobson T, Meyen C, Clementz AG, Ok EY, Choi E, Wilson K, Vitello LB, Erman JE (2008) Effect of single-site charge-reversal mutations on the catalytic properties of yeast cytochrome c peroxidase: evidence for a single, catalytically active, cytochrome c binding domain. Biochemistry 47:2766–2775
Pecina P, Borisenko GG, Belikova NA, Tyurina Y, Pecinova A, Lee I, Samhan-Arias AK, Przyklenk K, Kagan VE, Huttemann M (2010) Phosphomimetic substitution of cytochrome c tyrosine 48 decreases respiration and binding to cardiolipin and abolishes ability to trigger downstream caspase activation. Biochemistry 49:6705–6710
Pelicci G, Lanfrancone L, Grignani F, McGlade J, Cavallo F, Forni G, Nicoletti I, Grignani F, Pawson T, Pelicci PG (1992) A novel transforming protein (SHC) with an SH2 domain is implicated in mitogenic signal transduction. Cell 70:93–104
Pellegrini M, Pacini S, Baldari CT (2005) p66Shc: the apoptotic side of Shc proteins. Apoptosis 10:13–18
Pelletier H, Kraut J (1992) Crystal structure of a complex between electron transfer partners, cytochrome c peroxidase and cytochrome c. Science 258:1748–1755
Pereverzev MO, Vygodina TV, Konstantinov AA, Skulachev VP (2003) Cytochrome c, an ideal antioxidant. Biochem Soc Trans 31:1312–1315
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Pettigrew GW, Prazeres S, Costa C, Palma N, Krippahl L, Moura I, Moura JJG (1999) The structure of an electron transfer complex containing a cytochrome c and a peroxidase. J Biol Chem 274:11383–11389
Pinton P, Rimessi A, Marchi S, Orsini F, Migliaccio E, Giorgio M, Contursi C, Minucci S, Mantovani F, Wieckowski MR, Del Sal G, Pelicci PG, Rizzuto R (2007) Protein kinase C β and prolyl isomerase 1 regulate mitochondrial effects of the life-span determinant p66Shc. Science 315:659–663
Poulos TL, Freer ST, Alden RA, Edwards SJ, Skoglands U, Takio K, Eriksson B, Xuong N-H, Yonetani T, Kraut J (1980) The crystal structure of cytochrome c peroxidase. J Biol Chem 255:575–580
Prudêncio M, Ubbink M (2004) Transient complexes of redox proteins: structural and dynamic details from NMR studies. J Mol Recognit 17:524–539
Reincke B, Perez C, Pristovsek P, Lucke C, Ludwig C, Lohr F, Rogov VV, Ludwig B, Ruterjans H (2001) Solution structure and dynamics of the functional domain of Paracoccus denitrificans cytochrome c(552) in both redox states. Biochemistry 40:12312–12320
Riemer J, Fischer M, Hermann JM (2011) Oxidation-driven protein import into mitochondria: insights and blind spots. Biochim Biophys Acta 1808:981–989
Roberts VA, Pique ME (1999) Definition of the interaction domain for cytochrome c on cytochrome c oxidase. J Biol Chem 274:38051–38060
Robinson NC (1993) Functional binding of cardiolipin to cytochrome c oxidase. J Bioenerg Biomembr 25:153–163
Rödiger A, Baudisch B, Langner U, Klösgen RB (2011) Dual targeting of a mitochondrial protein: the case study of cytochrome c 1. Mol Plant 4:679–687
Rodríguez-Roldán V, García-Heredia JM, Navarro JA, De la Rosa MA, Hervás M (2008) Effect of nitration on the physicochemical and kinetic features of wild-type and mono-tyrosine mutants of human respiratory cytochrome c. Biochemistry 47:12371–12379
Rytömaa M, Mustonen P, Kinnunen PK (1992) Reversible, nonionic, and pH-dependent association of cytochrome c with cardiolipin-phosphatidylcholine liposomes. J Biol Chem 267:22243–22248
Sakamoto K, Kamiya M, Imai M, Shinzawa-Itoh K, Uchida T, Kawano K, Yoshikawa S, Ishimori K (2011) NMR basis for interprotein electron transfer gating between cytochrome c and cytochrome c oxidase. Proc Natl Acad Sci USA 108:12271–12276
Saraste M (1999) Oxidative phosphorylation at the fin de siecle. Science 283:1488–1493
Schug ZT, Gottlieb E (2009) Cardiolipin acts as a mitochondrial signaling platform to launch apoptosis. Biochim Biophys Acta 1788:2022–2031
Scorolli L, Meduri A, Morara M, Scalinci SZ, Meduri RA (2007) Effect of cytochrome c peroxidase on the corneal epithelial healing process after excimer laser photoablation in transgenic mice. Eur Surg Res 39:82–87
Shao W, Im S-C, Zuiderweg ERP, Waskell L (2003) Mapping the binding interface of the cytochrome b 5–cytochrome c complex by nuclear magnetic resonance. Biochemistry 42:14774–14784
Shidoji Y, Hayashi K, Komura S, Ohishi N, Yagi K (1999) Loss of molecular interaction between cytochrome c and cardiolipin due to lipid peroxidation. Biochem Biophys Res Commun 264:343–347
Sinibaldi F, Fiorucci L, Patriarca A, Lauceri R, Ferri T, Coletta M, Santucci R (2008) Insights into cytochrome c-cardiolipin interaction. Role played by ionic strength. Biochemistry 47:6928–6935
Sinibaldi F, Howes BD, Piro MC, Polticelli F, Bombelli C, Ferri T, Coletta M, Smulevich G, Santucci R (2010) Extended cardiolipin anchorage to cytochrome c: a model for protein-mitochondrial membrane binding. J Biol Inorg Chem 15:689–700
Solmaz SRN, Hunte C (2008) Structure of complex III with bound cytochrome c in reduced state and definition of a minimal core interface for electron transfer. J Biol Chem 283:17542–17549
Stemp EDA, Hoffman BM (1993) Cytochrome c peroxidase binds two molecules of cytochrome c: evidence for a low-affinity, electron transfer-active site on cytochrome c peroxidase. Biochemistry 32:10848–10865
Stepanov G, Gnedenko O, Mol’nar A, Ivanov A, Vladimirov Y, Osipov A (2009) Evaluation of cytochrome c affinity to anionic phospholipids by means of surface plasmon resonance. FEBS Lett 583:97–100
Sun Y-L, Wang Y-H, Yan M-M, Sun B-Y, Xie Y, Huang Z-X, Jiang S-K, Wu H-M (1999) Structure, interaction and electron transfer between cytochrome b 5, its E44A and/or E56A mutants and cytochrome c. J Mol Biol 285:347–359
Sun Y-L, Wang Y-H, Qian C, Lu J, Li E, Wang W, Lu J, Xie Y, Wang J, Zhu D, Huang Z-X, Tang W (2001) Solution structure of cytochrome b 5 mutant (E44/48/56A/D60A) and its interaction with cytochrome c. Eur J Biochem 268:1620–1630
Świerczek M, Cieluch E, Sarewicz M, Borek A, Moser CC, Dutton PL, Osyczka A (2010) An electronic bus bar lies in the core of cytochrome bc 1. Science 329:451–454
Tian H, Sadoski R, Zhang L, Yu C-A, Yu L, Durham B, Millet F (2000) Definition of the interaction domain for cytochrome c on the cytochrome bc(1) complex. Steady-state and rapid kinetic analysis of electron transfer between cytochrome c and Rhodobacter sphaeroides cytochrome bc(1) surface mutants. J Biol Chem 275:9587–9595
Trinei M, Giorgio M, Cicalese A, Barozzi S, Ventura A, Migliaccio E, Milia E, Martin Padura I, Raker VA, Maccarana M, Petronilli V, Minucci S, Bernardi P, Lanfrancone L, Pelicci PG (2002) A p53-p66Shc signalling pathway controls intracellular redox status, levels of oxidation-damaged DNA and oxidative stress-induced apoptosis. Oncogene 21:3872–3878
Tsukihara T, Aoyama H, Yamashita E, Tomizaki T, Yamaguchi H, Shinzawa-Itoh K, Nakashima R, Yaono R, Yoshikawa S (1996) The whole structure of the 13-subunit oxidized cytochrome c oxidase at 2.8 Ǻ. Science 272:1136–1144
Tuominen EKJ, Wallace CJA, Kinnunen PVJ (2002) Phospholipid-cytochrome c interaction: evidence for the extended lipid anchorage. J Biol Chem 277:8822–8826
Ubbink M (2009) The courtship of proteins: understanding the encounter complex. FEBS Lett 583:1060–1066
Ventura A, Maccarana M, Raker VA, Pelicci PG (2004) A cryptic targeting signal induces isoform-specific localization of p46Shc to mitochondria. J Biol Chem 279:2299–2306
Vik SB, Georgevich G, Capaldi RA (1981) Diphosphatidylglycerol is required for optimal activity of beef cytochrome c oxidase. Proc Natl Acad Sci USA 78:1456–1460
Volkov AN, Ferrari D, Worrall JAR, Bonvin AMJJ, Ubbink M (2005) The orientations of cytochrome c in the highly dynamic complex with cytochrome b 5 visualized by NMR and docking using HADDOCK. Prot Sci 14:799–811
Volkov AN, Worrall JAR, Holtzmann E, Ubbink M (2006) Solution structure and dynamics of the complex between cytochrome c and cytochrome c peroxidase determined by paramagnetic NMR. Proc Natl Acad Sci USA 103:18945–18950
Volkov AN, Bashir Q, Worrall JAR, Ubbink M (2009) Binding hot spot in the weak protein complex of physiological redox partners yeast cytochrome c and cytochrome c peroxidase. J Mol Biol 385:1003–1013
Volkov AN, Bashir Q, Worrall JAR, Ullmann GM, Ubbink M (2010) Shifting the equilibrium between the encounter state and the specific form of a protein complex by interfacial point mutations. J Am Chem Soc 132:11487–11495
Volkov AN, Nicholls P, Worrall JAR (2011) The complex of cytochrome c and cytochrome c peroxidase. The end of the road? Biochim Biophys Acta. doi:10.1016/j.bbabio.2011.07.010
Wang K, Mei H, Geren L, Miller MA, Saunders A, Wang X, Waldner JL, Pielak GJ, Durham B, Millett F (1996a) Design of a ruthenium-cytochrome c derivative to measure electron transfer to the radical cation and oxyferryl heme in cytochrome c peroxidase. Biochemistry 35:15107–15119
Wang J, Larsen RW, Moench SJ, Satterlee JD, Rousseau DL, Ondrias MR (1996b) Cytochrome c peroxidase complexed with cytochrome c has an unperturbed heme moiety. Biochemistry 35:453–463
Witt H, Malatesta F, Nicoletti F, Brunori M, Ludwig B (1998) Tryptophan 121 of subunit II is the electron entry site to cytochrome-c oxidase in Paracoccus denitrificans. Involvement of a hydrophobic patch in the docking reaction. J Biol Chem 273:5132–5136
Worrall JAR, Kolczak U, Canters GW, Ubbink M (2001) Interaction of yeast iso-1-cytochrome c with cytochrome c peroxidase investigated by [15N, 1H] heteronuclear NMR spectroscopy. Biochemistry 40:7069–7076
Worrall JAR, Reinle W, Bernhardt R, Ubbink M (2003) Transient protein interactions studied by NMR spectroscopy: the case of cytochrome c and adrenodoxin. Biochemistry 42:7068–7076
Xu X, Reinle W, Hannemann F, Konarev PV, Svergun DI, Bernhardt R, Ubbink M (2008) Dynamics in a pure encounter complex of two proteins studied by solution scattering and paramagnetic NMR spectroscopy. J Am Chem Soc 130:6395–6403
Yonetani T, Ohnishi T (1966) Cytochrome c peroxidase, a mitochondrial enzyme of yeast. J Biol Chem 241:2983–2984
Yoshikawa S, Shinzawa-Itoh K, Nakashima R, Yaono R, Yamashita E, Inoue N, Yao M, Jie Fei M, Libeu CP, Mizushima T, Yamaguchi H, Tomizaki T, Tsukihara T (1998) Redox coupled crystal structural changes in bovine heart cytochrome c oxidase. Science 280:1723–1729
Yu C-A, Wen X, Xiao K, Xi D, Yu L (2002) Inter- and intra-molecular electron transfer in the cytochrome bc 1 complex. Biochem Biophys Acta 1555:65–70
Yu H, Lee I, Salomon AR, Yu K, Hüttemann M (2008) Mammalian liver cytochrome c is tyrosine-48 phosphorylated in vivo, inhibiting mitochondrial respiration. Biochem Biophys Acta 1777:1066–1071
Zhao X, Leon IR, Bak S, Mogensen M, Wrzesinski K, Hojlund K, Jensen ON (2011) Phosphoproteome analysis of functional mitochondria isolated from resting human muscle reveals extensive phosphorylation of inner membrane protein complexes and enzymes. Mol Cell Proteomics M110.000299
Zhou JS, Hoffmann BM (1993) Cytochrome c peroxidase simultaneously binds cytochrome c at two different sites with strikingly different reactivities: titrating a “substrate” with an enzyme. J Am Chem Soc 115:11008–11009
Zhou JS, Hoffmann BM (1994) Stern-volmer in reverse: 2:1 stoichiometry of the cytochrome c–cytochrome c peroxidase electron-transfer complex. Science 265:1693–1696
Zhu Y, Li M, Wang X, Jin H, Liu S, Xu J, Chen Q (2011) Caspase cleavage of cytochrome c 1 disrupts mitochondrial function and enhances cytochrome c release. Cell Res. doi:10.1038/cr.2011.82
Acknowledgments
The authors wish to thank Jonathan Martínez-Fábregas for helpful advice and critical reading of the manuscript. This work was funded by the Spanish Ministry of Science and Innovation (BFU2009-07190) and the Andalusian Government (BIO198 and P08-CVI-3876).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Special Issue: Transient interactions in biology.
Rights and permissions
About this article
Cite this article
Díaz-Moreno, I., García-Heredia, J.M., Díaz-Quintana, A. et al. Cytochrome c signalosome in mitochondria. Eur Biophys J 40, 1301–1315 (2011). https://doi.org/10.1007/s00249-011-0774-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-011-0774-4