Abstract
The genome of Synechococcus elongatus PCC 7942 encodes six peroxiredoxins (Prx). Single genes are present each for a 1-Cys Prx and a 2-Cys Prx, while four genes code for PrxQ-like proteins (prxQ-A1, -A2, -A3 and B). Their transcript accumulation varies with growth conditions in a gene-specific manner (Stork et al. in J Exp Bot 56:3193–3206, 2005). To address their functional properties, members of the prx gene family were produced as recombinant proteins and analysed for their peroxide detoxification capacity and quaternary structure by size exclusion chromatography. Independent of the reduction state, the 2-Cys Prx separated as oligomer, the 1-Cys Prx as dimer and the PrxQ-A1 as monomer. PrxQ-A2 was inactive in our assays, 1-Cys Prx activity was unaffected by addition of TrxA, while all others were stimulated to a variable extent by addition of E. coli thioredoxin. Sensitivity towards cumene hydroperoxide treatment of E. coli BL21 cells expressing the cyanobacterial PrxQ-A1 to A3 proteins was greatly reduced, while expression of the other Prx had no effect. The study shows differentiation of Prx functions in S. elongatus PCC 7942 which is discussed in relation to potential roles in site- and stress-specific defence.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Following the evolution of the first cyanobacteria about 3.5 billion years ago the earth atmosphere enriched in oxygen and slowly reached the present day concentration of 21% (v/v) (Schopf 1993). From early times on cyanobacteria had to cope with three types of oxidative threats, (1) O2 released in the vicinity of the thylakoid membranes by their own oxygenic metabolism of photosynthesis, (2) the local O2 enrichment in photosynthesising cyanobacterial populations, and (3) the rising atmospheric O2 concentration. Since in addition a cyanobacterium served as endocytobiont that turned into the chloroplast of plants, exploring the antioxidant system of cyanobacteria promises important insight into the requirements for antioxidant defence of organisms with oxygenic photosynthesis (Koksharova et al. 2006; Olson and Blankenship 2004).
Cyanobacteria constitute a taxonomic group distinct from other eubacteria, namely the gram-negative and -positive bacteria. This separate classification is justified not only due to their photosynthetic capacity but also based on other characteristics such as the multilayer cell wall (Xiong et al. 2000; Hansmann and Martin 2000). Synechococcus elongatus PCC 7942 (synonymous to Anacystis nidulans R2) which was used in this study is an obligate photoautotrophic cyanobacterium that as early as in 1970 was transformed with foreign DNA (Shestakov and Khyen 1970). Its full genome sequence is available at NCBI (NC_007604).
Imbalances in energy metabolism result in increased production rates of reactive oxygen species (ROS) such as the superoxide anion radical, hydrogen peroxide and hydroxyl radical. ROS in turn damage macromolecules and elicit lipid peroxidation or peroxinitrite generation. In respiratory and photosynthetic electron transport particularly under conditions of overreduction ROS are produced at various sites such as photosystem II, the plastoquinon pool, photosystem I in photosynthesis, and complex I and III in respiration (Elstner 1990). To counteract oxidative stress, cells maintain an elaborate antioxidant defence system. Synechocystis PCC 6803 is the cyanobacterium studied best with respect to its antioxidant system. It expresses a superoxide dismutase (slr1516), a catalase peroxidase (sll1987) and five peroxiredoxins (Prx) (Kaneko et al. 1996; Kobayashi et al. 2004; Hosoya-Matsuda et al. 2005; Dietz et al. 2005; Stork et al. 2005). Perelman et al. (2003) analysed a double mutant of S. elongatus PCC 7942 deficient in both the catalase and the 2-Cys Prx homologue. The cells survived in the presence of toxic H2O2. The authors concluded that additional H2O2 detoxification pathways must coexist with the catalase and 2-Cys Prx in S. elongatus PCC 7942 cells. In the light of the presence of a complex Prx protein family with six members, such a functional redundancy may not be surprising.
Prx constitute a protein family of evolutionary old peroxidases with high affinity to their peroxide substrates (Hofmann et al. 2002). Prx are present in all organisms. Their enzymatic mechanism of peroxide reduction depends on a catalytic cysteinyl residue in conserved amino acid environment that reacts with the peroxide substrate under formation of a sulphenic acid intermediate (Dietz 2003). Hydrogen peroxide, alkyl hydroperoxides and peroxinitrite are reported Prx substrates. The catalytic efficiency with diverse substrates varies among the four different Prx forms which are distinguished based on the intramolecular location of their catalytic Cys residues and the subsequent mechanism of active site regeneration: (1) The first identified Prx group was later on denominated 2-Cys Prx. 2-Cys Prx function as homodimers in a head-to-tail arrangement. Each subunit has a catalytic and a resolving cysteinyl residue (Kim et al. 1988; Chae et al. 1993; Baier and Dietz 1996). Upon reaction with the peroxide substrate, an intermolecular disulphide bridge is formed between both subunits following the intermittent formation of the sulphenic acid derivative of the catalytic Cys. Reductive regeneration of 2-Cys Prx is achieved by thioredoxins and cyclophilins (Collin et al. 2003; Dietz 2003; Laxa et al. 2007). Plant 2-Cys Prx are posttranslationally imported into chloroplasts (Baier and Dietz 1997); (2) 1-Cys Prx lack a resolving Cys. The sulphenic acid intermediate is stabilised within the molecule prior to the direct regeneration by a yet unknown thiol regenerant (Stacy et al. 1996, 1999); (3) The PrxQ protein carries the two interacting Cys in close vicinity spaced by only four amino acids. Higher plant PrxQ is a chloroplast enzyme (Rouhier et al. 2004) attached to the thylakoid membrane (Lamkemeyer et al. 2006) and possibly imported into the thylakoid lumen (Petersson et al. 2006). PrxQ are absent from animals. In prokaryotic systems, the PrxQ is often named bacterioferritin comigratory protein (Bcp). Some bacterial PrxQ isoforms lack the resolving Cys. Plant PrxQ functions in context of photosynthesis (Lamkemeyer et al. 2006); (4) Type II Prx, together with PrxQ, also called atypical 2-Cys Prx form the disulphide bridge within one polypeptide. In this case the catalytic Cys is in 24 aa distance from the resolving Cys. Type II Prx are regenerated by glutaredoxins, thioredoxins or glutathione (Rouhier et al. 2001; Bréhélin et al. 2003; Finkemeier et al. 2005).
With its five members in Synechocystis PCC 6803, six in S. elongatus PCC 7942 (Stork et al. 2005) and seven in Anabaena sp. 7120 (Latifi et al. 2007), respectively, the peroxiredoxin gene families in cyanobacteria are larger than in other prokaryotic organisms. All organisms appear to possess at least one Prx, which most frequently is a 2-Cys Prx. In addition and with a high degree of variability most organisms contain an atypical 2-Cys Prx, i.e. either a type II Prx or a PrxQ-like protein, and a 1-Cys Prx, respectively. For example, in gram-negative and -positive bacteria, the Prx count gives figures between one (Lactobacillus plantarum) and four (Mycobacterium bovis AF2111/97, Engemann 2006) with most often two or three members. As representative example, Helicobacter pylori expresses one 2-Cys Prx (AhpC thiol-specific antioxidant), a Bcp and a thiol peroxidase (Tpx) (Wang et al. 2006). It is tempting to assume that the large Prx number in cyanobacteria is linked to various metabolic requirements in context of oxygenic photosynthesis.
This work aimed at characterising the prx gene family previously identified in S. elongatus PCC 7942 (named Se-Prx; Stork et al. 2005). To this end prx genes were cloned as cDNAs, heterologously expressed in E. coli, purified and analysed for their catalytic properties, as well as characterised for their functional aggregation state. In addition a peroxide tolerance test was performed with E. coli BL21 that expressed cyanobacterial prx genes. The study shows pronounced differences for these Prx proteins and suggests distinct functions in S. elongatus PCC 7942.
Materials and methods
Strains and cDNA cloning
S. elongatus PCC 7942 was originally obtained from the Pasteur Culture Collection (Paris, France) and maintained in the collection of the institute. Cells were cultivated in 250 ml gas wash-bottles containing BG11 medium and aerated with air enriched in 2% CO2 at 30°C (Stephan et al. 2000). The medium was inoculated with cells at an optical density of OD750 nm = 0.35. The RNA was isolated using a hot acidic phenol extraction procedure (Reddy et al. 1990) and the RNeasy Kit (Qiagen, Hilden, Germany). Using the gene-specific primers compiled in Table 1, the coding regions were reverse-transcribed from the RNA with MMLV reverse transcriptase (Promega) and the obtained cDNA amplified with Phusion™-polymerase (Finnzymes Oy, Espoo, Finland). PCR products were purified with ‘Wizard SV Gel and PCR Clean-Up System’ (Promega, Mannheim) and cloned into pCR T7/NT-TOPO-TA vector (Invitrogen). TOP10 F′-E. coli cells were transformed and tested for the presence of insertions by colony PCR. Following confirmation of the sequence fidelity by DNA sequencing, heterologous expression of the proteins was conducted in BL21(DE3)pLysS E. coli cells (Horling et al. 2002).
Production of recombinant proteins
Each E. coli BL21 strain harbouring the specified Prx/pCR T7/NT-TOPO-TA vector was grown over night in two Erlenmeyer flasks with 100 ml LB medium each supplemented with 100 μg/ml ampicillin. The precultures were transferred to 1 L medium and grown further to an OD 0.6–0.8. To induce expression IPTG (isopropyl-β-D-thiogalactopyranoside) was added to 0.36 mmol/L final concentration. Protein accumulation was monitored in harvested aliquots by SDS-PAGE analysis. After 4 h, the cells were sedimented by centrifugation and stored at −20°C. The thawed cell pellets were lysed in 0 mmol/L NaH2PO4, 300 mmol/L NaCl, 10 mmol/L imidazole, 10 mmol/L ascorbate, pH 8.0 with repeated ultrasonication. Recombinant proteins were purified by Ni-NTA affinity chromatography as described before (Horling et al. 2003). Purified and quantified proteins were dialysed against 40 mmol/L potassium phosphate buffer (pH 7.0) three times for 4 h, each. Protein concentrations were determined using the BioRad protein reagent with bovine serum albumin as standard. Analysis by SDS polyacrylamide gel electrophoresis was conducted according to standard protocols. E. coli TrxA and thioredoxin reductase were overexpressed and purified as described by Yamamoto et al. (1999). The anti his-tag antibody was obtained from SIGMA Chemicals.
Peroxiredoxin activity
The enzyme assay mixture contained 40 mmol/L potassium phosphate buffer (pH 7.0), 10 mmol/L dithiothreitol and 2.5 to 5 μmol/L recombinant peroxiredoxin. Hydrogen peroxide, butylhydroperoxide and cumene hydroperoxide at the concentrations as indicated were employed as substrates for the Prx reaction. For activity determination, 10 μl aliquots were taken after 0, 30, 60, 90, 120, 180, 240 and 300 s of incubation and injected to 160 μl 12.5% trichloroacetic acid. After addition of 40 μl of 10 mmol/L (NH4)2Fe(SO4)2 and 20 μl of 2.5 mol/L KSCN, the colour development was assayed at 480 nm by use of a microtiterplate reader. Appropriate blanks and controls were subtracted. Quantification was achieved by comparing the absorption obtained in the enzyme assay with absorption values of defined peroxide standards. The NADPH-coupled peroxidase activity was conducted in 40 mmol/L K-phosphate buffer, pH 7.0 supplemented with 1 mmol/L ethylenediamine tetraacetic acid (EDTA), 220 μmol/L NADPH + H+, 0.5 μmol/L E. coli thioredoxin reductase, 3 μmol/L E. coli thioredoxin TrxA, 2 μmol/L 2-Cys Prx or 5 μmol/L PrxQ-A1/3. The reaction was started by addition of H2O2 with concentrations varying between 10 and 100 μmol/L. The change in absorption was monitored at 340 nm and rates were calculated with the molar extinction coefficient εNADPH = 6,220 L/mol cm.
Size exclusion chromatography
Prx proteins (100 μl at 1 μg/μL) were loaded at the flow rate of 0.5 ml/min on the Superose 12 (10/300) column equilibrated with 40 mmol/L K-phosphate buffer (pH 7.0). The proteins were reduced in the presence of 10 mmol/L dithiothreitol (20 min, 4°C) or oxidised with 500 μmol/L H2O2 (15 min) with subsequent addition of 2.5 mmol/L ascorbate (final concentration, 5 min) to quench the H2O2 in the test solution. The well established separation behaviour of the system (König et al. 2003) was routinely confirmed by recalibration of the column with ribonuclease A and albumin.
Peroxide tolerance test of transgenic E.coli
Single colonies of E. coli BL21 carrying the pCR T7/NT-TOPO-TA vectors with inserted prx genes were transferred to 20 ml LB medium supplemented with 35 μg/ml chloramphenicol and 100 μg/ml ampicillin for over night culture. 100 μl suspension was transferred to new 20 ml LB with 100 μg/ml ampicillin and grown for 4.5 h. The expression of prx genes was induced by adding isopropyl-β-D-thiogalactopyranoside at 10 μmol/L final concentration for 30 min. This regime had been optimised in series of test experiments with variable IPTG concentrations and induction times in order to get significant expression without loss of bacterial cell viability. The control E. coli cells contained the pCR T7/NT-TOPO-TA vector with a short synthetic 27 bp insertion, namely GGAATTGCTCTGCAAGGATTATTCATC. The control cells were treated like the prx-expressing strains. Cells were collected by centrifugation (12,000×g for 5 min), washed once with 10 mM MgCl2, and resuspended in 10 mM MgCl2 at a titre of 2 × 107 colony-forming units (cfu) mL−1. Bacteria (106 cfu/mL) were subsequently incubated at 37°C for 30 min in 10 mL potassium phosphate buffer (50 mmol/L, pH 7.4) in the presence or absence of 0.1 mmol/L cumene hydroperoxide and morpholinosydnonimine (SIN-1) as organic peroxide and peroxinitrite donor, respectively. After 30 min incubation, the cell suspension was diluted 1:10- and 1:100-fold, respectively, in 10 mmol/L MgCl2. In triplicates, 10 μl were spread on 1.5% (w/v) LB agar plates, grown over night at 37°C, and number of surviving colonies was counted.
Statistical tests
Results from activity determinations were evaluated by Fisher LSD-test with one-way ANOVA (Statistica 6.0) with if not indicated otherwise p < 0.2. Significance of difference was calculated for the resistance test using Student’s t test.
Results and discussion
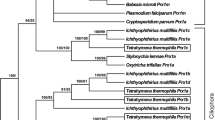
From the six Prx proteins encoded in the S. elongatus PCC 7942 genome, the 1- and 2-Cys Prx are highly similar to their Arabidopsis thaliana orthologues with 68% and 83% (Stork et al. 2005). The four PrxQ isoforms, denominated A1-3 and B, have similarities of 66, 58, 71 and 54% to At-PrxQ, respectively. Figure 1 depicts the aa sequence alignment of the four Se-PrxQ, the four PrxQ identified in the Anabaena sp. PCC 7120 genome, the two PrxQ sequences from Synechocystis PCC 6803 and the single Q-type Prx At-PrxQ and Os-PrxQ present in A. thaliana and rice. SePrxQ-B lacks the resolving Cys and is the protein most dissimilar to the cognate At-PrxQ. Like Se-PrxQ, also PrxQ-A and PrxQ-D from Anabaena PCC7120 and PrxQ-B1 and PrxQ-B2 from Synechococcus PCC6803 each of which lack the resolving Cys residue.
PrxQ amino acid sequence alignment. The alignment was constructed with the PrxQ sequences from S. elongatus PCC 7942 (PrxQ-A1, -A2, -A3 and B), Synechocystis sp. PCC 6803 (PrxQ-B1 and -B2) (Stork et al. 2005), Anabaena sp. PCC 7120 (PrxQ-A, -B, -C and -D) (Latifi et al. 2007), Arabidopsis thaliana (PrxQ) and Oryza sativa (PrxQ“217”). Cys residues are in black, amino acid residues conserved in at least six sequences are in grey. Asterisks denote fully conserved, colon denotes conserved and dot denotes partly conserved amino acid positions. The chloroplast PrxQ-sequence of A. thaliana was included in the comparison without the transit sequence
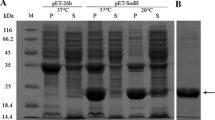
Since none of the S. elongatus PCC 7942 Prx proteins has been characterised up to now, a comprehensive biochemical analysis was sought with recombinant proteins. The coding regions of the 1-cys prx, 2-cys prx and the four prxQ-A1, -A2, -A3 and -B1 from S. elongatus PCC 7942 were fused in frame to the 6xHis-extension of the pCR T7/NT-TOPO-vector. Recombinant proteins of the Se-Prxs and Ec-TrxA were purified by affinity chromatography on Ni-NTA material, and highly enriched fractions were obtained (Fig. 2).
Heterologous expression of the six S. elongatus PCC 7942 Prx proteins and the E. coli TrxA protein. Protein expression was induced by addition of IPTG to E. coli lines harbouring the Prx-constructs. Cells were lysed and the Prx proteins and the TrxA protein enriched by Ni-NTA affinity chromatography. The obtained proteins corresponded to the expected size. The complete purification is represented. M 4 μl Seablue protein marker, 0, 2, 4 denote the time in hours elapsed after IPTG induction, S supernatant, which was passed through the Ni-NTA column, P pellet-containing particulate cell components like membrane fragments after centrifugation and separation of the protein-containing supernatant for the column passage, E washing eluate from Ni-NTA-column, last lane purified protein after dialysis. The gels were stained with Coomassie brilliant blue
Peroxide reduction assays were conducted to characterise the enzymatic properties of the six Prx (Table 2). To this end two tests were performed with hydrogen peroxide at 100 μmol/L concentration as substrate. In the first test, 10 mmol/L dithiothreitol as artificial electron donor was added as immediate regenerant. Peroxidase activities of five Se-Prx ranged between 1 and 7.8 μmol H2O2decomposed/(μmol Prx min), while PrxQ-A2 revealed no detectable activity (not shown). To check for thioredoxin-dependent regeneration, the DTT-containing assay was supplemented with E. coli thioredoxin TrxA. Prx-A2 showed no activity with or without TrxA, as with any other tested potential regenerant such as plant glutaredoxins (At-Grx as in Finkemeier et al. 2005, not shown). The peroxidase activity of the 1-Cys Prx was unchanged with TrxA. All other Prx showed significant activity stimulation after addition of the TrxA protein despite the rather high concentration of 10 mmol DTT/L (Table 2). The least increase was seen with PrxQ-B with 1.3-fold stimulation, while a 5.5-fold stimulation was measured for PrxQ-A3, a 7.1-fold one for PrxQ-A1 and an 8.7-fold one for 2-Cys Prx, showing efficient regeneration by the heterologous E. coli thioredoxin.
Three different peroxide substrates, i.e. hydrogen peroxide, butylhydroperoxide (tBOOH) and cumene hydroperoxide (COOH), were compared in the TrxA-dependent assay, except for the case of 1-Cys Prx where TrxA was omitted since TrxA had failed to stimulate the activity (Fig. 3). The 1-Cys Prx detoxified H2O2, tBOOH and COOH with similar efficiencies. All three PrxQ-A revealed the lowest activity with tBOOH, but high efficiencies with COOH and equal or slightly lower activity with H2O2. S. elongatus 2-Cys Prx revealed a significant substrate preference for H2O2 relative to tBOOH and COOH, which exceeded that of plant 2-Cys Prx (König et al. 2003) but resembled that, e.g. of the malarial parasite Plasmodium falciparum (Rahlfs and Becker 2001).
Peroxidase activity of S. elongatus PCC 7942 Prx proteins. Activity measurements were performed in a colorimetric test with 1-Cys Prx, 2-Cys Prx, PrxQ-A1, -A3, -B and 2-Cys Prx, while PrxQ-A2 had no activity. Peroxide substrates were added at 100 μmol/L concentrations, DTT at a concentration of 10 mmol/L and TrxA at 5 μmol/L. The data are means of n = 28–45 determinations, ±SE. Different letters (a, b) indicate significance of difference at p < 0.2. In addition, the NADPH-linked peroxidase assay was performed with thioredoxin reductase, TrxA and 2-Cys Prx. H2O2 was added at concentrations between 10 and 100 μmol/L. Time dependent absorption changes were monitored at 340 nm
In addition an NADPH-linked peroxidase activity test was performed, where thioredoxin reductase, TrxA were coupled to H2O2 detoxification. Reliable results were obtained with 2-Cys Prx, PrxQ-A1 and PrxQ-A3. Figure 3 shows the data for 2-Cys Prx. The initial slope of the NADPH oxidation curve was highly similar irrespective of the H2O2 concentration. The reaction proceeded with same speed until the substrate was almost completely exhausted. The experiments allowed us to derive the kinetic constants for these three Prx (Table 3). Concentration dependency revealed KM values between 2 and 4.1 μmol/L, turnover numbers between 4.2 and 14.9 and values for the catalytic efficiency that decreased from 2-Cys Prx to PrxQ-A1 and PrxQ-A3. The turnover numbers of PrxQ-A1 and PrxQ3 were similar for the colorimetric and NADPH-dependent test. The lower activity of 2-Cys Prx in the NADPH-test might be caused by the lower TrxA concentration of 3 μmol/L as compared to 5 μmol/L in the colorimetric test and may indicate a regeneration limitation. The very low stimulation by TrxA of the monocysteinic Se-PrxQ-B had to be expected since TrxA most efficiently catalyses two electron transfer reactions. Glutaredoxins, glutathione-S-transferases in conjunction with GSH may serve as electron donors as in the case of 1-Cys Prx (Manevich et al. 2004; Noguera-Mazon et al. 2006).
Prx function as monomers, dimers or oligomers that partly undergo dynamic transitions depending on the redox state. To characterise the protein family of S. elongatus PCC 7942, heterologously expressed proteins in reduced and oxidised state from each group, respectively, were separated by size exclusion chromatography (Fig. 4). 1-Cys Prx separated as dimer of 42 kDa, 2-Cys Prx as oligomer of ≥200 kDa and PrxQ-A1 as monomer of 20 kDa independent on the redox state. It should be noted that chaperones eventually present in Synechococcus but absent in E. coli during heterologous expression might affect the correct folding and aggregation. The unexpected stability of the cyanobacterial oxidised 2-Cys Prx is partly in line with results from a recent isothermal titration microcalorimetric study showing a critical assembly concentration for reduced human and plant 2-Cys Prx, that was not seen for the 2-Cys Prx from S. elongatus 2-Cys Prx (Barranco-Medina et al. 2008).
Aggregation state of S. elongatus PCC 7942 1-Cys Prx, 2-Cys Prx and PrxQ-A1. Recombinant Prx proteins in reduced (10 mM DTT for 20 min) or oxidised state (0.5 mM H2O2 for 15 min, subsequent treatment with 2.5 mM ascorbate for 5 min) were separated by size exclusion chromatography on Superose 12 column. Elution of protein was monitored at 280 nm. Ascorbate was detected as a peak between 20–25 ml elution volume. Peaks representing Prx protein are labelled with the apparent molecular masses in kDa. The given curves are representative of 2–5 different independent fractionations
E. coli expresses two cytoplasmic peroxidoxins, the 2-Cys Prx AhpC (P0AE08) (Michan et al. 1999) and the PrxQ-like Bcp (P0AE52) (Jeong et al. 2000). In addition, a thioredoxin-dependent peroxidase is targeted to the periplasmic space (Cha et al. 1995). In the background of these endogenous Prx, the protective function of the Se-Prx isoforms was tested in vivo by assessing the viability of E. coli cells expressing the various cyanobacterial Prx in the presence of cumene hydroperoxide and peroxinitrite (Fig. 5). These peroxides were chosen based on the established toxicity in E. coli and the reported protective effect, e.g. of overexpressed plant 2-Cys Prx, e.g. against cumene hydroperoxide (Baier and Dietz 1997). Peroxinitrite has been investigated by Sakamoto et al. (2003) and Bryk et al. (2000). In initial experiments, the IPTG concentration routinely used to induce rapid and strong accumulation of recombinant protein was shown to drastically impair cell viability. To optimise the test system, the minimum IPTG concentration was searched and subsequently applied to induce significant synthesis of the respective Prx protein but to avoid detrimental overproduction. Figure 5 exemplarily shows a basal expression of 2-Cys Prx protein in vector-containing cells even in the absence of IPTG. IPTG at 10 μmol/L concentration increased the amount of recombinant protein but did not affect cell viability (not shown). Application for 30 min of 100 μmol/L COOH resulted in 50–60% cell survival. Expression of 1-Cys Prx, 2-Cys Prx and PrxQ-B did not affect cell sensitivity towards COOH. In a converse manner expression of PrxQ-A1, −2 and −3 strongly increased cell survival almost to the level of the untreated control. 2-Cys Prx and PrxQ-B showed a trend towards protection of the cells against peroxinitrite; however, the difference to the treated control was statistically not significant.
Western Blot detection of 2-Cys Prx in E. coli lysates with and without IPTG (A) and in vivo protection of E. coli expressing Se-Prx against peroxynitrite (ONOO−) and cumene hydrogen peroxide (COOH). a Transgenic E. coli BL21 cells with and without 2-cys prx gene-carrying expression vector were grown for 4 h 30 min and then incubated with or without IPTG for 30 min. After washing and centrifugation lysates were obtained, which were separated after protein amount determination on SDS-PAGE. Detection of 2-Cys Prx was performed after blotting on nitrocellulose membrane with anti His antibody. As control 100 ng purified recombinant 2-Cys Prx protein was separated on the gel. b E. coli cells with T7/NT-TOPO-TA vector-containing one of each prx genes of S. elongatus PCC 7942 or vector with a small insertion (control) were grown for 4 h 30 min, incubated for 30 min with 10 μM IPTG, washed and diluted in 40 mM KPi buffer pH 7.4 up to 1 × 106 cells. After 30 min treatment with 100 μM COOH (grey bars), 1 mM SIN1 (open bars) or water (control, black bars) dilutions of the cell cultures were spread on agar plates, incubated at 37°C over night, colonies counted and survival relative to control calculated. Mean values ± SE were as follows: 1-Cys Prx, PrxQ-A2, PrxQ-A3 and PrxQ-B: n = 6 from two independent experiments; 2-Cys Prx and PrxQ-A1: n = 9 from three independent experiments; control: n = 42. Significant differences (p < 0.01) were calculated in the case of COOH in comparison to the control and indicated by different letters (a, b)
Conclusions
Reverse genetic approaches using insertional inactivation of individual prx genes have implicated Prx in antioxidant defence of bacteria (Seaver and Imlay 2001; Wang et al. 2006), animals (Rhee et al. 2005) and plants (Baier and Dietz 1999; Finkemeier et al. 2005; Lamkemeyer et al. 2006). In Synechocystis PCC 6803, both deletion of the 1-cys prx (slr1198) and the type II prx (sll1621) impaired cell survival (Hosoya-Matsuda et al. 2005). S. elongatus PCC 7942 lacks a type II Prx. This is interesting, since plant chloroplasts generally appear to contain a PrxII E (Horling et al. 2003). PrxII E is soluble in the stroma and was suggested to detoxify peroxides escaping from the thylakoids, generated in the stroma or diffusing from the cytosol into the chloroplast (Dietz et al. 2006). It is tempting to hypothesise that the lack of type II Prx is compensated by PrxQ in S. elongatus. The three Se-PrxQ-A proteins contain the reactive centre typical for PrxQ with two Cys residues spaced by four intervening amino acids. Se-PrxQ-A1 and -A3 showed significant activity stimulation in the presence of TrxA. Se-PrxQ-A2 revealed no activity under the various assay conditions tested, i.e. with DTT, DTT/TrxA, NADPH/thioredoxin reductase/TrxA, GSH and GSH/Grx. The reason for this inactivity is unknown. Inefficient regeneration is the most likely explanation. Surprisingly heterologous expression of all Se-PrxQ-A increased the tolerance of E. coli towards cumene hydroperoxide, showing that the required regeneration systems are present in E. coli cells. Comparing the gene family composition of Synechocystis PCC 6803 and S. elongatus PCC 7942 and Prx activities a functional substitution of PrxQ-A for the type II Prx present in some cyanobacteria may be assumed. This is in line with a strong transcript accumulation of the Synechocystis type II prx upon exposure to various stresses, e.g. in response to high light and H2O2 treatment. A similar pattern is seen for prxQ-A, particularly the A1-form in S. elongatus (Stork et al. 2005). The results show the structural and functional differentiation within the Prx family in S. elongatus. Future work should address the subcellular localisation and the processes affected by disturbed Prx function in single and multiple mutants in order to arrive at the next level of mechanistic understanding of the large Prx family in Cyanobaceteria.
Abbreviations
- COOH:
-
Cumene hydroperoxide
- DTT:
-
Dithiothreitol
- Grx:
-
Glutaredoxin
- GSH:
-
Glutathione
- IPTG:
-
Isopropyl-β-D-thiogalactopyranoside
- Prx:
-
Peroxiredoxin
- tBOOH:
-
Tertiary butylhydroperoxide
- TrxA:
-
Thioredoxin
References
Baier M, Dietz KJ (1996) Primary structure and expression of plant homologues of animal and fungal thioredoxin-dependent peroxide reductases and bacterial alkyl hydroperoxide reductases. Plant Mol Biol 31:553–564
Baier M, Dietz KJ (1997) The plant 2-Cys peroxiredoxin BAS1 is a nuclear-encoded chloroplast protein: its expressional regulation, phylogenetic origin, and implications for its specific physiological function in plants. Plant J 12:179–190
Baier M, Dietz KJ (1999) Protective function of chloroplast 2-cysteine peroxiredoxin in photosynthesis. Evidence from transgenic Arabidopsis. Plant Physiol 119:1407–1414
Barranco-Medina S, Kakorin S, Lazaro JJ, Dietz KJ (2008) Thermodynamics of the dimer-decamer transition of reduced human and plant 2-Cys peroxiredoxin. Biochem 47:7196–7204
Bréhélin C, Meyer EH, de Souris JP, Bonnard G, Meyer Y (2003) Resemblance and dissemblance of Arabidopsis type II peroxiredoxins: similar sequences for divergent gene expression, protein localization, and activity. Plant Physiol 132:2045–2057
Bryk R, Griffin P, Nathan C (2000) Peroxynitrite reductase activity of bacterial peroxiredoxins. Nature 407:211–215
Cha MK, Kim HK, Kim IH (1995) Thioredoxin-linked thiol peroxidase from periplasmic space of Escherichia coli. J Biol Chem 270:28635–28641
Chae HZ, Kim IH, Kim K, Rhee SG (1993) Cloning, sequencing and mutation of thiol-specific antioxidant gene of Saccharomyces cerevisiae. J Biol Chem 268:16815–16821
Collin V, Issakidis-Bourguet E, Marchand C, Hirasawa M, Lancelin JM, Knaff DB, Miginiac-Maslow M (2003) The Arabidopsis plastidial thioredoxins. J Biol Chem 278:23747–23752
Dietz KJ (2003) Plant peroxiredoxins. Annu Rev Plant Biol 54:93–107
Dietz KJ, Stork T, Finkemeier I, Lamkemeyer P, Li WX, El-Tayeb MA, Michel KP, Pistorius EK, Baier M (2005) The role of peroxiredoxins in oxygenic photosynthesis of cyanobacteria and higher plants: peroxide detoxification or redox sensing? In: Demmig-Adams B, Adams WWIII, Mattoo AK (eds) Photoprotection, photoinhibition, gene regulation, and environment. Springer, Dordrecht, pp 303–319
Dietz KJ, Jacob S, Oelze ML, Laxa M, Tognetti V, de Miranda SMN, Baier M, Finkemeier I (2006) The function of peroxiredoxins in plant organelle redox metabolism. J Exp Bot 57:1697–1709
Elstner EF (1990) Der Sauerstoff. Biochemie, Biologie Medizin. Wissenschaftsverlag, Mannheim
Engemann J (2006) Charakterisierung des Peroxiredoxins Bcp von Clavibacter michiganensis subsp. michiganensis. PhD thesis (http://bieson.ub.uni-bielefeld.de/volltexte/2006/896/pdf/Dissertation.pdf)
Finkemeier I, Goodman M, Lamkemeyer P, Kandlbinder A, Sweetlove LJ, Dietz KJ (2005) The mitochondrial type II peroxiredoxin F is essential for redox homeostasis and root growth of Arabidopsis thaliana under stress. J Biol Chem 280:12168–12180
Hansmann S, Martin W (2000) Phylogeny of 33 ribosomal and six other proteins encoded in an ancient gene cluster that is conserved across prokaryotic genomes: influence of excluding poorly alignable sites from analysis. Int J Syst Evol Microbiol 50:1655–1663
Hofmann B, Hecht HJ, Flohé L (2002) Peroxiredoxins. Biol Chem 383:347–464
Horling F, König J, Dietz KJ (2002) Type II peroxiredoxin C, a member of the peroxiredoxin family of Arabidopsis thaliana: its expression and activity in comparison with other peroxiredoxins. Plant Physiol Biochem 40:491–499
Horling F, Lamkemeyer P, König J, Finkemeier I, Kandlbinder A, Baier M, Dietz KJ (2003) Divergent light-, ascorbate-, and oxidative stress-dependent regulation of expression of the peroxiredoxin gene family in Arabidopsis. Plant Physiol 131:317–325
Hosoya-Matsuda N, Motohashi K, Yoshimura H, Nozaki A, Inoue K, Ohmori M, Hisabori T (2005) Anti-oxidative stress system in cyanobacteria—significance of type II peroxiredoxin and the role of 1-Cys peroxiredoxin in Synechocystis sp. strain PCC 6803. J Biol Chem 280:840–846
Jeong W, Cha MK, Kim IH (2000) Thioredoxin-dependent hydroperoxide peroxidase activity of bacterioferritin comigratory protein (BCP) as a new member of the thiol-specific antioxidant protein (TSA)/alkyl hydroperoxide peroxidase C (AhpC) family. J Biol Chem 275:2924–2930
Kaneko T, Sato S, Kotani H et al (1996) Sequence analysis of the genome of the unicellular cyanobacterium Synechocystis sp. strain PCC 6803. II. Sequence determination of the entire genome and assignment of potential protein-coding regions (supplement). DNA Res 3:185–209
Kim K, Kim IH, Lee KY, Rhee SG, Stadtman ER (1988) The isolation and purification of a specific “protector” protein which inhibits enzyme inactivation by a thiol/Fe(III)/O2 mixed-function oxidation system. J Biol Chem 263:4704–4711
Kobayashi M, Ishizuka T, Katayama M, Kanehisa M, Bhattacharyya-Pakrasi M, Pakrasi HB, Ikeuchi M (2004) Response to oxidative stress involves a novel peroxiredoxin gene in the unicellular cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol 45:290–299
Koksharova OA, Klint J, Rasmussen U (2006) The first protein map of Synechococcus sp. strain PCC 7942. Microbiol 75:664–672
König J, Lotte K, Plessow R, Brockhinke A, Baier M, Dietz KJ (2003) Reaction mechanism of the 2-Cys peroxiredoxins. Role of the C terminus and the quaternary structure. J Biol Chem 278:24409–24420
Lamkemeyer P, Laxa M, Collin V, Li WX, Finkemeier I, Schöttler MA, Holtkamp V, Tognetti VB, Issakidis-Bourguet E, Kandlbinder A, Weis E, Miginiac-Maslow M, Dietz KJ (2006) Peroxiredoxin Q of Arabidopsis thaliana is attached to the thylakoids and functions in context of photosynthesis. Plant J 45:968–981
Latifi A, Ruiz M, Jeanjean R, Zhang CC (2007) PrxQ-A, a member of the peroxiredoxin Q family, plays a major role in defense against oxidative stress in the cyanobacterium Anabaena sp. strain PCC 7120. Free Radic Biol Med 42:424–431
Laxa M, König J, Dietz KJ, Kandlbinder A (2007) Role of the cysteine residues in Arabidopsis thaliana cyclophilin CYP20-3 in peptidyl-prolyl cis-trans isomerase and redox-related functions. Biochem J 401:287–297
Manevich Y, Feinstein SI, Fisher AB (2004) Activation of the antioxidant enzyme 1-CYS peroxiredoxin requires glutathionylation mediated by heterodimerization with pi GST. Proc Natl Acad Sci USA 101:3780–3785
Michan C, Manchado M, Dorado G, Pueyo C (1999) In vivo transcription of the Escherichia coli oxyR regulon as a function of growth phase and in response to oxidative stress. J Bacteriol 181:2759–2764
Noguera-Mazon V, Lemoine J, Walker O, Rouhier N, Salvador A, Jacquot JP, Lancelin JM, Krimm I (2006) Glutathionylation induces the dissociation of 1-Cys D-peroxiredoxin non-covalent homodimer. J Biol Chem 281:31736–31742
Olson JM, Blankenship RE (2004) Thinking about the evolution of photosynthesis. Photosynth Res 80:373–386
Perelman A, Uzan A, Hacohen D, Schwarz R (2003) Oxidative stress in Synechococcus sp strain PCC 7942: various mechanisms for H2O2 detoxification with different physiological roles. J Bacteriol 185:3654–3660
Petersson UA, Kieselbach T, García-Cerdán JG, Schröder WP (2006) The Prx Q protein of Arabidopsis thaliana is a member of the luminal chloroplast proteome. FEBS Lett 580:6055–6061
Rahlfs S, Becker K (2001) Thioredoxin peroxidases of the malarial parasite Plasmodium falciparum. Eur J Biochem 268:1404–1409
Reddy KJ, Webb R, Sherman LA (1990) Bacterial RNA isolation with one hour centrifugation in a table-top ultracentrifuge. BioTechniques 8:250–251
Rhee SG, Chae HZ, Kim K (2005) Peroxiredoxins: a historical overview and speculative preview of novel mechanisms and emerging concepts in cell signaling. Free Radic Biol Med 38:1543–1552
Rouhier N, Gelhaye E, Sautiere PE, Brun A, Laurent P, Tagu D, Gerard J, de Fay E, Meyer Y, Jacquot JP (2001) Isolation and characterization of a new peroxiredoxin from poplar sieve tubes that uses either glutaredoxin or thioredoxin as a proton donor. Plant Physiol 127:1299–1309
Rouhier R, Gelhaye E, Gualberto JM, Jordy MN, De Fay E, Hirasawa M, Duplessis S, Lemaire SD, Frey P, Martin F, Manieri W, Knaff DB, Jacquot JP (2004) Poplar peroxiredoxin Q. A thioredoxin-linked chloroplast antioxidant functional in pathogen defence. Plant Physiol 134:1027–1038
Sakamoto A, Tsukamoto S, Yamamoto H, Ueda-Hashimoto M, Takahashi M, Suzuki H, Morikawa H (2003) Functional complementation in yeast reveals a protective role of chloroplast 2-Cys peroxiredoxin against reactive nitrogen species. Plant J 33:841–851
Schopf JW (1993) Microfossils of the early archean apex chert: new evidence of the antiquity of life. Science 260:640–646
Seaver LC, Imlay JA (2001) Alkyl hydroperoxide reductase is the primary scavenger of endogenous hydrogen peroxide in Escherichia coli. J Bacteriol 183:7173–7181
Shestakov SV, Khyen NT (1970) Evidence for genetic transformation in blue-green alga Anacystis nidulans. Mol Gen Genet 107:372–375
Stacy RAP, Munthe E, Steinum T, Sharma B, Aalen RB (1996) A peroxiredoxin antioxidant is encoded by a dormancy-related gene, Per1, expressed during late development in the aleuron and embryo of barley grains. Plant Mol Biol 31:1205–1216
Stacy RAP, Nordeng TW, Culianez-Macia FA, Aalen RB (1999) The dormancy-related peroxiredoxin anti-oxidant, PER1, is localized to the nucleus of barley embryo and aleurone cells. Plant J 19:1–8
Stephan DP, Ruppel HG, Pistorius EK (2000) Interrelation between cyanophycin synthesis, L-arginine catabolism and photosynthesis in the cyanobacterium Synechocystis sp. strain PCC 6803. Z Naturforsch 55:927–942
Stork T, Michel KP, Pistorius EK, Dietz KJ (2005) Bioinformatic analysis of the genomes of the cyanobacteria Synechocystis sp. PCC 6803 and Synechococcus elongatus PCC 7942 for the presence of peroxiredoxins and their transcript regulation under stress. J Exp Bot 56:3193–3206
Wang G, Hong Y, Johnson MK, Maier RJ (2006) Lipid peroxidation as a source of oxidative damage in Helicobacter pylori: protective roles of peroxiredoxins. Biochim Biophys Acta 1760:1596–1603
Xiong J, Fischer WM, Inoue K, Nakahara M, Bauer CE (2000) Molecular evidence for the early evolution of photosynthesis. Science 289:1724–1730
Yamamoto H, Miyake C, Dietz KJ, Tomizawa KI, Murata N, Yokota A (1999) Thioredoxin peroxidase in the cyanobacterium Synechocystis sp. PCC 6803. FEBS Lett 447:269–273
Acknowledgments
The authors gratefully acknowledge support by the DFG (SFB 613, Di 346).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Jack Meeks.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Stork, T., Laxa, M., Dietz, M.S. et al. Functional characterisation of the peroxiredoxin gene family members of Synechococcus elongatus PCC 7942. Arch Microbiol 191, 141–151 (2009). https://doi.org/10.1007/s00203-008-0438-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-008-0438-7