Abstract
Ion mobility coupling to mass spectrometry facilitates enhanced identification certitude. Further coupling to liquid chromatography results in multi-dimensional analytical methods, especially suitable for complex matrices with structurally similar compounds. Modified nucleosides represent a large group of very similar members linked to aberrant proliferation. Besides basal production under physiological conditions, they are increasingly excreted by transformed cells and subsequently discussed as putative biomarkers for various cancer types. Here, we report a method for modified nucleosides covering 37 species. We determined collisional cross-sections with high reproducibility from pure analytical standards. For sample purification, we applied an optimized phenylboronic acid solid-phase extraction on media obtained from four different pancreatic cancer cell lines. Our analysis could discriminate different subtypes of pancreatic cancer cell lines. Importantly, they could clearly be separated from a pancreatic control cell line as well as blank medium. m1A, m27G, and Asm were the most important features discriminating cancer cell lines derived from well-differentiated and poorly differentiated cancers. Eventually, we suggest the analytical method reported here for future tumor-marker identification studies.

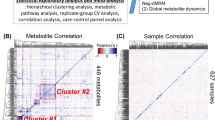
Graphical abstract






Similar content being viewed by others
References
Rahib L, Smith BD, Aizenberg R, Rosenzweig AB, Fleshman JM, Matrisian LM. Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res. 2014;74:2913–21. https://doi.org/10.1158/0008-5472.CAN-14-0155.
Siegel R, Ma J, Zou Z, Jemal A. Cancer statistics, 2014. CA Cancer J Clin. 2014;64:9–29. https://doi.org/10.3322/caac.21208.
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2013. CA Cancer J Clin. 2013;63:11–30. https://doi.org/10.3322/caac.21166.
Neoptolemos JP, Kleeff J, Michl P, Costello E, Greenhalf W, Palmer DH. Therapeutic developments in pancreatic cancer: current and future perspectives. Nat Rev Gastroenterol Hepatol. 2018;15:333–48. https://doi.org/10.1038/s41575-018-0005-x.
Kleeff J, Korc M, Apte M, La Vecchia C, Johnson CD, Biankin AV, et al. Pancreatic cancer. Nat Rev Dis Primers. 2016;2:16022. https://doi.org/10.1038/nrdp.2016.22.
Partensky C. Toward a better understanding of pancreatic ductal adenocarcinoma: glimmers of hope? Pancreas. 2013;42:729–39. https://doi.org/10.1097/MPA.0b013e318288107a.
Schnelldorfer T, Ware AL, Sarr MG, Smyrk TC, Zhang L, Qin R, et al. Long-term survival after pancreatoduodenectomy for pancreatic adenocarcinoma: is cure possible? Ann Surg. 2008;247:456–62. https://doi.org/10.1097/SLA.0b013e3181613142.
Oettle H, Neuhaus P, Hochhaus A, Hartmann JT, Gellert K, Ridwelski K, et al. Adjuvant chemotherapy with gemcitabine and long-term outcomes among patients with resected pancreatic cancer: the CONKO-001 randomized trial. JAMA. 2013;310:1473–81. https://doi.org/10.1001/jama.2013.279201.
Gillen S, Schuster T, Zum Meyer Büschenfelde C, Friess H, Kleeff J. Preoperative/neoadjuvant therapy in pancreatic cancer: a systematic review and meta-analysis of response and resection percentages. PLoS Med. 2010;7:e1000267. https://doi.org/10.1371/journal.pmed.1000267.
Scarà S, Bottoni P, Scatena R. CA 19-9: biochemical and clinical aspects. Adv Exp Med Biol. 2015;867:247–60. https://doi.org/10.1007/978-94-017-7215-0_15.
Takai E, Yachida S. Circulating tumor DNA as a liquid biopsy target for detection of pancreatic cancer. World J Gastroenterol. 2016;22:8480–8. https://doi.org/10.3748/wjg.v22.i38.8480.
Cote GA, Gore AJ, McElyea SD, Heathers LE, Xu H, Sherman S, et al. A pilot study to develop a diagnostic test for pancreatic ductal adenocarcinoma based on differential expression of select miRNA in plasma and bile. Am J Gastroenterol. 2014;109:1942–52. https://doi.org/10.1038/ajg.2014.331.
Nagayoshi Y, Nakamura M, Matsuoka K, Ohtsuka T, Mori Y, Kono H, et al. Profiling of autoantibodies in sera of pancreatic cancer patients. Ann Surg Oncol. 2014;21(Suppl 3):S459–65. https://doi.org/10.1245/s10434-014-3574-0.
Pan S, Chen R, Crispin DA, May D, Stevens T, McIntosh MW, et al. Protein alterations associated with pancreatic cancer and chronic pancreatitis found in human plasma using global quantitative proteomics profiling. J Proteome Res. 2011;10:2359–76. https://doi.org/10.1021/pr101148r.
Melo SA, Luecke LB, Kahlert C, Fernandez AF, Gammon ST, Kaye J, et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature. 2015;523:177–82. https://doi.org/10.1038/nature14581.
Ritchie SA, Akita H, Takemasa I, Eguchi H, Pastural E, Nagano H, et al. Metabolic system alterations in pancreatic cancer patient serum: potential for early detection. BMC Cancer. 2013;13:416. https://doi.org/10.1186/1471-2407-13-416.
Mayerle J, Kalthoff H, Reszka R, Kamlage B, Peter E, Schniewind B, et al. Metabolic biomarker signature to differentiate pancreatic ductal adenocarcinoma from chronic pancreatitis. Gut. 2018;67:128–37. https://doi.org/10.1136/gutjnl-2016-312432.
Di Gangi IM, Mazza T, Fontana A, Copetti M, Fusilli C, Ippolito A, et al. Metabolomic profile in pancreatic cancer patients: a consensus-based approach to identify highly discriminating metabolites. Oncotarget. 2016;7:5815–29. https://doi.org/10.18632/oncotarget.6808.
Mehta KY, Wu H-J, Menon SS, Fallah Y, Zhong X, Rizk N, et al. Metabolomic biomarkers of pancreatic cancer: a meta-analysis study. Oncotarget. 2017;8:68899–915. https://doi.org/10.18632/oncotarget.20324.
Kobayashi T, Nishiumi S, Ikeda A, Yoshie T, Sakai A, Matsubara A, et al. A novel serum metabolomics-based diagnostic approach to pancreatic cancer. Cancer Epidemiol Biomark Prev. 2013;22:571–9. https://doi.org/10.1158/1055-9965.EPI-12-1033.
Borek E, Baliga BS, Gehrke CW, Kuo CW, Belman S, Troll W, et al. High turnover rate of transfer RNA in tumor tissue. Cancer Res. 1977;37:3362–6.
Tusup M, Kundig T, Pascolo S. Epitranscriptomics of cancer. World J Clin Oncol. 2018;9:42–55. https://doi.org/10.5306/wjco.v9.i3.42.
Kammerer B, Frickenschmidt A, Muller CE, Laufer S, Gleiter CH, Liebich H. Mass spectrometric identification of modified urinary nucleosides used as potential biomedical markers by LC-ITMS coupling. Anal Bioanal Chem. 2005;382:1017–26. https://doi.org/10.1007/s00216-005-3232-2.
Zhang Y-R, Shi L, Wu H, Tang D-D, Wang S-M, Liu H-M, et al. Urinary modified nucleosides as novel biomarkers for diagnosis and prognostic monitoring of urothelial bladder cancer. Tumori. 2014;100:660–6. https://doi.org/10.1700/1778.19274.
Willmann L, Erbes T, Halbach S, Brummer T, Jäger M, Hirschfeld M, et al. Exometabolom analysis of breast cancer cell lines: metabolic signature. Sci Rep. 2015;5:13374. https://doi.org/10.1038/srep13374.
Hsu W-Y, Chen WT-L, Lin W-D, Tsai F-J, Tsai Y, Lin C-T, et al. Analysis of urinary nucleosides as potential tumor markers in human colorectal cancer by high performance liquid chromatography/electrospray ionization tandem mass spectrometry. Clin Chim Acta. 2009;402:31–7. https://doi.org/10.1016/j.cca.2008.12.009.
Frickenschmidt A, Frohlich H, Bullinger D, Zell A, Laufer S, Gleiter CH, et al. Metabonomics in cancer diagnosis: mass spectrometry-based profiling of urinary nucleosides from breast cancer patients. Biomarkers. 2008;13:435–49. https://doi.org/10.1080/13547500802012858.
Lu Z, Wang Q, Wang M, Fu S, Zhang Q, Zhang Z, et al. Using UHPLC Q-rap/MS as a complementary technique to in-depth mine UPLC Q-TOF/MS data for identifying modified nucleosides in urine. J Chromatogr B Analyt Technol Biomed Life Sci. 2017;1051:108–17. https://doi.org/10.1016/j.jchromb.2017.03.002.
Pringle SD, Giles K, Wildgoose JL, Williams JP, Slade SE, Thalassinos K, et al. An investigation of the mobility separation of some peptide and protein ions using a new hybrid quadrupole/travelling wave IMS/oa-ToF instrument. Int J Mass Spectrom. 2007;261:1–12. https://doi.org/10.1016/j.ijms.2006.07.021.
Stephan S, Jakob C, Hippler J, Schmitz OJ. A novel four-dimensional analytical approach for analysis of complex samples. Anal Bioanal Chem. 2016;408:3751–9. https://doi.org/10.1007/s00216-016-9460-9.
Stark TD, Ranner J, Stiglbauer B, Weiss P, Stark S, Balemba OB, et al. Construction and application of a database for a five dimensional identification of natural compounds in Garcinia species by means of UPLC-ESI-TWIMS-TOF-MS-introducing gas phase polyphenol conformer drift time distribution intensity ratios. J Agric Food Chem. 2018. https://doi.org/10.1021/acs.jafc.8b06157 .
Rister AL, Martin TL, Dodds ED. Application of group I metal adduction to the separation of steroids by traveling wave ion mobility spectrometry. J Am Soc Mass Spectrom. 2019;30:248–55. https://doi.org/10.1007/s13361-018-2085-9.
Keelor JD, Zambrzycki S, Li A, Clowers BH, Fernandez FM. Atmospheric pressure drift tube ion mobility-Orbitrap mass spectrometry: initial performance characterization. Anal Chem. 2017;89:11301–9. https://doi.org/10.1021/acs.analchem.7b01866.
Jeanne Dit Fouque K, Bisram V, Hegemann JD, Zirah S, Rebuffat S, Fernandez-Lima F. Structural signatures of the class III lasso peptide BI-32169 and the branched-cyclic topoisomers using trapped ion mobility spectrometry–mass spectrometry and tandem mass spectrometry. Anal Bioanal Chem. 2019. https://doi.org/10.1007/s00216-019-01613-8.
Kanu AB, Hampikian G, Brandt SD, Hill HH Jr. Ribonucleotide and ribonucleoside determination by ambient pressure ion mobility spectrometry. Anal Chim Acta. 2010;658:91–7. https://doi.org/10.1016/j.aca.2009.10.058.
Quinn R, Basanta-Sanchez M, Rose RE, Fabris D. Direct infusion analysis of nucleotide mixtures of very similar or identical elemental composition. J Mass Spectrom. 2013;48:703–12. https://doi.org/10.1002/jms.3207.
Rose RE, Quinn R, Sayre JL, Fabris D. Profiling ribonucleotide modifications at full-transcriptome level: a step toward MS-based epitranscriptomics. RNA. 2015;21:1361–74. https://doi.org/10.1261/rna.049429.114.
Loukopoulos P, Kanetaka K, Takamura M, Shibata T, Sakamoto M, Hirohashi S. Orthotopic transplantation models of pancreatic adenocarcinoma derived from cell lines and primary tumors and displaying varying metastatic activity. Pancreas. 2004;29:193–203.
Berrozpe G, Schaeffer J, Peinado MA, Real FX, Perucho M. Comparative analysis of mutations in the p53 and K-ras genes in pancreatic cancer. Int J Cancer. 1994;58:185–91.
Kyriazis AA, Kyriazis AP, Sternberg CN, Sloane NH, Loveless JD. Morphological, biological, biochemical, and karyotypic characteristics of human pancreatic ductal adenocarcinoma Capan-2 in tissue culture and the nude mouse. Cancer Res. 1986;46:5810–5.
Fahrmann JF, Bantis LE, Capello M, Scelo G, Dennison JB, Patel N, et al. A plasma-derived protein-metabolite multiplexed panel for early-stage pancreatic cancer. J Natl Cancer Inst. 2018. https://doi.org/10.1093/jnci/djy126.
Metzgar RS, Gaillard MT, Levine SJ, Tuck FL, Bossen EH, Borowitz MJ. Antigens of human pancreatic adenocarcinoma cells defined by murine monoclonal antibodies. Cancer Res. 1982;42:601–8.
Moore PS, Sipos B, Orlandini S, Sorio C, Real FX, Lemoine NR, et al. Genetic profile of 22 pancreatic carcinoma cell lines. Analysis of K-ras, p53, p16 and DPC4/Smad4. Virchows Arch. 2001;439:798–802.
Yunis AA, Arimura GK, Russin DJ. Human pancreatic carcinoma (MIA PaCa-2) in continuous culture: sensitivity to asparaginase. Int J Cancer. 1977;19:128–35.
Sun C, Yamato T, Furukawa T, Ohnishi Y, Kijima H, Horii A. Characterization of the mutations of the K-ras, p53, p16, and SMAD4 genes in 15 human pancreatic cancer cell lines. Oncol Rep. 2001;8:89–92.
Lieber M, Mazzetta J, Nelson-Rees W, Kaplan M, Todaro G. Establishment of a continuous tumor-cell line (panc-1) from a human carcinoma of the exocrine pancreas. Int J Cancer. 1975;15:741–7.
Dodds JN, May JC, McLean JA. Correlating resolving power, resolution and collision cross section: unifying cross platform assessment of separation efficiency in ion mobility spectrometry. Anal Chem. 2017;89:12176–84. https://doi.org/10.1021/acs.analchem.7b02827.
Smith DP, Knapman TW, Campuzano I, Malham RW, Berryman JT, Radford SE, et al. Deciphering drift time measurements from travelling wave ion mobility spectrometry-mass spectrometry studies. Eur J Mass Spectrom (Chichester). 2009;15:113–30. https://doi.org/10.1255/ejms.947.
Chong J, Soufan O, Li C, Caraus I, Li S, Bourque G, et al. MetaboAnalyst 4.0: towards more transparent and integrative metabolomics analysis. Nucleic Acids Res. 2018;46:W486–94. https://doi.org/10.1093/nar/gky310.
Cantara WA, Crain PF, Rozenski J, McCloskey JA, Harris KA, Zhang X, et al. The RNA modification database, RNAMDB: 2011 update. Nucleic Acids Res. 2011;39:D195–201. https://doi.org/10.1093/nar/gkq1028.
Willmann L, Erbes T, Krieger S, Trafkowski J, Rodamer M, Kammerer B. Metabolome analysis via comprehensive two-dimensional liquid chromatography: identification of modified nucleosides from RNA metabolism. Anal Bioanal Chem. 2015;407:3555–66. https://doi.org/10.1007/s00216-015-8516-6.
Li H-Y, Wang S-M, Liu H-M, Li J, Han D, Bu S-S, et al. Analysis of modified nucleosides in the urine of patients with malignant cancer by liquid chromatography/electrospray ionization mass spectrometry. Rapid Commun Mass Spectrom. 2008;22:3161–71. https://doi.org/10.1002/rcm.3721.
Schlimpert M, Lagies S, Budnyk V, Müller B, Walz G, Kammerer B. Metabolic phenotyping of Anks3 depletion in mIMCD-3 cells - a putative nephronophthisis candidate. Sci Rep. 2018;8:9022. https://doi.org/10.1038/s41598-018-27389-y.
Paglia G, Williams JP, Menikarachchi L, Thompson JW, Tyldesley-Worster R, Halldórsson S, et al. Ion mobility derived collision cross sections to support metabolomics applications. Anal Chem. 2014;86:3985–93. https://doi.org/10.1021/ac500405x.
Acknowledgements
This work was supported by Project B01 of the collaborative research initiative (SFB 1140 - KIDGEM) to M.S. and B.K. by the German Research Foundation (DFG).
Author information
Authors and Affiliations
Contributions
Simon Lagies, Manuel Schlimpert, Lukas M. Braun, Thalia Erbes, Uwe A. Wittel, and Bernd Kammerer designed the study. Simon Lagies, Manuel Schlimpert, Lukas M. Braun, and Michel Kather performed the experiments and data analysis. Johannes Plagge aided in data processing and image creation. Simon Lagies, Lukas M. Braun, and Manuel Schlimpert wrote the manuscript.
Corresponding author
Ethics declarations
Informed consent
Informed consent was obtained for KHM1106 cell line usage and approved by the local Ethics Committee Freiburg (126/17 and 371/14) and registered at the German Clinical Trials Register (DRKS-ID DRKS00007561).
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Published in the topical collection Close-Up of Current Developments in Ion Mobility Spectrometry with guest editor Gérard Hopfgartner.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 3044 kb)
Rights and permissions
About this article
Cite this article
Lagies, S., Schlimpert, M., Braun, L.M. et al. Unraveling altered RNA metabolism in pancreatic cancer cells by liquid-chromatography coupling to ion mobility mass spectrometry. Anal Bioanal Chem 411, 6319–6328 (2019). https://doi.org/10.1007/s00216-019-01814-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-019-01814-1