Abstract
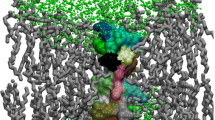
The membrane disruption and pore-forming mechanism of melittin has been widely explored by experiments and computational studies. However, the precise mechanism is still enigmatic, and further study is required to turn antimicrobial peptides into future promising drugs against microbes. In this study, unbiased microsecond (µs) time scale (total 17 µs) atomistic molecular dynamics simulation were performed on multiple melittin systems in 1,2-dimyristoyl-sn-glycero-3-phosphocholine membrane to capture the various events during the membrane disorder produced by melittin. We observed bent U-shaped conformations of melittin, penetrated deeply into the membrane in all simulations, and a special double U-shaped structure. However, no peptide transmembrane insertion, nor pore formation was seen, indicating that these processes occur on much longer timescales, and suggesting that many prior computational studies of melittin were not sufficiently unbiased.






Similar content being viewed by others
References
Allende D, Simon SA, McIntosh TJ (2005) Melittin-induced bilayer leakage depends on lipid material properties: evidence for toroidal pores. Biophys J 88:1828–1837
Andersson M, Ulmschneider JakobP, Ulmschneider MartinB, White StephenH (2013) Conformational states of melittin at a bilayer interface. Biophys J 104:L12–L14
Berendsen HJC, van der Spoel D, van Drunen R (1995) GROMACS: a message-passing parallel molecular dynamics implementation. Comput Phys Commun 91:43–56
Berneche S, Nina M, Roux B (1998) Molecular dynamics simulation of melittin in a dimyristoylphosphatidylcholine bilayer membrane. Biophys J 75:1603–1618
Brogden KA (2005) Antimicrobial peptides: pore formers or metabolic inhibitors in bacteria? Nat Rev Microbiol 3:238–250
Brown KL, Hancock RE (2006) Cationic host defense (antimicrobial) peptides. Curr Opin Immunol 18:24–30
Brown LR, Lauterwein J, Wuthrich K (1980) High-resolution 1H-NMR studies of self-aggregation of melittin in aqueous solution. Biochim Biophys Acta 622:231–244
Bussi G, Donadio D, Parrinello M (2007) Canonical sampling through velocity rescaling. J Chem Phys 126:014101
Chen X, Wang J, Boughton AP, Kristalyn CB, Chen Z (2007) Multiple orientation of melittin inside a single lipid bilayer determined by combined vibrational spectroscopic studies. J Am Chem Soc 129:1420–1427
Chromek M, Slamova Z, Bergman P, Kovacs L, Podracka L, Ehren I, Hokfelt T, Gudmundsson GH, Gallo RL, Agerberth B, Brauner A (2006) The antimicrobial peptide cathelicidin protects the urinary tract against invasive bacterial infection. Nat Med 12:636–641
Demchenko AP, Kostrzhevskaia EG (1986) Melittin: structure, properties, interaction with a membrane. Ukr Biokhim Zh 58:92–103
Hancock RE, Scott MG (2000) The role of antimicrobial peptides in animal defenses. Proc Natl Acad Sci USA 97:8856–8861
Hristova K, Dempsey CE, White SH (2001) Structure, location, and lipid perturbations of melittin at the membrane interface. Biophys J 80(2):801–811
Huang HW (2000) Action of antimicrobial peptides: two-state model. Biochemistry 39:8347–8352
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(33–8):27–28
Irudayam SJ, Berkowitz ML (2011) Influence of the arrangement and secondary structure of melittin peptides on the formation and stability of toroidal pores. Biochim Biophys Acta 1808:2258–2266
Irudayam SJ, Berkowitz ML (2012) Binding and reorientation of melittin in a POPC bilayer: computer simulations. Biochim Biophys Acta 1818:2975–2981
Iwadate M, Asakura T, Williamson MP (1998) The structure of the melittin tetramer at different temperatures—an NOE-based calculation with chemical shift refinement. Eur J Biochem 257:479–487
Langham AA, Ahmad AS, Kaznessis YN (2008) On the nature of antimicrobial activity: a model for protegrin-1 pores. J Am Chem Soc 130:4338–4346
Lee MT, Sun TL, Hung WC, Huang HW (2013) Process of inducing pores in membranes by melittin. Proc Natl Acad Sci USA 110:14243–14248
Leontiadou H, Mark AE, Marrink SJ (2006) Antimicrobial peptides in action. J Am Chem Soc 128:12156–12161
Leveritt JM III (2015) The structure of a melittin-stabilized toroidal pore. Biophys J 108(2):249a. doi:10.1016/j.bpj.2014.11.1380
Lin JH, Baumgaertner A (2000) Stability of a melittin pore in a lipid bilayer: a molecular dynamics study. Biophys J 78:1714–1724
Lohner K, Blondelle SE (2005) Molecular mechanisms of membrane perturbation by antimicrobial peptides and the use of biophysical studies in the design of novel peptide antibiotics. Comb Chem High Throughput Screen 8:241–256
Manna M, Mukhopadhyay C (2009) Cause and effect of melittin-induced pore formation: a computational approach. Langmuir 25:12235–12242
Matsuzaki K, Yoneyama S, Miyajima K (1997) Pore formation and translocation of melittin. Biophys J 73:831–838
Park Y, Hahm KS (2005) Antimicrobial peptides (AMPs): peptide structure and mode of action. J Biochem Mol Biol 38:507–516
Pronk S, Pall S, Schulz R, Larsson P, Bjelkmar P, Apostolov R, Shirts MR, Smith JC, Kasson PM, van der Spoel D, Hess B, Lindahl E (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29:845–854
Rex S (1996) Pore formation induced by the peptide melittin in different lipid vesicle membranes. Biophys Chem 58(1–2):75–85
Sanchez-Martinez S, Huarte N, Maeso R, Madan V, Carrasco L, Nieva JL (2008) Functional and structural characterization of 2B viroporin membranolytic domains. Biochemistry 47:10731–10739
Santo KP, Irudayam SJ, Berkowitz ML (2013) Melittin creates transient pores in a lipid bilayer: results from computer simulations. J Phys Chem B 117:5031–5042
Schwarz G, Zong RT, Popescu T (1992) Kinetics of melittin induced pore formation in the membrane of lipid vesicles. Biochim Biophys Acta 1110(1):97–104
Sengupta D, Leontiadou H, Mark AE, Marrink SJ (2008) Toroidal pores formed by antimicrobial peptides show significant disorder. Biochim Biophys Acta 1778:2308–2317
Sharon M, Oren Z, Shai Y, Anglister J (1999) 2D-NMR and ATR-FTIR study of the structure of a cell-selective diastereomer of melittin and its orientation in phospholipids. Biochemistry 38:15305–15316
Son DJ, Lee JW, Lee YH, Song HS, Lee CK, Hong JT (2007) Therapeutic application of anti-arthritis, pain-releasing, and anti-cancer effects of bee venom and its constituent compounds. Pharmacol Ther 115:246–270
van den Bogaart G, Guzman JV, Mika JT, Poolman B (2008) On the mechanism of pore formation by melittin. J Biol Chem 283:33854–33857
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718
Vogel H, Jahnig F (1986) The structure of melittin in membranes. Biophys J 50:573–582
Wang Y, Zhao T, Wei D, Strandberg E, Ulrich AS, Ulmschneider JP (2014) How reliable are molecular dynamics simulations of membrane active antimicrobial peptides? Biochim Biophys Acta Biomembr 1838:2280–2288
Yang L, Harroun TA, Weiss TM, Ding L, Huang HW (2001) Barrel-stave model or toroidal model? A case study on melittin pores. Biophys J 81:1475–1485
Yeaman MR, Yount NY (2003) Mechanisms of antimicrobial peptide action and resistance. Pharmacol Rev 55:27–55
Zhu L, Kemple MD, Yuan P, Prendergast FG (1995) N-terminus and lysine side chain pKa values of melittin in aqueous solutions and micellar dispersions measured by 15 N NMR. Biochemistry 34:13196–13202
Acknowledgments
This research was supported by a grant from the National Natural Science Foundation of China (91230105) and a 1000 Plan’s Program for Young Talents (13Z127060001) to J.P.U.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Upadhyay, S.K., Wang, Y., Zhao, T. et al. Insights from Micro-second Atomistic Simulations of Melittin in Thin Lipid Bilayers. J Membrane Biol 248, 497–503 (2015). https://doi.org/10.1007/s00232-015-9807-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-015-9807-8