Abstract
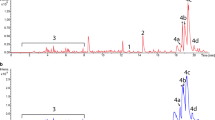
Activation and silencing of antibiotic production was achieved in Streptomyces albus J1074 and Streptomyces lividans TK21 after introduction of genes within the thienamycin cluster from S. cattleya. Dramatic phenotypic and metabolic changes, involving activation of multiple silent secondary metabolites and silencing of others normally produced, were found in recombinant strains harbouring the thienamycin cluster in comparison to the parental strains. In S. albus, ultra-performance liquid chromatography purification and NMR structural elucidation revealed the identity of four structurally related activated compounds: the antibiotics paulomycins A, B and the paulomenols A and B. Four volatile compounds whose biosynthesis was switched off were identified by gas chromatography–mass spectrometry analyses and databases comparison as pyrazines; including tetramethylpyrazine, a compound with important clinical applications to our knowledge never reported to be produced by Streptomyces. In addition, this work revealed the potential of S. albus to produce many others secondary metabolites normally obtained from plants, including compounds of medical relevance as dihydro-β-agarofuran and of interest in perfume industry as β-patchoulene, suggesting that it might be an alternative model for their industrial production. In S. lividans, actinorhodins production was strongly activated in the recombinant strains whereas undecylprodigiosins were significantly reduced. Activation of cryptic metabolites in Streptomyces species might represent an alternative approach for pharmaceutical drug discovery.







Similar content being viewed by others
References
Argoudelis AD, Brinkley TA, Brodasky TF, Buege JA, Meyer HF, Mizsak SA (1982) Paulomycins A and B. Isolation and characterization. J Antibiot 35:285–294
Argoudelis AD, Baczynskyj L, Mizsak SA, Shilliday FB (1988) O-demethylpaulomycins A and B, U-77,802 and U-77,803, paulomenols A and B, new metabolites produced by Streptomyces paulus. J Antibiot 41:1316–1330
Baltz RH (2008) Renaissance in antibacterial discovery from actinomycetes. Curr Opin Pharmacol 8:557–563
Barbe V, Bouzon M, Mangenot S, Badet B, Poulain J, Segurens B, Vallenet D, Marlière P, Weissenbach J (2011) Complete genome sequence of Streptomyces cattleya NRRL 8057, a producer of antibiotics and fluorometabolites. J Bacteriol 193:5055–5056
Blanco G (2012) Comparative analysis of a cryptic thienamycin-like gene cluster identified in Streptomyces flavogriseus by genome mining. Arch Microbiol 194:549–555
Blin K, Medema MH, Kazempour D, Fischbach MA, Breitling R, Takano E, Weber T (2013) antiSMASH 2.0—a versatile platform for genome mining of secondary metabolite producers. Nucleic Acids Res 41:204–212
Bodner JJ, Phelan RM, Freeman MF, Li R, Townsend CA (2010) Non-heme iron oxygenases generate natural structural diversity in carbapenem antibiotics. J Am Chem Soc 132:12–13
Brakhage AA, Schroeckh V (2011) Fungal secondary metabolites—strategies to activate silent gene clusters. Fungal Genet Biol 48:15–22
Cane DE, Ikeda H (2011) Exploration and mining of the bacterial terpenome. Acc Chem Res 45:463–472
Chiang Y-M, Chang S-L, Oakley BR, Wang CC (2011) Recent advances in awakening silent biosynthetic gene clusters and linking orphan clusters to natural products in microorganisms. Curr Opin Chem Biol 15:137–143
Demain AL (2009) Antibiotics: natural products essential to human health. Med Res Rev 29:821–842
Demain AL, Jackson M, Trenner NR (1967) Thiamine-dependent accumulation of tetramethylpyrazine accompanying a mutation in the isoleucine-valine pathway. J Bacteriol 94:323–326
Fernández E, Weissbach U, Reillo CS, Braña AF, Méndez C, Rohr J, Salas JA (1998) Identification of two genes from Streptomyces argillaceus encoding two glycosyltransferases involved in the transfer of a disaccharide during the biosynthesis of the antitrumor drug mithramycin. J Bacteriol 180:4929–4937
Freeman MF, Moshos KA, Bodner MJ, Li R, Townsend CA (2008) Four enzymes define the incorporation of coenzyme A in thienamycin biosynthesis. Proc Natl Acad Sci USA 105:11128–11133
Gao JM, Wu WJ, Zhang JW, Konishi Y (2007) The dihydro-beta-agarofuran sesquiterpenoids. Nat Prod Rep 24:1153–1189
Gust B, Challis GL, Fowler K, Kieser T, Chater K (2003) PCR-targeted Streptomyces gene replacement identifies a protein domain needed for biosynthesis of the sesquiterpene soil odor geosmin. Proc Natl Acad Sci USA 100:1541–1546
Hamed RB, Batchelar ET, Mecinovic J, Claridge TD, Schofield CJ (2009) Evidence that thienamycin biosynthesis proceeds via C-5 epimerization: ThnE catalyzes the formation of (2S, 5S)-trans-carboxymethylproline. ChemBioChem 10:246–250
Hamed RB, Gomez-Castellanos JR, Henry L, Ducho C, McDonough MA, Schofield CJ (2013) The enzymes of β-lactam biosynthesis. Nat Prod Rep 30:21–107
Holden MTG, McGowan SJ, Bycroft BW, Stewart GSAB, Williams P, Salmond GPC (1998) Cryptic carbapenem antibiotic production genes are widespread in Erwinia carotovora: facile trans activation by carR transcriptional regulator. Microbiology 144:1495–1508
Horinouchi S, Beppu T (1984) Production in large quantities of actinorhodin and undecylprodigiosin induced by afsR in Streptomyces lividans. Agric Biol Chem 48:2131–2133
Huang J, Shi J, Molle V, Sohlberg B, Weaver D, Bibb MJ, Karoonuthaisiri N, Lih CJ, Kao CM, Buttner MJ, Cohen SN (2005) Cross-regulation among disparate antibiotic biosynthetic pathways of Streptomyces coelicolor. Mol Microbiol 58:1276–1287
Jachymova J, Votruba J, Viden I, Rezanka T (2002) Identification of Streptomyces odor spectrum. Folia Microbiol 47:37–41
Kahan JS, Kahan FM, Goegelman R, Currie SA, Jackson M, Stapley EO, Miller TW, Miller AK, Hendlin D, Mochales S, Hernandez S, Woodruff HB, Birnbaum J (1979) Thienamycin, a new beta-lactam antibiotic. I. Discovery, taxonomy, isolation and physical properties. J Antibiot 23:1255–1265
Kieser T, Bibb MJ, Buttner MJ, Chater KF, Hopwood DA (2000) Practical Streptomyces genetics. John Innes Foundation, Norwich
Majer J, Chater K (1987) Streptomyces albus G produces an antibiotic complex identical to paulomycins A and B. J Gen Microbiol 133:2503–2507
Medema MH, Blin K, Cimermancic P, Jager V, Zakrzewski P, Fischbach MA, Weber T, Takano E, Breitling R (2011) antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res 39:339–346
Müller R, Rappert S (2010) Pyrazines: occurrence, formation and biodegradation. Appl Microbiol Biotechnol 85:1315–1320
Novak E (1988) Treating Chlamydia infections with paulomycin. The Upjohn Company. Patent PCT/US1987/002420
Núñez LE, Méndez C, Braña AF, Blanco G, Salas JA (2003) The biosynthetic gene cluster for the beta-lactam carbapenem thienamycin in Streptomyces cattleya. Chem Biol 10:1–20
Ou X, Zhan B, Zhan L, Zhao G, Xiaoming D (2009) Characterization of rrdA, a TetR family protein gene involved in the regulation of secondary metabolism in Streptomyces coelicolor. Appl Environ Microbiol 75:2158–2165
Penn J, Li X, Whiting A, Latif M, Gibson T, Silva CJ, Brian P, Davies J, Miao V, Wrigley SK, Baltz RH (2006) Heterologous production of daptomycin in Streptomyces lividans. J Ind Microbiol Biotechnol 33:121–128
Perez J, Muñoz-Dorado J, Braña AF, Shimkets LJ, Sevillano L, Santamaría RI (2011) Myxococcus xanthus induces actinorhodin overproduction and aerial mycelium formation by Streptomyces coelicolor. Microb Biotechnol 4:175–183
Pollak FC, Berger RG (1996) Geosmin and related volatiles in bioreactor-cultured Streptomyces citreus CBS 109.60. Appl Environ Microbiol 62:1295–1299
Rajini KS, Aparna P, Salikala Ch, Ramana ChV (2011) Microbial metabolism of pyrazynes. Crit Rev Microbiol 37:99–112
Rodríguez M, Núñez LE, Braña AF, Méndez C, Salas JA, Blanco G (2008) Identification of transcriptional activators for thienamycin and cephamycin C biosynthetic genes within the thienamycin gene cluster from Streptomyces cattleya. Mol Microbiol 69:633–645
Rodríguez M, Méndez Salas JA, Blanco G (2010) Transcriptional organization of ThnI-regulated thienamycin biosynthetic genes in Streptomyces cattleya. J Antibiot 63:135–138
Rodríguez M, Núñez LE, Braña AF, Méndez C, Salas JA, Blanco G (2011) Mutational analysis of the thienamycin biosynthetic gene cluster from Streptomyces cattleya. Antimicrob Agents Chemother 55:1638–1649
Romero NM, Parro V, Malpartida F, Mellado RP (1992) Heterologous activation of the actinorhodin biosynthetic pathway in Streptomyces lividans. Nucleic Acids Res 20:2767–2772
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Shin S-K, Xu D, Kwo H-J, Syh J-W (2006) S-adenosylmethionine activates adpA transcription and promotes streptomycin biosynthesis in Streptomyces griseus. FEMS Microbiol Lett 259:53–59
van Wezel GP, McDowall KJ (2011) The regulation of secondary metabolism of Streptomyces: new links and experimental advances. Nat Prod Rep 28:1311–1333
Walsh CT, Fischbach MA (2010) Natural products version 2.0: connecting genes to molecules. J Am Chem Soc 132:2469–2493
Wiley FP (1983) Paulomycin derivatives. The Upjohn Company. Patent PCT/US1983/4520191
Zaburannyi N, Rabyk M, Ostash B, Fedorenko V, Luzhetskyy A (2014) Insights into naturally minimised Streptomyces albus J1074 genome. BMC Genom. doi:10.1186/1471-2164-15-97
Zerikly M, Challis GL (2009) Strategies for the discovery of new natural products by genome mining. ChemBioChem 10:625–633
Zhang Z, Wei T, Hou J, Li G, Yu S, Xin W (2003) Tetramethylpyrazine scavenges superoxide anion and decreases nitric oxide production in human polymorphonuclear leukocytes. Life Sci 72:2465–2472
Zhu H, Sandiford SK, vanWezel GP (2013) Triggers and cues that activate antibiotic production by actinomycetes. J Ind Microbiol Biotechnol. doi:10.1007/s10295-013-1309-z
Acknowledgments
This study was financial supported by the University of Oviedo (Ayuda Puente UNOV-10-MA-4). Thanks to Miguel Campoamor for his excellent GC–MS technical assistance. The authors are grateful to José L. Caso and José A. Guijarro for their continuous support. We also want to thank José A. Salas, Carmen Méndez and Luz E. Núñez for previous work on thienamycin.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Braña, A.F., Rodríguez, M., Pahari, P. et al. Activation and silencing of secondary metabolites in Streptomyces albus and Streptomyces lividans after transformation with cosmids containing the thienamycin gene cluster from Streptomyces cattleya . Arch Microbiol 196, 345–355 (2014). https://doi.org/10.1007/s00203-014-0977-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-014-0977-z