Abstract
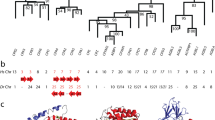
Fourteen different pepsinogen-A cDNAs and one pepsinogen-C cDNA have been cloned from gastric mucosa of the orangutan, Pongo pygmaeus. Encoded pepsinogens A were classified into two groups, i.e., types A1 and A2, which are different in acidic character. The occurrence of 9 and 5 alleles of A1 and A2 genes (at least 5 and 3 loci), respectively was anticipated. Respective orthologous genes are present in the chimpanzee genome although their copy numbers are much smaller than those of the orangutan genes. Only A1 genes are present in the human probably due to the loss of the A2 gene. Molecular phylogenetic analyses showed that A1 and A2 genes diverged before the speciation of great hominoids. Further reduplications of respective genes occurred several times in the orangutan lineage, with much higher frequencies than those occurred in the chimpanzee and human lineages. The rates of non-synonymous substitutions were higher than those of synonymous ones in the lineage of A2 genes, implying the contribution of the positive selection on the encoded enzymes. Several sites of pepsin moieties were indeed found to be under positive selection, and most of them locate on the surface of the molecule, being involved in the conformational flexibility. Deduced from the known genomic structures of pepsinogen-A genes of primates and other mammals, the duplication/loss were frequent during their evolution. The extreme multiplication in the orangutan might be advantageous for digestion of herbaceous foods due to the increase in the level of enzymes in stomach and the diversification of enzyme specificity.






Similar content being viewed by others
References
Athauda SB, Tanji M, Kageyama T, Takahashi K (1989) A comparative study on the NH2-terminal amino acid sequences and some other properties of six isozymic forms of human pepsinogens and pepsins. J Biochem 106:920–927
Bankowska A, Roszkowska-Jakimiec W, Worowki K (1998) Inhibitors of pepsin, trypsin and chymotrypsin in seeds of plants consumed by humans and animals. I. Evaluation of pepsin, trypsin, and chymotrypsin inhibitors activity in seeds of 26 plant species. Rocz Akad Med Bialymst 43:278–286
Borrelli L, De Stasio R, Filosa S, Parisi E, Riggio M, Scudiero R, Trinchella F (2006) Evolutionary fate of duplicate genes encoding aspartic proteinases. Nothepsin case study. Gene 368:101–109
Carginale V, Trinchella F, Capasso C, Scudiero R, Riggio M, Parisi E (2004) Adaptive evolution and functional divergence of pepsin gene family. Gene 333:81–90
Chirgwin JM, Przybyla AE, MacDonald RJ, Rutter WJ (1979) Isolation of biologically active ribonucleic acid from sources enriched in ribonuclease. Biochemistry 18:5294–5299
Christeller JT, Farley PC, Ramsay RJ, Sullivan PA, Laing WA (1998) Purification, characterization and cloning of an aspartic proteinase inhibitor from squash phloem exudate. Eur J Biochem 254:160–167
Cottrell TJ, Harris LJ, Tanaka T, Yada RY (1995) The sole lysine residue in porcine pepsin works as a key residue for catalysis and conformational flexibility. J Biol Chem 270:19974–19978
Dumas L, Kim YH, Karimpour-Fard A, Cox M, Hopkins J, Pollack JR, Sikela JM (2007) Gene copy number variation spanning 60 million years of human and primate evolution. Genome Res 17:1266–1277
Evers MP, Zelle B, Bebelman JP, van Beusechem V, Kraakman L, Hoffer MJ, Pronk JC, Mager WH, Planta RJ, Eriksson AW, Frants RR (1989) Nucleotide sequence comparison of five human pepsinogen A (PGA) genes: evolution of the PGA multigene family. Genomics 4:232–239
Feng S, Li W, Lin H (2008) Characterization and expression of the pepsinogen C gene and determination of pepsin-like enzyme activity from orange-spotted grouper (Epinephelus coioides). Comp Biochem Physiol B Biochem Mol Biol 149:275–284
Finch CE, Stanford CB (2004) Meat-adaptive genes and the evolution of slower aging in humans. Q Rev Biol 79:3–50
Foltmann B (1981) Gastric proteinases—structure, function, evolution and mechanism of action. Essays Biochem 17:52–84
Foltmann B (1992) Chymosin: a short review on foetal and neonatal gastric proteases. Scand J Clin Lab Invest 52(Suppl. 210):65–79
Fortna A, Kim Y, MacLaren E, Marshall K, Hahn G, Meltesen L, Brenton M, Hink R, Burgers S, Hernandez-Boussard T, Karimpour-Fard A, Glueck D, McGavran L, Berry R, Pollack J, Sikela JM (2004) Lineage-specific gene duplication and loss in human and great ape evolution. PLoS Biol 2:E207
Frazer KA, Chen X, Hinds DA, Pant PV, Patil N, Cox DR (2003) Genomic DNA insertions and deletions occur frequently between humans and nonhuman primates. Genome Res 13:341–346
Galdikas BMF (1988) Orangutan diet, range, and activity at Tanjung Puting, Central Borneo. Int J Primatol 9:1–35
Goodman M, Bailey WJ, Hayasaka K, Stanhope MJ, Slightom J, Czelusniak J (1994) Molecular evidence on primate phylogeny from DNA sequences. Am J Phys Anthropol 94:3–24
Goodman M, Porter CA, Czelusniak J, Page SL, Schneider H, Shoshani J, Gunnell G, Groves CP (1998) Toward a phylogenetic classification of primates based on DNA evidence complemented by fossil evidence. Mol Phylogenet Evol 9:585–598
Gubler U, Hoffman BJ (1983) A simple and very efficient method for generating cDNA libraries. Gene 25:263–269
Harding RSO (1981) An order of omnivores: nonhuman primate diets in the wild. In: Harding RSO, Teleki G (eds) Omnivorous primates. Columbia University Press, New York, pp 191–214
Hasegawa M, Kishino H, Yano T (1985) Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol 22:160–174
Kageyama T (2000) New world monkey pepsinogens A and C, and prochymosins. Purification, characterization of enzymatic properties, cDNA cloning, and molecular evolution. J Biochem 127:761–770
Kageyama T (2002) Pepsinogens, progastricsins, and prochymosins: structure, function, evolution, and development. Cell Mol Life Sci 59:288–306
Kageyama T (2004) Role of S’1 loop residues in the substrate specificities of pepsin A and chymosin. Biochemistry 43:15122–15130
Kageyama T (2006) Roles of Tyr13 and Phe219 in the unique substrate specificity of pepsin B. Biochemistry 45:14415–14426
Kageyama T, Takahashi K (1976) Pepsinogens and pepsins from gastric mucosa of Japanese Monkey. Purification and characterization. J Biochem 79:455–468
Kageyama T, Takahashi K (1977) The carbohydrate moiety of Japanese monkey pepsinogens. Its composition and site of attachment to protein. Biochem Biophys Res Commun 74:789–795
Kageyama T, Takahashi K (1984) Rabbit pepsinogens. Purification, characterization, analysis of the conversion process to pepsin and determination of the NH2-terminal amino-acid sequences. Eur J Biochem 141:261–269
Kageyama T, Tanabe K, Koiwai O (1990) Structure and development of rabbit pepsinogens. Stage-specific zymogens, nucleotide sequences of cDNAs, molecular evolution, and gene expression during development. J Biol Chem 265:17031–17038
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kishino H, Hasegawa M (1989) Evaluation of the maximum likelihood estimate of the evolutionary tree topologies from DNA sequence data, and the branching order in hominoidea. J Mol Evol 29:170–179
Miyamoto MM, Koop BF, Slightom JL, Goodman M, Tennant MR (1988) Molecular systematics of higher primates: genealogical relations and classification. Proc Natl Acad Sci USA 85:7627–7631
Narita Y, Kageyama T (2003) Diversity of ape pepsinogen genes (in Japanese with English summary). Primate Res 19:125–133
Narita Y, Oda S, Moriyama A, Takenaka O, Kageyama T (1997) Pepsinogens and pepsins from house musk shrew, Suncus murinus: purification, characterization, determination of the amino-acid sequences of the activation segments, and analysis of proteolytic specificities. J Biochem 121:1010–1017
Narita Y, Oda S, Takenaka O, Kageyama T (2000) Multiplicities and some enzymatic characteristics of ape pepsinogens and pepsins. J Med Primatol 29:402–410
Narita Y, Oda S, Takenaka O, Kageyama T (2001) Phylogenetic position of Eulipotyphla inferred from the cDNA sequences of pepsinogens A and C. Mol Phylogenet Evol 21:32–42
Narita Y, Oda S, Moriyama A, Kageyama T (2002) Primary structure, unique enzymatic properties, and molecular evolution of pepsinogen B and pepsin B. Arch Biochem Biophys 404:177–185
O’hUigin C, Satta Y, Takahata N, Klein J (2002) Contribution of homoplasy and of ancestral polymorphism to the evolution of genes in anthropoid primates. Mol Biol Evol 19:1501–1513
Ordonez GR, Hillier LW, Warren WC, Grutzner F, Lopez-Otin C, Puente XS (2008) Loss of genes implicated in gastric function during platypus evolution. Genome Biol 9:R81
Perry GH, Dominy NJ, Claw KG, Lee AS, Fiegler H, Redon R, Werner J, Villanea FA, Mountain JL, Misra R, Carter NP, Lee C, Stone AC (2007) Diet and the evolution of human amylase gene copy number variation. Nat Genet 39:1256–1260
Robinson-Rechavi M, Huchon D (2000) RRTree: relative-rate tests between groups of sequences on a phylogenetic tree. Bioinformatics 16:296–297
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sali A, Blundell TL (1993) Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol 234:779–815
Samloff IM (1971) Pepsinogens, pepsins, and pepsin inhibitors. Gastroenterology 60:586–604
Schoniger M, von Haeseler A (1994) A stochastic model for the evolution of autocorrelated DNA sequences. Mol Phylogenet Evol 3:240–247
Steiper ME, Young NM (2006) Primate molecular divergence dates. Mol Phylogenet Evol 41:384–394
Strimmer K, von Haeseler M (1996) Quartet puzzling: a quartet maximum-likelihood method for reconstructing tree topologies. Mol Biol Evol 13:964–969
Suchodolski JS, Steiner JM, Ruaux CG, Boari A, Williams DA (2002) Purification and partial characterization of canine pepsinogen A and B. Am J Vet Res 63:1585–1590
Swofford DL (1998) PAUP*. Phylogenetic analysis using parsimony and other methods. Sinauer Associates, Sunderland, MA
Tang J, Sepulveda P, Marciniszyn J Jr, Chen KC, Huang WY, Tao N, Liu D, Lanier JP (1973) Amino-acid sequence of porcine pepsin. Proc Natl Acad Sci U S A 70:3437–3439
Yang Z (1998) Likelihood ratio tests for detecting positive selection and application to primate lysozyme evolution. Mol Biol Evol 15:568–573
Yang Z (2007) PAML 4: a phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Yang Z, Nielsen R (1998) Synonymous and nonsynonymous rate variation in nuclear genes of mammals. J Mol Evol 46:409–418
Zelle B, Evers MP, Groot PC, Bebelman JP, Mager WH, Planta RJ, Pronk JC, Meuwissen SG, Hofker MH, Eriksson AW, Frants RR (1988) Genomic structure and evolution of the human pepsinogen A multigene family. Hum Genet 78:79–82
Acknowledgments
This study was supported in part by Grants-in-Aid for Scientific Research (19370102 to T.K.) and the Global Center of Excellence Program “Formation of a Strategic Base for Biodiversity and Evolutionary Research: from Genome to Ecosystem” from the Ministry of Education, Science, Sports and Culture of Japan, and by Grants for the co-operative research program of the Primate Research Institute, Kyoto University.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Narita, Y., Oda, Si., Takenaka, O. et al. Lineage-Specific Duplication and Loss of Pepsinogen Genes in Hominoid Evolution. J Mol Evol 70, 313–324 (2010). https://doi.org/10.1007/s00239-010-9320-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-010-9320-8