Abstract
In this paper we provide a detailed biochemical and structural characterization of the active site of recombinant human prolidase, a dimeric metalloenzyme, whose misfunctioning causes a recessive connective tissue disorder (prolidase deficiency) characterized by severe skin lesions, mental retardation and respiratory tract infections. It is known that the protein can host two metal ions in the active site of each constituent monomer. We prove that two different kinds of metals (Mn and Zn) can be simultaneously present in the protein active sites with the protein partially maintaining its enzymatic activity. Structural information extracted from X-ray absorption spectroscopy measurements have been used to yield a full reconstruction of the atomic environment around each one of the two monomeric active sites. In particular, as for the metal ion occupation configuration of the recombinant human prolidase, we have found that one of the two active sites is occupied by two Zn ions and the second one by one Zn and one Mn ion. In both dinuclear units a histidine residue is bound to a Zn ion.






Similar content being viewed by others
Notes
The weight of the recombinant dimeric prolidase is 123,600 Da (Lupi et al. 2008).
The relation between the measured absorption coefficient, μ(E), and the derived signal χ(k) is given in Appendix.
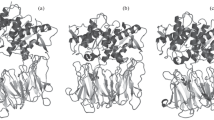
As already commented in “Introduction,” we indicated that we have decided to employ the 3D crystallographic data of P. horikoshii OT3 as for the latter (unlike P. furiosus) both monomers have been resolved.
See the web site: http://webusers.fis.uniroma3.it/~meneghini/. For recent application see (Maret et al. 2005).
See Error Reporting Recommendations: A Report of the Standards and Criteria Committee (2002), URL http://ixs.csrri.iit.edu/
Abbreviations
- APPro:
-
Aminopeptidase proline
- Asp:
-
Aspartic acid
- DESY:
-
Deutsches Elektronen-Synchrotron
- DTT:
-
Dithiothreitrol
- DW:
-
Debye–Waller
- EMBL:
-
European Molecular Biology Laboratory
- EXAFS:
-
Extended X-ray absorption fine structure
- FT:
-
Fourier transform
- Glu:
-
Glutamic acid
- Gly:
-
Glycine
- His:
-
Histidine
- ICP-MS:
-
Inductively coupled plasma-mass spectrometry
- MetAP:
-
Methionine aminopeptidase
- MS:
-
Multiple scattering
- PDB:
-
Protein Data Bank
- PEPD:
-
Peptidase D: prolidase gene
- Pfprol:
-
Pyrococcus furiosus prolidase
- XAS:
-
X-ray absorption spectroscopy
- Pro:
-
Proline
- XANES:
-
X-ray absorption near edge structure
References
Benfatto M, Natoli CR, Bianconi A, Garcia J, Marcelli A, Fanfoni M, Davoli I (1986) Multiple-scattering regime and higher-order correlations in X-ray-absorption spectra of liquid solutions. Phys Rev B 34:5774–5781. doi:10.1103/PhysRevB.34.5774
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242. doi:10.1093/nar/28.1.235
Bianconi A, Congiu-Castellano A, Dell’Ariccia M, Giovannelli A, Morante S, Burattini E, Durham PJ (1986) Local Fe site structure in the tense-to-relaxed transition in carp deoxyhemoglobin: a XANES (X-ray absorption near edge structure) study. Proc Natl Acad Sci USA 83:7736–7740. doi:10.1073/pnas.83.20.7736
Boland JJ, Crane SE, Baldeschwieler JD (1982) Theory of extended X-ray absorption fine structure: single and multiple scattering formalisms. J Chem Phys 77:142–153. doi:10.1063/1.443662
Browne P, O’Cuinn G (1983) The purification and characterization of a proline dipeptidase from guinea pig brain. J Biol Chem 258:6147–6154
Chinard FP (1952) Photometric estimation of praline and ornithine. J Biol Chem 199:91–95
Cosper NJ, D’souza VM, Scott RA, Holz RC (2001) Structural evidence that the methionyl aminopeptidase from Escherichia coli is a mononuclear metalloprotease. Biochemistry 40:13302–13309. doi:10.1021/bi010837m
Cunningham DF, O’Connor B (1997) Proline specific peptidases. Biochim Biophys Acta 1343:160–186
D’souza VM, Bennet B, Copik AJ, Holz RC (2000) Divalent metal binding properties of the methionyl aminopeptidase from Escherichia coli. Biochemistry 39:3817–3826. doi:10.1021/bi9925827
Du X, Tove S, Kast-Hutcheson K, Grunden AM (2005) Characterization of the dinuclear metal center of Pyrococcus furiosus prolidase by analysis of targeted mutants. FEBS Lett 579:6140–6146. doi:10.1016/j.febslet.2005.09.086
Fernandez-Espla MD, Martin-Hernandez MC, Fox PF (1997) Purification and characterization of a prolidase from Lactobacillus casei subsp. casei IFPL 731. Appl Environ Microbiol 63:314–316
Fujii M, Nagaoka Y, Imamura S, Shimizu T (1996) Purification and characterization of a prolidase from Aureobacterium esteraromaticum. Biosci Biotechnol Biochem 60:1118–1122
Ghosh M, Grunden AM, Dunn DM, Weiss R, Adams MW (1998) Characterization of native and recombinant forms of an unusual cobalt-dependent proline dipeptidase (prolidase) from the hyperthermophilic archaeon Pyrococcus furiosus. J Bacteriol 180:4781–4789
Gurman SJ, Binsted N, Ross I (1986) A rapid, exact, curved-wave theory for EXAFS calculations II. The multiple-scattering contributions. J Phys Chem 19:1845–1861
James F (1994) MINUIT: function minimization and error analysis reference manual version 94.1, CERN Program Library D506
Kobayashi M, Shimizu S (1999) Cobalt proteins. Eur J Biochem 261:1–9. doi:10.1046/j.1432-1327.1999.00186.x
Koningsberger DC, Prins R (eds) (1988) X-ray absorption. Principles, applications, techniques of EXAFS, SEXAFS and XANES. Wiley, New York
Korbas M, Fulla Marsa D, Meyer-Klaucke W (2006) KEMP: a program script for automated biological X-ray absorption spectroscopy data reduction. Rev Sci Instrum 77:063105. doi:10.1063/1.2209954
Lee PA, Pendry JB (1975) Theory of the extended X-ray absorption fine structure. Phys Rev B 11:2795–2811. doi:10.1103/PhysRevB.11.2795
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the folin phenol reagent. J Biol Chem 193:265–275
Lowther WT, Matthews BW (2002) Metalloaminopeptidases: common functional themes in disparate structural surroundings. Chem Rev 102:4581–4608. doi:10.1021/cr0101757
Lupi A, Della Torre S, Campari E, Tenni R, Cetta G, Rossi A, Forlino A (2006) Human recombinant prolidase from eukaryotic and prokaryotic sources Expression, purification, characterization and long-term stability studies. FEBS J 273:5466–5478. doi:10.1111/j.1742-4658.2006.05538.x
Lupi A, Tenni R, Rossi A, Cetta G, Forlino A (2008) Human prolidase and prolidase deficiency: an overview on the characterization of the enzyme involved in proline recycling and on the effects of its mutations. Amino Acids 35:739–752. doi:10.1007/s00726-008-0055-4
Maher MJ, Ghosh M, Grunden AM, Menon AL, Adams MW, Freeman HC, Guss JM (2004) Structure of the prolidase from Pyrococcus furiosus. Biochemistry 43:2771–2783. doi:10.1021/bi0356451
Maret M, Bley F, Meneghini C, Albrecht M, Khler J, Bucher E, Hazemann JL (2005) The Cr local structure in epitaxial CrPt3(111) films probed using polarized X-ray absorption fine structure. J Phys Condens Matter 17:2529–2541. doi:10.1088/0953-8984/17/17/001
Meneghini C, Morante S (1998) The active site structure of tetanus neurotoxin resolved by multiple scattering analysis in X-ray absorption spectroscopy. Biophys J 75:1953–1963. doi:10.1016/S0006-3495(98)77636-2
Myara I, Charpentier C, Lemonnier A (1982) Optimal conditions for prolidase assay by proline calorimetric determination: application to iminodipeptiduria. Clin Chim Acta 125:193–205. doi:10.1016/0009-8981(82)90196-6
O’Cuinn G, Fortrell PF (1975) Purification and characterization of an aminoacyl proline hydrolase from guinea-pig intestinal mucosa. Biochim Biophys Acta 391:388–395
Ravel B, Newville M (2005) ATHENA, ARTEMIS, HEPHAESTUS: data analysis for X-ray absorption spectroscopy using IFEFFIT. J Sync Rad 12:537–541. doi:10.1107/S0909049505012719
Rehr JJ, Albers RC (1990) Scattering-matrix formulation of curved-wave multiple-scattering theory: application to X-ray-absorption fine structure. Phys Rev B 41:8139–8149. doi:10.1103/PhysRevB.41.8139
Roderick SL, Matthews BW (1993) Structure of the cobalt-dependent methionine aminopeptidase from Escherichia coli: a new type of proteolytic enzyme. Biochemistry 32:3907–3912. doi:10.1021/bi00066a009
Royce PM, Steinmann B (2002) Prolidase deficiency. In: Royce PM, Steinmann B (eds) Connective tissue and its heritable disorders. Wiley-Liss, New York
Tahirov TH, Oki H, Tsukihara T, Ogasahara K, Yutani K, Ogata K, Izu Y, Tsunasawa S, Kato I (1998) Crystal structure of methionine aminopeptidase from hyperthermophile, Pyrococcus furiosus. J Mol Biol 284:101–124. doi:10.1006/jmbi.1998.2146
Teo BK, Lee PA (1979) Ab initio calculations of amplitude and phase functions for extended X-ray absorption fine structure spectroscopy. J Am Chem Soc 101:2815–2832. doi:10.1021/ja00505a003
Yang SI, Tanaka T (2008) Characterization of recombinant prolidase from Lactococcus lactis—changes in substrate specificity by metal cations and allosteric behaviour of the peptides. FEBS J 275:271–280. doi:10.1111/j.1742-4658.2008.06344.x
Yoshimoto T, Matsubara F, Kawano E, Tsuru D (1983) Prolidase from bovine intestine: purification and characterization. J Biochem 94:1889–1896
Wilcox DE (1996) Binuclear metallohydrolases. Chem Rev 96:2435–2458. doi:10.1021/cr950043b
Willingham K, Maher MJ, Grunden AM, Ghosh M, Adams MW, Freeman HC, Guss JM (2001) Crystallization and characterization of the prolidase from Pyrococcus furiosus. Acta Crystallogr D Biol Crystallogr 57:428–430. doi:10.1107/S0907444900020187
Zabinsky SI, Rehr JJ, Ankudinov A, Albers RC, Eller MJ (1995) Multiple scattering calculations of X-ray absorption spectra. Phys Rev B 52:2995–3009. doi:10.1103/PhysRevB.52.2995
Acknowledgments
We wish to thank G.C. Rossi for a careful reading of the manuscript. V. Minicozzi, S. Morante and F. Stellato gratefully acknowledge INFN (Italy) for partial financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Proceedings of the XIX Congress of the Italian Society of Pure and Applied Biophysics (SIBPA), Rome, September 2008.
S. Alleva is the co-first author.
Appendix
Appendix
The EXAFS signal,\( \chi (k ) , \) is defined by the formula
where \( \mu (k ) \) is the measured total absorption coefficient and \( \mu_{0} (k ) \) is the absorption coefficient of the isolated absorber. k is the wave vector of the extracted electron which is related to the incident photon energy, \( E, \) and the ionization energy, \( E_{0} , \) by the obvious relation
with m the electron mass and ħ the (reduced) Planck constant.
For the reader’s convenience we recall the theoretical formula describing the EXAFS signal. We give it for simplicity in the single scattering approximation which is valid for photon energies sufficiently higher than the threshold energy. Following the standard notation of Boland et al. (1982), it reads
where the sum runs over the different coordination shells around the absorber. N l is the number of scatterers of the l-th shell, located at a distance r l from the absorber and \( \sigma_{l}^{2} \) is the Debye–Waller factor. \( \left| {f_{l} (k,{{\uppi}})} \right| \) is the modulus of the back-scattering amplitude and \( \phi_{l} (k) \) the total scattering phase. \( S_{0}^{2} \) is an empirical quantity that accounts for all the many-body losses in photo-absorption processes and \( \lambda (k) \) is the photo-electron mean free path. In cases where also MS processes become important a formally similar expression holds in which, however, r l represents the length of the full MS path. Modulus and phase functions are in this case rather complicated expressions which depend on all the scattering events occurring along the MS path (see Lee and Pendry 1975; Benfatto et al. 1986; Gurman et al. 1986; Koningsberger and Prins 1988; Rehr and Albers 1990).
The quantitative analysis of the structural EXAFS signals is performed with the help of the FITEXA code, developed by one of the authors (C. Meneghini).Footnote 4 FITEXA exploits the FEFF8.2 package (Zabinsky et al. 1995) for the calculation of backscattering amplitudes, total phase shifts and photoelectron mean free path. Best fit is achieved by minimizing the quantity
where \( \chi_{ \exp } \) and \( \chi_{\text{th}} \) are the experimental and theoretical data, respectively, and the sum is over the number, N p, of the values of k at which data were collected. The MINUIT (James 1994) subroutine of CERN library is used for data refinement and statistical error analysis.
The fit quality is measured by the associated R 2-factor defined by the formulaFootnote 5
A value of R 2 of about 10% is to be considered satisfactory for complex biological molecules.
Rights and permissions
About this article
Cite this article
Besio, R., Alleva, S., Forlino, A. et al. Identifying the structure of the active sites of human recombinant prolidase. Eur Biophys J 39, 935–945 (2010). https://doi.org/10.1007/s00249-009-0459-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-009-0459-4