Abstract
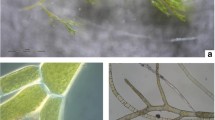
Although several studies have investigated bacterial–algal interactions, the bacterial component has often not been identified, and the ecological role of bacterial–algal associations is still unclear. In the present study two different approaches (molecular and culture) have been used to characterize the bacterial community associated with the invasive alga Caulerpa cylindracea (Sonder) over time, in a coastal area of the Mediterranean basin. C. cylindracea is an invasive macroalga in the Mediterranean Sea, able to colonize several types of substrates. Traditional culture-based and PCR-SSCP methods have been used to analyze the bacterial community. Molecular traces of Gammaproteobacteria belonging to the genera Shewanella and Vibrio have been found by both approaches on the surface of C. cylindracea consistently in time, along with those of an unknown species belonging to the Rhodobacteraceae family. Other taxa belonging to Bacillus, Pseudoalteromonas, Tropicibacter, Photobacterium, Exiguobacterium, Kocuria, Ruegeria, and Marinobacter genera have been discovered by culture-based approach. PCR-SSCP method has shown traces of an unknown species of the Bacteroidetes phylum and the Granulosicoccus genus. Our results suggest that C. cylindracea hosts a bacterial assemblage scarcely variable with time. Further studies are needed to clarify the nature of this alga–bacteria association and the potential role in the spreading of this alga using a holistic view considering the seaweed with associated bacteria as an essentially unique meta-organism.





Similar content being viewed by others
References
Abriouel H, Franz CM, Omar NB, Gálvez A (2011) Diversity and applications of Bacillus bacteriocins. FEMS Microbiol Rev 35(1):201–232. doi:10.1111/j.1574-6976.2010.00244.x
Aires T, Serrão EA, Kendrick G, Duarte CM, Arnaud-Haond S (2013) Invasion is a community affair: Clandestine followers in the bacterial community associated to green algae, Caulerpa racemosa. Track the invasion source. PLoS One 8(7):e68429. doi:10.1371/journal.pone.0068429
Ashen JB, Goff LJ (1996) Molecular identification of a bacterium associated with gall formation in the marine red alga Prionitis lanceolata. J Phycol 32:286–297. doi:10.1111/j.0022-3646.1996.00286.x
Ast JC, Cleenwerck I, Engelbeen K, Urbanczyk H, Thompson FL, De Vos P, Dunlap PV (2007) Photobacterium kishitanii sp. nov., a luminous marine bacterium symbiotic with deep-sea fishes. Int J Syst Evol Microbiol 57:2073–2078. doi:10.1099/ijs.0.65153-0
Baumann P, Baumann L, Bang SS, Woolkalis MJ (1980) Reevaluation of the taxonomy of Vibrio, Beneckea, and Photobacterium: abolition of the genus Beneckea. Curr Microbiol 4(3):127–132. doi:10.1007/BF02602814
Belton GS, Prud’homme van Reine WF, Huisman JM, Draisma SGA, Gurgel CFD (2014) Resolving phenotypic plasticity and species designation in the morphology challenging Caulerpa racemosa-peltata complex (Caulerpaceae, Chlorophyta). J Phycol 50(1):32–54. doi:10.1111/jpy.12132
Bengtsson MM, Sjøtun K, Lanzén A, Øvreås L (2012) Bacterial diversity in relation to secondary production and succession on surfaces of the kelp Laminaria hyperborea. ISME J 6(12):2188–2198. doi:10.1038/ismej.2012.67
Bernbom N, Ng YY, Olsen SM, Gram L (2013) Pseudoalteromonas spp. serve as initial bacterial attractants in mesocosms of coastal waters but have subsequent antifouling capacity in mesocosms and when embedded in paint. Appl Environ Microbiol 79(22):6885–6893. doi:10.1128/AEM.01987-13
Burke C, Steinberg P, Rusch D, Kjelleberg S, Thomas T (2011a) Bacterial community assembly based on functional genes rather than species. Proc Natl Acad Sci USA 108:14288–14293. doi:10.1073/pnas.1101591108
Burke C, Thomas T, Lewis M, Steinberg P, Kjelleberg S (2011b) Composition, uniqueness and variability of the epiphytic bacterial community of the green alga Ulva australis. ISME J 5:590–600. doi:10.1038/ismej.2010.164
Cano-Gómez A, Goulden EF, Owens L, Høj L (2010) Vibrio owensii sp. nov., isolated from cultured crustaceans in Australia. FEMS Microbiol Lett 302(2):175–181. doi:10.1111/j.1574-6968.2009.01850.x
Cebrian E, Linares C, Marchal C, Garrabou J (2012) Exploring the effects of invasive algae on the persistence of gorgonian populations. Biol Invasions 14:2647–2656. doi:10.1007/s10530-012-0261-6
Chimetto LA, Cleenwerck I, Alves N, Silva BS, Brocchi M, Willems A, De Vos P, Thompson FL (2011) Vibrio communis sp. nov., isolated from the marine animals Mussismilia hispida, Phyllogorgia dilatata, Palythoa caribaeorum, Palythoa variabilis and Litopenaeus vannamei. Int J Syst Evol Microbiol 61(2):362–368. doi:10.1099/ijs.0.019729-0
Dheilly A, Soum-Soutéra E, Klein GL, Bazire A, Compére C, Haras D, Dufour A (2010) Antibiofilm activity of the marine bacterium Pseudoalteromonas sp. strain 3J6. Appl Environ Microbiol 76:3452–3461. doi:10.1128/AEM.02632-09
Dobretsov SV, Dahms HU, Harder T, Qian PY (2006) Allelochemical defense against epibiosis in the macroalga Caulerpa racemosa var. turbinata. Mar Ecol Prog Ser 318:165–175. doi:10.3354/meps318165
Editorial (2013) The cultural revolution. Nat Rev Microbiol 11:1. doi:10.1038/nrmicro2948
Egan S, Thomas T, Holmstrom C, Kjelleberg S (2000) Phylogenetic relationship and antifouling activity of bacterial epiphytes from the marine alga Ulva lactuca. Environ Microbiol 2:343–347. doi:10.1046/j.1462-2920.2000.00107.x
Felline S, Caricato R, Cutignano A, Gorbi S, Lionetto MG, Mollo E, Regoli F, Terlizzi A (2012) Subtle effects of biological invasions: cellular and physiological responses of fish eating the exotic pest Caulerpa racemosa. PLoS One 7:e38763. doi:10.1371/journal.pone.0038763
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376. doi:10.1007/BF01734359
Gauthier G, Gauthier M, Christen R (1995) Phylogenetic analysis of the genera Alteromonas, Shewanella, and Moritella using genes coding for small-subunit rRNA sequences and division of the genus Alteromonas into two genera, Alteromonas (emended) and Pseudoalteromonas gen. nov., and proposal of twelve new species combinations. Int J Syst Bacteriol 45:755–761. doi:10.1099/00207713-45-4-755
Goecke F, Labes A, Wiese J, Imhoff JF (2010) Chemical interactions between marine macroalgae and bacteria. Mar Ecol Prog Ser 409:267–299. doi:10.3354/meps08607
Goecke F, Labes A, Wiese J, Imhoff JF (2013a) Phylogenetic analysis and antibiotic activity of bacteria isolated from the surface of two co-occurring macroalgae from the Baltic Sea. Eur J Phycol 48(1):47–60. doi:10.1080/09670262.2013.767944
Goecke F, Thiel V, Wiese J, Labes A, Imhoff JF (2013b) Algae as an important environment for bacteria: phylogenetic relationships among new bacterial species isolated from algae. Phycologia 52:14–24. doi:10.2216/12-24.1
Gomez-Gil B, Thompson FL, Thompson CC, Swings J (2003) Vibrio rotiferianus sp. nov., isolated from cultures of the rotifer Brachionus plicatilis. Int J Syst Evol Microbiol 53(1):239–243. doi:10.1099/ijs.0.02430-0
Goulden EF, Hall MR, Bourne DG, Pereg LL, Høj L (2012) Pathogenicity and infection cycle of Vibrio owensii in larviculture of the ornate spiny lobster (Panulirus ornatus). Appl Environ Microbiol 78(8):2841–2849. doi:10.1128/AEM.07274-11
Gouy M, Guindon S, Gascuel O (2010) SeaView Version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27(2):221–224. doi:10.1093/molbev/msp259
Guerrero-Ferreira R, Gorman C, Chavez AA, Willie S, Nishiguchi MK (2013) Characterization of the bacterial diversity in Indo-West Pacific loliginid and sepiolid squid light organs. Microb Ecol 65(1):214–226. doi:10.1007/s00248-012-0099-6
Hollants J, Leroux O, Leliaert F, Decleyre H, De Clerck O, Willems A (2011) Who is in there? Exploration of endophytic bacteria within the siphonous green seaweed Bryopsis (Bryopsidales, Chlorophyta). PLoS One 6(10):e26458. doi:10.1371/journal.pone.0026458
Holmström C, Egan S, Franks A, McCloy S, Hjelleberg S (2002) Antifouling activities expressed by marine surface associated Pseudoalteromonas species. FEMS Microbiol Ecol 41:47–58. doi:10.1111/j.1574-6941.2002.tb00965.x
Ivanova EP, Nedashkovskaya OI, Zhukova NV, Nicolau DV, Christen R, Mikhailov VV (2003) Shewanella waksmanii sp. nov., isolated from a sipuncula (Phascolosoma japonicum). Int J Syst Evol Microbiol 53(5):1471–1477. doi:10.1099/ijs.0.02630-0
Kim IG, Lee MH, Jung SY, Song JJ, Oh TK, Yoon JH (2005) Exiguobacterium aestuarii sp. nov. and Exiguobacterium marinum sp. nov., isolated from a tidal flat of the Yellow Sea in Korea. Int J Syst Evol Microbiol 55(2):885–889. doi:10.1099/ijs.0.63308-0
Kim YO, Kim KK, Park S, Kang SJ, Lee JH, Lee SJ, Oh TK, Yoon JH (2010) Photobacterium gaetbulicola sp. nov., a lipolytic bacterium isolated from a tidal flat sediment. Int J Syst Evol Microbiol 60(11):2587–2591. doi:10.1099/ijs.0.016923-0
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721. doi:10.1099/ijs.0.038075-0
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. doi:10.1007/BF01731581
Klein J, Verlaque M (2008) The Caulerpa racemosa invasion: a critical review. Mar Pollut Bull 56:205–225. doi:10.1016/j.marpolbul.2007.09.043
Kurilenko VV, Christen R, Zhukova NV, Kalinovskaya NI, Mikhailov VV, Crawford RJ, Ivanova EP (2010) Granulosicoccus coccoides sp. nov., isolated from leaves of seagrass (Zostera marina). Int J Syst Evol Microbiol 60(4):972–976. doi:10.1099/ijs.0.013516-0
Lachnit T, Meske D, Wahl M, Harder T, Schmitz R (2011) Epibacterial community patterns on marine macroalgae are host-specific but temporally variable. Environ Microbiol 13:655–665. doi:10.1111/j.1462-2920.2010.02371.x
Lane DJ, Pace B, Olsen GJ, Stahl DA, Sogin ML, Pace NR (1985) Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci USA 82:6955–6959. doi:10.1073/pnas.82.20.6955
Lee K, Lee HK, Choi T-H, Kim K-MT, Cho J-C (2007) Granulosicoccaceae fam. nov., to include Granulosicoccus antarcticus gen. nov., sp. nov., a non-phototrophic, obligately aerobic chemoheterotroph in the order Chromatiales, isolated from antarctic Seawater. J Microbiol Biotechnol 17:1483–1490
Lee OO, Lai PY, Wu HX, Zhou XJ, Miao L, Wang H, Qian PY (2012) Marinobacter xestospongiae sp. nov., isolated from the marine sponge Xestospongia testudinaria collected from the Red Sea. Int J Syst Evol Microbiol 62(8):1980–1985. doi:10.1099/ijs.0.028811-0
Lenk S, Moraru C, Hahnke S, Arnds J, Richter M, Kube M, Reinhardt R, Brinkhoff Harder TJ, Amann R, Mußmann M (2012) Roseobacter clade bacteria are abundant in coastal sediments and encode a novel combination of sulfur oxidation genes. ISME J 6(12):2178–2187. doi:10.1038/ismej.2012.66
Longford SR, Tujula NA, Crocetti GR, Holmes AJ, Holmström C, Kjelleberg S, Steinberg PD, Taylor MW (2007) Comparisons of diversity of bacterial communities associated with three sessile marine eukaryotes. Aquat Microb Ecol 48:217–229. doi:10.3354/ame048217
Lucena T, Pujalte MJ, Ruvira MA, Garay E, Macián MC, Arahal DR (2012) Tropicibacter multivorans sp. nov., an aerobic alphaproteobacterium isolated from surface seawater. Int J Syst Evol Microbiol 62(4):844–848. doi:10.1099/ijs.0.030973-0
MacDonell MT, Colwell RR (1985) Phylogeny of the Vibrionaceae, and recommendation for two new genera, Listonella and Shewanella. Syst Appl Microbiol 6:171–182
Martin M, Portetelle D, Michel G, Vandenbol M (2014) Microorganisms living on macroalgae: diversity, interactions, and biotechnological applications. Appl Microbiol Biotechnol 98(7):2917–2935. doi:10.1007/s00253-014-5557-2
Meusnier I, Olsen JL, Stam WT, Destombe C, Valero M (2001) Phylogenetic analyses of Caulerpa taxifolia (Chlorophyta) and of its associated bacterial microflora provide clues to the origin of the Mediterranean introduction. Mol Ecol 10(4):931–946. doi:10.1046/j.1365-294X.2001.01245.x
Myers CR, Nealson KH (1988) Bacterial manganese reduction and growth with manganese oxide as the sole electron acceptor. Science 240:1319–1321. doi:10.1126/science.240.4857.1319
Nocker A, Burr M, Camper AK (2007) Genotypic microbial community profiling: a critical technical review. Microb Ecol 54(2):276–289. doi:10.1007/s00248-006-9199-5
Park S, Yoon JH (2012) Ruegeria arenilitoris sp. nov., isolated from the seashore sand around a seaweed farm. Antonie Van Leeuwenhoek 102(4):581–589. doi:10.1007/s10482-012-9753-8
Petrovskis EA, Vogel TM, Adriaens P (1994) Effects of electron acceptors and donors on transformation of tetrachloromethane by Shewanella putrefaciens MR-1. FEMS Microbiol Lett 121:357–364. doi:10.1111/j.1574-6968.1994.tb07126.x
Piazzi L, Ceccherelli G, Cinelli F (2001) Threat to macroalgal diversity: effects of the introduced green alga Caulerpa racemosa in the Mediterranean. Mar Ecol Prog Ser 210:161–165. doi:10.3354/meps210149
Porter KG, Feig YS (1980) The use of DAPI for identifying and counting aquatic microflora. Limnol Oceanogr 25:943–948
Prado S, Romalde JL, Montes J, Barja JL (2005) Pathogenic bacteria isolated from disease outbreaks in shellfish hatcheries: first description of Vibrio neptunius as an oyster pathogen. Dis Aquat Organ 67:209–215. doi:10.3354/dao067209
Preston GM, Haubold B, Rainey PR (1998) Bacterial genomics and adaptation to life on plants: implications for the evolution of pathogenicity and symbiosis. Curr Opin Microbiol 1:589–597. doi:10.1016/S1369-5274(98)80094-5
Rivas R, Garcıa-Fraile P, Mateos PF, Martınez-Molina E, Velazquez E (2006) Photobacterium halotolerans sp. nov., isolated from Lake Martel in Spain. Int J Syst Evol Microbiol 56:1067–1071. doi:10.1099/ijs.0.64099-0
Ruitton S, Verlaque M, Boudouresque CF (2005) Seasonal changes of the introduced Caulerpa racemosa var. cylindracea (Caulerpales, Chlorophyta) at the northwest limit of its Mediterranean distribution. Aquat Bot 82:55–70. doi:10.1016/j.aquabot.2005.02.008
Russell NJ, Nichols DS (1999) Polyunsaturated fatty acids in marine bacteria: a dogma rewritten. Microbiology 145:767–779
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Salemi M, Vandamme AM (2003) The phylogenetic handbook: a practical approach to DNA and protein phylogeny. Cambridge University Press, Cambridge
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbour Laboratory Press, New York
Sauvage T, Payri C, Draisma SGA, van Reine WFPH, Verbruggen H, Belton GS, Gurgel CFD, Gabriel D, Sherwood AR, Fredericq S (2013) Molecular diversity of the Caulerpa racemosa–Caulerpa peltata complex (Caulerpaceae, Bryopsidales) in New Caledonia, with new Australasian records for C. racemosa var. cylindracea. Phycologia 52(1):6–13. doi:10.2216/11-116.1
Schumann P, Spröer C, Burghardt J, Kovacs G, Stackebrandt E (1999) Reclassification of the species Kocuria erythromyxa (Brooks and Murray 1981) as Kocuria rosea (Flügge 1886). Int J Syst Bacteriol 49(2):393–396
Schumann P, Tindall BJ, Mendrock U, Kramer I, Stackebrandt E (2000) Pelczaria aurantia ATCC 49321T (=DSM 12801T) is a strain of Kocuria rosea (Flügge 1886) Stackebrandt et al. 1995. Int J Syst Evol Microbiol 50(4):1421–1424
Semple KM, Westlake DWS (1987) Characterization of iron reducing Alteromonas putrefaciens strains from oil field fluids. Can J Microbiol 35:925–931
Seo HJ, Bae SS, Lee JH, Kim SJ (2005) Photobacterium frigidiphilum sp. nov., a psychrophilic, lipolytic bacterium isolated from deep-sea sediments of Edison Seamount. Int J Syst Evol Microbiol 55:1661–1666. doi:10.1099/ijs.0.63338-0
Shade A, Hogan CS, Klimowicz AK, Linske M, McManus PS, Handelsman J (2012) Culturing captures members of the soil rare biosphere. Environ Microbiol 14(9):2247–2252. doi:10.1111/j.1462-2920.2012.02817.x
Singh RP, Reddy CRK (2014) Seaweed–microbial interactions: key functions of seaweed-associated bacteria. FEMS Microbiol Ecol 88(2):213–230. doi:10.1111/1574-6941.12297
Slepecky RA, Hemphill HE (2006) The genus Bacillus: nonmedical. In: Dworkin M, Falkow S (eds) The prokaryotes: bacteria: firmicutes, cyanobacteria, vol 4. Springer Science & Business Media, New York, pp 530–562. doi:10.1007/0-387-30744-3_16
Sober E (1983) Parsimony in systematics: philosophical issues. Annu Rev Ecol Evol Syst 14:335–357
Stabili L, Gravili C, Tredici SM, Piraino S, Talà A, Boero F, Alifano P (2008) Epibiotic Vibrio luminous bacteria isolated from some Hydrozoa and Bryozoa species. Microb Ecol 56(4):625–636
Stabili L, Rizzo L, Alifano P, Fraschetti S (2014) Functional diversity of the bacterial community associated to the invasive alien seaweed Caulerpa racemosa. Biol Mar Mediterr 21(1):127–128
Staufenberger T, Thiel V, Wiese J, Imhoff JF (2008) Phylogenetic analysis of bacteria associated with Laminaria saccharina. FEMS Microbiol Ecol 64:65–77. doi:10.1111/j.1574-6941.2008.00445.x
Talà A, Lenucci M, Gaballo A, Durante M, Tredici SM, Debowles DA, Pizzolante G, Marcuccio C, Carata E, Piro G, Carpita NC, Mita G, Alifano P (2013) Sphingomonas cynarae sp. nov., a proteobacterium that produces an unusual type of sphingan. Int J Syst Evol Microbiol 63:72–79. doi:10.1099/ijs.0.032060-0
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. doi:10.1093/nar/22.22.4673
Thompson FL, Li Y, Gomez-Gil B, Thompson CC, Hoste B, Vandemeulebroecke K, Rupp GS, Pereira A, De Bem MM, Sorgeloos P, Swings J (2003) Vibrio neptunius sp. nov., Vibrio brasiliensis sp. nov. and Vibrio xuii sp. nov., isolated from the marine aquaculture environment (bivalves, fish, rotifers and shrimps). Int J Syst Evol Microbiol 53(1):245–252. doi:10.1099/ijs.0.02447-0
Thompson FL, Thompson CC, Naser S, Hoste B, Vandemeulebroecke K, Munn C, Bourne D, Swings J (2005) Photobacterium rosenbergii sp. nov. and Enterovibrio coralii sp. nov., vibrios associated with coral bleaching. Int J Syst Evol Microbiol 55:913–917. doi:10.1099/ijs.0.63370-0
Urbanczyk H, Yoshitoshi O, Hayashi T (2013) Taxonomic revision of Harveyi clade bacteria (family Vibrionaceae) based on analysis of whole genome sequences. Int J Syst Evol Microbiol 63(7):2742–2751. doi:10.1099/ijs.0.051110-0
Venkateswaran K, Dohmoto N (2000) Pseudoalteromonas peptidolytica sp. nov., a novel marine mussel-thread-degrading bacterium isolated from the Sea of Japan. Int J Syst Evol Microbiol 50(2):565–574. doi:10.1099/00207713-50-2-565
Venkateswaran K, Moser DP, Dollhopf ME, Lies DP, Saffarini DA, MacGregor BJ, Ringelberg DB, White DC, Nishijima M, Sano H, Burghardt J, Stackebrandt E, Nealson KH (1999) Polyphasic taxonomy of the genus Shewanella and description of Shewanella oneidensis sp. nov. Int J Syst Bacteriol 49:705–724. doi:10.1099/00207713-49-2-705
Verlaque M, Durand C, Huisman JM, Boudouresque CF, Le Parco Y (2003) On the identity and origin of the Mediterranean invasive Caulerpa racemosa (Caulerpales, Chlorophyta). Eur J Phycol 38:325–339. doi:10.1080/09670260310001612592
Wahl M, Goecke F, Labes A, Dobretsov S, Weinberger F (2012) The second skin: ecological role of epibiotic biofilms on marine organisms. Front Microbiol 3:292. doi:10.3389/fmicb.2012.00292
Wichard T (2015) Exploring bacteria-induced growth and morphogenesis in the green macroalga order Ulvales (Chlorophyta). Front Plant Sci. doi:10.3389/fpls.2015.00086
Xiao P, Jiang M, Liu Y, Sun M, Zhang L, Jie L, Li G, Mo Z (2015) Splenic necrosis signs and pathogen detection in cultured half-smooth tongue sole, Cynoglossus semilaevis Günther. J Fish Dis 38(1):103–106. doi:10.1111/jfd.12213
Yoon JH, Oh TK (2005) Bacillus litoralis sp. nov., isolated from a tidal flat of the Yellow Sea in Korea. Int J Syst Evol Microbiol 55(5):1945–1948. doi:10.1099/ijs.0.63332-0
Yoshizawa S, Tsuruya Y, Fukui Y, Sawabe T, Yokota A, Kogure K, Higgins M, Carson J, Thompson FL (2012) Vibrio jasicida sp. nov., a member of the Harveyi clade, isolated from marine animals (packhorse lobster, abalone and Atlantic salmon). Int J Syst Evol Microbiol 62(8):1864–1870. doi:10.1099/ijs.0.025916-0
Acknowledgments
Financial support was provided by RITMARE Flagship Project and by the PRIN TETRIS funded by the Italian Ministry of University and Research. The European Community’s 7th Framework Programmes (FP7/2007–2013) for the project COCONET (Grant agreement No. 287844, http://www.coconet-fp7.eu/) and PERSEUS (Grant agreement No. 287600, http://www.perseus-net.eu/site/content.php) are also acknowledged. Christian Vaglio and the staff of the Marine Protected Area of Torre Guaceto helped in the field. Thanks are due to Dr. Maurizio Salvatore Tredici and Dr. Marco Lezzi for the support in the laboratory.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: F. Weinberger.
Reviewed by T. Aires and an undisclosed expert.
Rights and permissions
About this article
Cite this article
Rizzo, L., Fraschetti, S., Alifano, P. et al. The alien species Caulerpa cylindracea and its associated bacteria in the Mediterranean Sea. Mar Biol 163, 4 (2016). https://doi.org/10.1007/s00227-015-2775-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00227-015-2775-9