Abstract
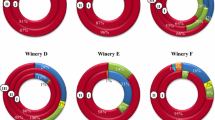
Diversity of lactic acid bacteria (LAB) species has been analyzed for three consecutive years (2006, 2007, and 2008) during alcoholic and malolactic fermentations of Tempranillo wine in a winery at La Rioja. The results showed differences in malolactic fermentation duration, and in both diversity of LAB species and diversity of Oenococcus oeni genotypes. O. oeni was shown to be the predominant species (73% of total isolates). Monitoring the different strains of O. oeni using pulsed-field gel electrophoresis of chromosomal DNA digested with SfiI and ApaI allowed detection of a total of 37 distinct genotypes, most of them comprised at least two isolates. Six appeared in more than one vintage, one of them being present in the three studied years. Moreover, four genotypes were indistinct of the strains isolated from the air of this same winery in 2007 vintage. The frequency of participation of each genotype varied from year to year, thus dominant genotypes at one year were minority or not present at another year. This suggests that distinct indigenous O. oeni strains are better adapted to the different winery conditions every year. Predominant genotypes that appeared in more than one vintage and lead to quality wines with low histamine contents could be considered as interesting for selecting of new malolactic starter cultures.



Similar content being viewed by others
References
Carrascosa AV, Muñoz R, González R (2005) Microbiología del vino. AMV Ediciones, Madrid
Lonvaud-Funel A (1999) Lactic acid bacteria in the quality improvement and depreciation of wine. Antonie Van Leeuwenhoek, Int J Gen Mol Microbiol 76:317–331
Fugelsang KC, Edwards CG (2007) Microbial ecology during vinification. In: Fugelsang KC, Edwards CG (eds) Wine microbiology. Practical applications and procedures, second edition. Springer, New York, pp 88–91
Guerrini S, Bastianini A, Blaiotta G, Granchi L, Moschetti G, Coppola S, Romano P, Vincenzini M (2003) Phenotypic and genotypic characterization of Oenococcus oeni strains isolated from Italian wines. Int J Food Microbiol 83:1–14
Larisika M, Claus H, König H (2008) Pulsed-field gel electrophoresis for the discrimination of Oenococcus oeni isolates from different wine-growing regions in Germany. Int J Food Microbiol 123:171–176
Solieri L, Genova F, De Paola M, Giudici P (2010) Characterization and technological properties of Oenococcus oeni strains from wine spontaneous malolactic fermentations: a framework for selection of new starter cultures. J Appl Microbiol 108:285–298
Reguant C, Carreté R, Constantí M, Bordons A (2005) Population dynamics of Oenococcus oeni strains in a new winery and the effect of SO2 and yeast strain. FEMS Microbiol Lett 246:111–117
Renouf V, Delaherche A, Claisse O, Lonvaud-Funel A (2008) Correlation between indigenous Oenococcus oeni strain resistance and the presence of genetic markers. J Ind Microbiol Biotechnol 35:27–33
Coucheney F, Desroche N, Bou M, Tourdot-Maréchal R, Dulau L, Guzzo J (2005) A new approach for selection of Oenococcus oeni strains in order to produce malolactic starters. Int J Food Microbiol 105:463–470
Izquierdo PM, Garcia E, Martinez J, Chacon JL (2004) Selection of lactic bacteria to induce malolactic fermentation in red wine of cv Cencibel. Vitis 43:149–153
Ruiz P, Izquierdo PM, Sesena S, Palop ML (2010) Selection of autochthonous Oenococcus oeni strains according to their oenological properties and vinification results. Int J Food Microbiol 137:230–235
Izquierdo PM, Ruiz P, Seseña S, Palop ML (2009) Ecological study of lactic acid microbiota isolated from Tempranillo wines of Castilla–La Mancha. J Biosc Bioeng 108:220–224
López I, Tenorio C, Zarazaga M, Dizy M, Torres C, Ruiz-Larrea F (2007) Evidence of mixed wild populations of Oenococcus oeni strains during wine spontaneous malolactic fermentations. Eur Food Res Technol 226:215–223
Ruiz P, Izquierdo PM, Sesena S, Palop ML (2010) Analysis of lactic acid bacteria populations during spontaneous malolactic fermentation of Tempranillo wines at five wineries during two consecutive vintages. Food Control 21:70–75
European Community (1990) Community methods for the analysis of wine. Commission Regulation (ECC) no. 26/76/90 of 17 September 1990. Off J Eur Commun 33(L272):1–191
López R (2009) Control de la fermentación maloláctica en vinos tintos de Rioja. Influencia en su calidad higiénica, físico-química y sensorial. Thesis, University of La Rioja, Spain
Holt JG, Krieg NR, Sneath PHA, Staley JT, Williams ST (1994) In: Hensyl WR (ed) Bergey's manual of determinative bacteriology, 9th edn. Baltimore, Williams and Wilkins
Beneduce L, Spano G, Vernile A, Tarantino D, Massa S (2004) Molecular characterization of lactic acid populations associated with wine spoilage. J Basic Microbiol 44:10–16
Zapparoli G, Torriani S, Pesente P, Dellaglio F (1998) Design and evaluation of malolactic enzyme gene targeted primers for rapid identification and detection of Oenococcus oeni in wine. Lett Appl Microbiol 27:243–246
López I, Ruiz-Larrea F, Cocolin L, Orr E, Phister T, Marshall M, VanderGheynst J, Mills DA (2003) Design and evaluation of PCR primers for analysis of bacterial populations in wine by denaturing gradient gel electrophoresis. Appl Environ Microbiol 69:6801–6807
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Birren B, Lai E (1993) Preparation of DNA for pulsed field analysis. In: Pulsed field gel electrophoresis. A practical guide. Academic Press Inc, San Diego
Ruiz P, Izquierdo PM, Sesena S, Palop ML (2008) Intraspecific genetic diversity of lactic acid bacteria from malolactic fermentation of Cencibel wines as derived from combined analysis of RAPD-PCR and PFGE patterns. Food Microbiol 25:942–948
Boletin de maduración del Consejo Regulador de la D.O.Ca. Rioja. http://www.riojawine.com/es/pdfs/2007_boletin7.pdf Accessed 20 June 2011
López I, López R, Santamaría P, Torres C, Ruíz-Larrea F (2008) Performance of malolactic fermentation by inoculation of selected Lactobacillus plantarum and Oenococcus oeni strains isolated from Rioja red wines. Vitis 47:123–129
Du Plessis HW, Dicks LMT, Pretorius IS, Lambrechts MG, Du Toit M (2004) Identification of lactic acid bacteria isolated from South African brandy base wines. Int J Food Microbiol 91:19–29
Garijo P, Lopez R, Santamaria P, Ocon E, Olarte C, Sanz S, Gutierrez AR (2009) Presence of lactic bacteria in the air of a winery during the vinification period. Int J Food Microbiol 136:142–146
Gutiérrez AR, Santamaría P, Epifanio S, Garijo P, López R (1999) Ecology of spontaneous fermentation in one winery during 5 consecutive years. Lett Appl Microbiol 29:411–415
Renouf V, Vayssieres LC, Claisse O, Lonvaud-Funel A (2009) Genetic and phenotypic evidence for two groups of Oenococcus oeni strains and their prevalence during winemaking. Appl Microbiol Biotechnol 83:85–97
Acknowledgments
This work was supported by funding and predoctoral grant (B.O.R., 6 March, 2009) of the Government of La Rioja, the I.N.I.A. project RTA2007-00104-00-00, and FEDER of the European Community and was made possible by the collaborating winery.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
González-Arenzana, L., López, R., Santamaría, P. et al. Dynamics of Indigenous Lactic Acid Bacteria Populations in Wine Fermentations from La Rioja (Spain) During Three Vintages. Microb Ecol 63, 12–19 (2012). https://doi.org/10.1007/s00248-011-9911-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-011-9911-y