Abstract
The influence of temporal and spatial variations on the microbial community composition was assessed in the unique coastal mangrove of Sundarbans using parallel 16S rRNA gene pyrosequencing. The total sediment DNA was extracted and subjected to the 16S rRNA gene pyrosequencing, which resulted in 117 Mbp of data from three experimental stations. The taxonomic analysis of the pyrosequencing data was grouped into 24 different phyla. In general, Proteobacteria were the most dominant phyla with predominance of Deltaproteobacteria, Alphaproteobacteria, and Gammaproteobacteria within the sediments. Besides Proteobacteria, there are a number of sequences affiliated to the following major phyla detected in all three stations in both the sampling seasons: Actinobacteria, Bacteroidetes, Planctomycetes, Acidobacteria, Chloroflexi, Cyanobacteria, Nitrospira, and Firmicutes. Further taxonomic analysis revealed abundance of micro-aerophilic and anaerobic microbial population in the surface layers, suggesting anaerobic nature of the sediments in Sundarbans. The results of this study add valuable information about the composition of microbial communities in Sundarbans mangrove and shed light on possible transformations promoted by bacterial communities in the sediments.





Similar content being viewed by others
References
Schmitt S, Tsai P, Bell J, Fromont J, Ilan M, Lindquist N, Perez T, Rodrigo A, Schupp PJ, Vacelet J, Webster N, Hentschel U, Taylor MW (2012) Assessing the complex sponge microbiota: core, variable and species-specific bacterial communities in marine sponges. ISME J 6:564–576
Godoy-Vitorino F, Goldfarb KC, Karaoz U, Leal S, Garcia-Amado MA, Hugenholtz P, Tringe SG, Brodie EL, Dominguez-Bello MG (2012) Comparative analyses of foregut and hindgut bacterial communities in hoatzins and cows. ISME J 6:531–541
Sogin ML, Morrison HG, Huber JA, Mark Welch D, Huse SM, Neal PR, Arrieta JM, Herndl GJ (2006) Microbial diversity in the deep sea and the underexplored “rare biosphere”. Proc Natl Acad Sci U S A 103:12115–12120
Gomes NC, Cleary DF, Calado R, Costa R (2011) Mangrove bacterial richness. Commun Integr Biol 4:419–423
Alongi DM (1988) Bacterial productivity and microbial biomass in tropical mangrove sediments. Microb Ecol 15:59–79
Holguin G, Gonzalez-Zamorano P, De-Bashan LE, Mendoza R, Amador E, Bashan Y (2006) Mangrove health in an arid environment encroached by urban development—a case study. Sci Total Environ 363:260–274
Duke NC, Meynecke JO, Dittmann S, Ellison AM, Anger K, Berger U, Cannicci S, Diele K, Ewel KC, Field CD, Koedam N, Lee SY, Marchand C, Nordhaus I, Dahdouh-Guebas F (2007) A world without mangroves? Science 317:41–42
Bouchez A, Pascault N, Chardon C, Bouvy M, Cecchi P, Lambs L, Herteman M, Fromard F, Got P, Leboulanger C (2013) Mangrove microbial diversity and the impact of trophic contamination. Mar Pollut Bull 66:39–46
Feller IC, Lovelock CE, Berger U, McKee KL, Joye SB, Ball MC (2010) Biocomplexity in mangrove ecosystems. Ann Rev Mar Sci 2:395–417
Giri C, Ochieng E, Tieszen LL, Zhu Z, Singh A, Loveland T, Masek J, Duke N (2011) Status and distribution of mangrove forests of the world using earth observation satellite data. Glob Ecol Biogeogr 20:154–159
Gomes NC, Flocco CG, Costa R, Junca H, Vilchez R, Pieper DH, Krogerrecklenfort E, Paranhos R, Mendonca-Hagler LC, Smalla K (2010) Mangrove microniches determine the structural and functional diversity of enriched petroleum hydrocarbon-degrading consortia. FEMS Microbiol Ecol 74:276–290
Ghosh A, Dey N, Bera A, Tiwari A, Sathyaniranjan K, Chakrabarti K, Chattopadhyay D (2010) Culture independent molecular analysis of bacterial communities in the mangrove sediment of Sundarban, India. Saline Syst 6:1
Yan B, Hong K, Yu ZN (2006) Archaeal communities in mangrove soil characterized by 16S rRNA gene clones. J Microbiol 44:566–571
Taketani RG, Yoshiura CA, Dias AC, Andreote FD, Tsai SM (2010) Diversity and identification of methanogenic archaea and sulphate-reducing bacteria in sediments from a pristine tropical mangrove. Antonie Van Leeuwenhoek 97:401–411
Dias AC, Pereira Silva Mde EC, Cotta SR, Dini-Andreote F, Soares FL Jr, Salles JF, Azevedo JL, Van Elsas JD, Andreote FD (2012) Abundance and genetic diversity of nifH gene sequences in anthropogenically affected Brazilian mangrove sediments. Appl Environ Microbiol 78:7960–7967
Zhang Y, Dong J, Yang Z, Zhang S, Wang Y (2008) Phylogenetic diversity of nitrogen-fixing bacteria in mangrove sediments assessed by PCR-denaturing gradient gel electrophoresis. Arch Microbiol 190:19–28
Marcial Gomes NC, Borges LR, Paranhos R, Pinto FN, Mendonca-Hagler LC, Smalla K (2008) Exploring the diversity of bacterial communities in sediments of urban mangrove forests. FEMS Microbiol Ecol 66:96–109
Dos Santos HF, Cury JC, Do Carmo FL, Dos Santos AL, Tiedje J, Van Elsas JD, Rosado AS, Peixoto RS (2011) Mangrove bacterial diversity and the impact of oil contamination revealed by pyrosequencing: bacterial proxies for oil pollution. PLoS ONE 6:e16943
Peixoto R, Chaer GM, Carmo FL, Araujo FV, Paes JE, Volpon A, Santiago GA, Rosado AS (2011) Bacterial communities reflect the spatial variation in pollutant levels in Brazilian mangrove sediment. Antonie Van Leeuwenhoek 99:341–354
Taketani RG, Franco NO, Rosado AS, van Elsas JD (2010) Microbial community response to a simulated hydrocarbon spill in mangrove sediments. J Microbiol 48:7–15
Brito EM, Guyoneaud R, Goni-Urriza M, Ranchou-Peyruse A, Verbaere A, Crapez MA, Wasserman JC, Duran R (2006) Characterization of hydrocarbonoclastic bacterial communities from mangrove sediments in Guanabara Bay, Brazil. Res Microbiol 157:752–762
Andreote FD, Jimenez DJ, Chaves D, Dias AC, Luvizotto DM, Dini-Andreote F, Fasanella CC, Lopez MV, Baena S, Taketani RG, de Melo IS (2012) The microbiome of Brazilian mangrove sediments as revealed by metagenomics. PLoS ONE 7:e38600
Gilbert JA, Meyer F, Bailey MJ (2011) The future of microbial metagenomics (or is ignorance bliss?). ISME J 5:777–779
Manna S, Chaudhuri K, Bhattacharyya S, Bhattacharyya M (2010) Dynamics of Sundarban estuarine ecosystem: eutrophication induced threat to mangroves. Saline Syst 6:8
Nelson DW, Somers LE (1975) Organic carbon. Academic, London
Black CA (1965) Methods of soil analysis. American Society of Agronomy, Wisconsin, USA
Knudsen M (1901) Hydrographical tables. G.E.C, Gad Copenhagen
Knap A, Michaels A, Close A, Ducklow H, Dickson A (eds.) (1996) Protocols for the Joint global ocean flux study (jgofs) core measurements
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4:9
Clark M (1998) Redox stratification and heavy metal partitioning in Avicennia-dominated mangrove sediments: a geochemical model. Chem Geol 149:147–171
Proctor LM (1997) Nitrogen-fixing, photosynthetic, anaerobic bacteria associated with pelagic copepods. Aquat Microb Ecol 12:105–113
Sauer JD, Shannon JG, Howe D, Hayes SF, Swanson MS, Heinzen RA (2005) Specificity of Legionella pneumophila and Coxiella burnetii vacuoles and versatility of Legionella pneumophila revealed by coinfection. Infect Immun 73:4494–4504
Dang H, Li T, Chen M, Huang G (2007) Cross-ocean distribution of Rhodobacterales bacteria as primary surface colonizers in temperate coastal marine waters. Appl Environ Microbiol 74:52–60
Liang Q, Lloyd-Jones G (2010) Sphingobium scionense sp. nov., an aromatic hydrocarbon-degrading bacterium isolated from contaminated sawmill soil. Int J Syst Evol Microbiol 60:413–416
Barua S, Tripathi S, Chakraborty A, Ghosh S, Chakrabarti K (2012) Characterization and crop production efficiency of diazotrophic bacterial isolates from coastal saline soils. Microbiol Res 167:95–102
Wu J, Guan T, Jiang H, Zhi X, Tang S, Dong H, Zhang L, Li W (2009) Diversity of Actinobacterial community in saline sediments from Yunnan and Xinjiang, China. Extremophiles 13:623–632
Arumugam M, Mitra A, Pramanik A, Saha M, Gachhui R, Mukherjee J (2010) Streptomyces Sundarbansensis sp. nov., an actinomycete that produces 2-allyloxyphenol. Int J Syst Evol Microbiol 61:2664–2669
Dunbar J, Takala S, Barns SM, Davis JA, Kuske CR (1999) Levels of bacterial community diversity in four arid soils compared by cultivation and 16S rRNA gene cloning. Appl Environ Microbiol 65:1662–1669
Lauber CL, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75:5111–5120
Schottner S, Pfitzner B, Grunke S, Rasheed M, Wild C, Ramette A (2011) Drivers of bacterial diversity dynamics in permeable carbonate and silicate coral reef sands from the Red Sea. Environ Microbiol 13:1815–1826
Bolhuis H, Stal LJ (2011) Analysis of bacterial and archaeal diversity in coastal microbial mats using massive parallel 16S rRNA gene tag sequencing. ISME J 5:1701–1712
Derakshani M, Lukow T, Liesack W (2001) Novel bacterial lineages at the (sub)division level as detected by signature nucleotide-targeted recovery of 16S rRNA genes from bulk soil and rice roots of flooded rice microcosms. Appl Environ Microbiol 67:623–631
Harris JK, Kelley ST, Pace NR (2004) New perspective on uncultured bacterial phylogenetic division OP11. Appl Environ Microbiol 70:845–849
Youssef NH, Blainey PC, Quake SR, Elshahed MS (2011) Partial genome assembly for a candidate division OP11 single cell from an anoxic spring (Zodletone Spring, Oklahoma). Appl Environ Microbiol 77:7804–7814
Acknowledgments
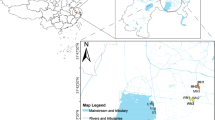
The authors would like to acknowledge the instrument facility provided by UGC-CAS, DST–FIST, DBT-IPLS, World Bank, ICZM project in the Department of Biochemistry, University of Calcutta, India. We acknowledge Mr. Tapas Paul, World Bank, for his continuous support and enthusiasm regarding our study in Sundarbans. The expenditure of this work and the research fellowships of P.B., S.N., A.B..., D.R., A.C., and R.P. were supported by the ICZM project, World Bank. A.G. was supported by Ramanujan Fellowship from the Department of Science and Technology, India (SR/S2/RJN-106/2012). The authors thank Ms. Anwesha Haldar, Department of Geography, University of Calcutta, for making a working map of the study sites in Sundarbans.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 13 kb)
ESM 2
(XLSX 11 kb)
Fig. S1
Mean nutrient concentrations [Ammonia (NH4 +), Silicate (SiO4 4−), Phosphate (PO4 3−), Nitrate (NO3 −), Nitrite (NO2 −), and sulfate (SO4 2−)] at the sampling stations during December 2011 (A) and July 2012 (B). (GIF 22 kb)
Fig. S2
Principal Component Analysis of samples based on environmental parameters using PAST. Percentages of variation explained are indicated in each axis in parentheses. (GIF 6 kb)
Rights and permissions
About this article
Cite this article
Basak, P., Majumder, N.S., Nag, S. et al. Spatiotemporal Analysis of Bacterial Diversity in Sediments of Sundarbans Using Parallel 16S rRNA Gene Tag Sequencing. Microb Ecol 69, 500–511 (2015). https://doi.org/10.1007/s00248-014-0498-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-014-0498-y