Abstract
Aims/hypothesis
Insufficient insulin secretion and reduced pancreatic beta cell mass are hallmarks of type 2 diabetes. Here, we focused on a family of serine-threonine kinases known as homeodomain-interacting protein kinases (HIPKs). HIPKs are implicated in the modulation of Wnt signalling, which plays a crucial role in transcriptional activity, and in pancreas development and maintenance. The aim of the present study was to characterise the role of HIPKs in glucose metabolism.
Methods
We used RNA interference to characterise the role of HIPKs in regulating insulin secretion and transcription activity. We conducted RT-PCR and western blot analyses to analyse the expression and abundance of HIPK genes and proteins in the islets of high-fat diet-fed mice. Glucose-induced insulin secretion and beta cell proliferation were measured in islets from Hipk3 −/− mice, which have impaired glucose tolerance owing to an insulin secretion deficiency. The abundance of pancreatic duodenal homeobox (PDX)-1 and glycogen synthase kinase (GSK)-3β phosphorylation in Hipk3 −/− islets was determined by immunohistology and western blot analyses.
Results
We found that HIPKs regulate insulin secretion and transcription activity. Hipk3 expression was most significantly increased in the islets of high-fat diet-fed mice. Furthermore, glucose-induced insulin secretion and beta cell proliferation were decreased in the islets of Hipk3 −/− mice. Levels of PDX1 and GSK-3β phosphorylation were significantly decreased in Hipk3 −/− islets.
Conclusions/interpretation
Depletion of HIPK3 impairs insulin secretion and glucose tolerance. Decreased levels of HIPK3 may play a substantial role in the pathogenesis of type 2 diabetes.
Similar content being viewed by others
Introduction
Type 2 diabetes is a complex disease characterised by chronic hyperglycaemia resulting from insulin resistance and impaired beta cell function [1–3]. Accumulating evidence shows that Wnt signalling modulates beta cell function [4–9]. The Wnts are a family of secreted glycoproteins that influence cell development via autocrine and paracrine mechanisms [10, 11]. Activation of the canonical pathway occurs by binding of Wnt ligands to the frizzled receptor. This binding triggers an intracellular signalling cascade, which leads to serine/threonine phosphorylation, inactivation of glycogen synthase kinase (GSK)-3β and destabilisation of the destruction complex, preventing GSK3β phosphorylation of β-catenin. This mechanism enables the accumulation and nuclear translocation of β-catenin. Once inside the nucleus, β-catenin acts in combination with the T cell factor to stimulate transcription of Wnt-responsive genes.
GSK-3 was originally identified as a serine/threonine kinase that inactivates glycogen synthase. GSK3 affects many cellular processes, including transcription, cell cycle regulation and apoptosis. GSK3β has been shown to affect beta cell mass, proliferation and apoptosis [12–14].
Homeodomain-interacting protein kinases (HIPKs) are implicated in the modulation of Wnt signalling in cultured cells [15], and in mouse [16, 17], Drosophila [18, 19] and Xenopus embryos [20, 21]. Members of the HIPK family (HIPK1, HIPK2, HIPK3, HIPK4) were originally identified as binding partners of the neurokinin (NK) homeodomain protein [22]. HIPKs have been reported to be essential for coordinated death in early developmental stages and for the regulation of proper cell number in diverse tissue types. Studies have shown that the HIPK family, and in particular its most studied member, HIPK2, interact with, phosphorylate and modulate the function of other homeodomain proteins, as well as other transcription factors, including p53 [23–25] and carboxy-terminal binding protein (CtBP)-1 [26, 27]. HIPK2 has been reported to modulate the transcriptional activity of pancreatic duodenal homeobox (PDX)-1 [28], which plays a crucial role in pancreas development [29–31].
Several studies have revealed important roles of HIPK3 in the regulation of transcription and phosphorylation in cultured cells [32–36]. HIPK3 overproduction has been reported to enhance androgen receptor-dependent transcription in various cell lines [32]. HIPK3 induced Fas-associated death domain (FADD) phosphorylation and inhibited Fas-mediated Jun NH2-terminal kinase activation [33]. HIPK3 also increased splicing factor 1 (SF-1) activity for steroidogenic gene transcription in response to cAMP through phosphorylation of death-associated protein 6 [35]. Interestingly, HIPK1, HIPK2 and HIPK3 expression in islets from patients with type 2 diabetes is decreased by 32% (p = 0.326), 30% (p = 0.082) and 46% (p = 0.064), respectively, according to data from the Diabetes Genomic Anatomic Project [3], suggesting that HIPK genes play a role in the control of beta cell function. However, the question of whether HIPKs might be involved in the effects of glucose metabolism has not been addressed.
The current study was undertaken to define the mechanisms by which HIPKs regulate glucose metabolism. Here we show that Hipk transcripts are expressed in pancreatic beta cells, and modulate insulin secretion and insulin promoter activity. We found that a high-fat diet enhanced Hipk3 expression. Moreover, our analysis of Hipk3 −/− mice revealed that they had impaired glucose tolerance due to a deficiency of insulin secretion. We also demonstrated that levels of PDX1 and phosphorylation of GSK3β were significantly decreased in Hipk3 −/− islets. Hence, our data provide evidence of a new mechanism by which insulin secretion is regulated.
Methods
RT-PCR
Total RNA was isolated from mouse islets and mouse pancreas using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). RT-PCR was performed using reverse transcriptase (RevTraAce; Toyobo, Osaka, Japan). The RT-PCR products were amplified using ExTaq polymerase (Takara, Kyoto, Japan). The gene-specific primer pairs were used, as previously described [37].
Quantitative RT-PCR
Quantification of the selected genes was performed using TaqMan gene expression assays with a commercially available system (7900HT; Applied Biosystems, Foster City, CA, USA). For each gene, an assay was selected to amplify the region corresponding to the location of the relevant probe. Gene expression analysis was performed using commercially available mouse assays as follows: Hipk1: Mm00501689_m1; Hipk2: Mm00439329_m1; Hipk3: Mm00468880_m1; Pdx1: Mm00435565_m1; Gck: Mm00439129_m1; Slc2a2: Mm00446229_m1; Cyclin D1: Mm00432359_m1; Hnf4 (also known as Hnf4a): m00433964_m1; Tcf7l2: Mm00501505_m1; Irs2: Mm03038438_m1; Ins1: Mm01950294_s1; Ins2: Mm00731595_g1; Hnf1a: Mm00493434_m1; Hnf1b: Mm00447459_m1; Irs1: Mm01278327_m1; Ucp2: Mm00627599_m1; and Gapdh: Mm99999915_g1 (Applied Biosystems). The assays for Gapdh and Pdx1 were performed in parallel as controls. Standard TaqMan cycling conditions were used and all reactions were performed in triplicate. Changes in expression levels in quantitative PCR were calculated as \( {2^{{ - \Delta \Delta {C_t}}}} \) (cycle threshold) values and presented relative to average changes.
Immunoprecipitation and immunoblotting analysis of mouse islets
Polyclonal anti-PDX1, anti-hepatocyte nuclear factor 4 alpha, anti-GSK3β, anti-phospho-GSK3β(Ser9), anti-β-catenin and anti-α-tubulin antibodies were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Polyclonal anti-IRS2 antibodies were purchased from Cell Signaling Technology (Beverly, MA, USA). Islets were sonicated, using an ultrasound sonicator, in 300 to 500 μl ice-cold buffer containing: 0.5% (vol./vol.) Triton X-100, 50 mmol/l TRIS-HCl (pH 8.0), 100 mmol/l NaCl, 1 mmol/l EDTA, 1 mmol/l Na3VO4, 10 mmol/l NaF and 1 mmol/l phenylmethylsulfonyl fluoride. Supernatant fractions were cleared at 12,000 g for 5 min, and incubated with protein A-Sepharose and each specific antibody. Samples were separated by SDS-polyacrylamide gel electrophoresis and immunodetection performed with a kit (ECL; Amersham Bioscience, Arlington Heights, IL, USA). Protein was prepared from more than 100 islets pooled from several mice of identical genotype.
RNA interference and transient transfection
Small interfering RNA (siRNA)-Lipofectamine 2000 complexes were prepared using 50 nmol/l siRNA for Hipk genes (Stealth Select RNAi; Invitrogen) and control siRNA (Invitrogen). Islets were precultured for 24 h, medium was changed to OptiMEM (Invitrogen) and siRNA-Lipofectamine 2000 complexes with or without DNA-Lipofectamine 2000 complexes were added. After 8 h incubation, the transfection medium was aspirated and replaced, for an additional 48 h, with fresh culture medium containing 2.8 mmol/l glucose.
Luciferase assays
Insulin reporter constructs were constructed from rat pancreas cDNA and genomic DNA using Advantage2 Taq (Invitrogen). The luciferase reporter plasmid harbouring the 5′-flanking region of rat Ins1 exon 1 (−450 to 51) was subcloned into the pGL4 vector (Promega Biosciences, Madison, WI, USA). Islets plated on to a 24 well plate were transfected with 0.5 μg of each luciferase reporter plasmid and 0.02 μg Renilla luciferase plasmid with the herpes simplex virus thymidine kinase promoter (phRL-TK) using Lipofectamine 2000. Cells were collected 48 h after transfection and luciferase assays were carried out (Promega).
Chromatin immunoprecipitation assay
Assays were performed in mouse islets. A chromatin immunoprecipitation (ChIP) assay kit (Upstate, Lake Placid, NY, USA) was used according to procedures suggested by the manufacturer. After crossing and sonication, cell lysates were immunoprecipitated with the indicated antibodies. The eluted genomic DNA from immunoprecipitates was subjected to PCR amplification. Anti-HIPK1, anti-HIPK2, anti-HIPK3 and anti-PDX1 antibodies, as well as control rabbit IgG (Santa Cruz Biotechnology) were used in ChIP assays. For PCR primer sequences, see electronic supplementary material (ESM) Table 1.
Analysis of Hipk3 −/− mice
Mice deficient for Hipk3 were generated using the gene trapping method developed by Lexicon Genetics (Woodlands, TX, USA) and supplied by CLEA (Tokyo, Japan). The artificial DNA was designed such that it could be inserted randomly into any gene, preventing the RNA splicing mechanism from working properly and thereby knocking out the gene’s function. The inserted DNA was specifically engineered to disrupt the function of the trapped gene, activate a selectable marker, enable the generation of knockout mice and allow precise identification of the chromosomal insertion site (www.lexpharma.com/research/drug-discovery/gene-knockouts.html, last accessed 8 August 2012). Mice were 8 to 20 weeks of age at the time of the experiment. Male 8-week-old BKS. Cg-+Lepr db/+Lepr db/Jcl (db/db) mice and BKS. Cg-m+/m+/Jcl mice (control) were purchased (CLEA). We used a high-fat diet consisting of 32% fat, 25.5% (wt/wt) protein, 2.9% fibre, 4.0% ash, 29.4% carbohydrates and 6.2% water (CLEA). For all other experiments, the diet consisted of standard chow (CLEA) with the following composition: 25.6% (wt/wt) protein, 3.8% fibre, 6.9% ash, 50.5% carbohydrates, 4% fat and 9.2% water.
Mice were fasted for >16 h before the glucose tolerance test, then loaded with 1.5 mg/g body weight glucose by oral or intraperitoneal administration. Blood samples were collected from the orbital sinus at various time points and glucose measured using an automatic blood glucose meter (Glutest Ace; Sanwa Kagaku, Nagoya, Japan). Insulin and GLP-1 levels were determined using a mouse insulin ELISA kit (Shibayagi, Shibukawa, Japan) and GLP-1 ELISA kit (Wako, Osaka, Japan), respectively. Mice were intraperitoneally challenged with 0.75 mU/g body weight human insulin (Novolin R; Novo Nordisk, Bagsvaerd, Denmark) in an insulin tolerance test. Animal care and the procedures of the experiments were approved by the Animal Care Committee of the University of Tokyo.
Histological and immunohistochemical analysis of mouse islets
Female mice (n = 4) aged 8 to 20 weeks were used for each genotype and 20 sections of islets were evaluated for morphometry. We studied four animals each per genotype and diet. Analysis was based on cell counts ranging from 1,500 to 2,000 per control and transgenic mouse islet samples. Tissues were routinely processed for paraffin embedding, and 4 μm sections were cut and mounted on silanised slides. Pancreatic sections were stained with anti-insulin, anti-HIPK3 anti-proliferating cell nuclear antigen (PCNA), anti-PDX1 and anti-phspho-GSK3β antibodies. Non-specific staining was blocked by 2 mg/ml poly-l-lysine pre-absorption [38]. Images of pancreatic tissue and islet beta cells were viewed on the monitor of a computer through a microscope connected to a charge-coupled device camera (Olympus, Tokyo, Japan). The areas of the pancreas, beta cells and non-beta cells were traced manually and analysed with Win ROOF software (Mitani, Fukui, Japan).
Islet isolation and culture
Mouse pancreas was incubated for 30 min at 37°C in 2.5 ml KRB buffer containing collagenase (Sigma-Aldrich, St Louis, MO, USA). The pancreas was dispersed by pipetting and washed twice with KRB buffer. Mouse recombinant Wnt3a was purchased from Wako. Glibenclamide, LiCl and 1-azakenpaullone were purchased form Sigma-Aldrich. Static incubation was performed for 1 h at 37°C with ten islets per tube after 20 min of preincubation. Insulin levels were determined with a mouse insulin ELISA kit (Shibayagi). For the perifusion experiment, ten islets of similar size were maintained in KRB buffer delivered using a peristaltic pump. Perifusion was as follows: 30 min with KRB buffer containing 2.8 mmol/l glucose; 20 min with KRB buffer containing 20 mmol/l glucose; 40 min with KRB buffer containing 2.8 mmol/l glucose; 10 min with KRB buffer containing 2.8 mmol/l KCl; and 30 min with KRB buffer containing 2.8 mmol/l glucose. Perifusate was collected every 5 min. At the end of the perifusion, islets were collected by handpicking and extracted with 0.18 mol/l HCl in 70% ethanol for determination of insulin content.
ATP measurement
ATP measurement of islets was performed as previously described with some modifications [39]. After preincubation, 1 ml fresh KRB buffer containing 2.8, 10 or 22 mmol/l glucose was introduced. Incubation was carried out for 1 h at 37°C and stopped by adding trichloroacetic acid. ATP content of the cells was assayed in duplicate by the luminometric method using an ATP bioluminescent assay kit (Sigma).
Two-photon excitation imaging
Two-photon excitation imaging of islets was performed as previously described [40]. Ca2+ measurements were performed using fura-2 as the Ca2+ indicator. Fura-2 post-stimulation fluorescence (F) was normalised to the resting fluorescence (F0). The increase of Ca2+ concentration was calculated as (F0 − F)/F0.
Statistical analysis
All data are presented as the mean ± SEM of independent replicates. All statistical analyses were performed by one-way ANOVA or χ2 analysis as indicated.
Results
The reduction of Hipk1, Hipk2 and Hipk3 expression by RNA interference attenuates insulin secretion
We assessed the expression of Hipk family genes by RT-PCR using cDNA prepared from isolated mouse islets. Hipk family expression was detected in islets (Fig. 1a). Quantitative RT-PCR using TaqMan gene expression assays revealed that the expression of Hipk2 and Hipk3 was two- and 2.5-fold higher, respectively, than that of Hipk1 (Fig. 1b,c). We estimated whether Hipk genes can also decrease insulin secretion from isolated islets. siRNA to Hipk1, Hipk2 and Hipk3 suppressed endogenous expression by 45%, 44% and 46%, respectively (Fig. 1d), leading upon stimulation with 20 mmol/l glucose to decreases in insulin secretion by 42%, 44% and 46%, respectively (Fig. 1d). We also found that simultaneous inhibition of Hipk1 and Hipk2, Hipk2 and Hipk3, and Hipk1 and Hipk3 inhibited insulin secretion by 52%, 54% and 56%, respectively (Fig. 1e). Simultaneous inhibition of Hipk1, Hipk2 and Hipk3 inhibited insulin secretion by 78% (Fig. 1e). These observations suggest that the additive inhibition of Hipk genes leads to impaired beta cell function. To evaluate whether Hipk genes also affect insulin promoter transcriptional activity, we conducted luciferase assays using RNA interference approaches. siRNA to Hipk1, Hipk2 and Hipk3 decreased insulin promoter transcriptional activity in isolated mouse islets by 38%, 44% and 52%, respectively (Fig. 1f). Endogenous insulin transcript levels were assessed by quantitative RT-PCR and correlated with the results obtained in luciferase reporter assays (Fig. 1g). ChIP assay revealed that HIPKs interacted with the insulin promoter (Fig. 1h).
(a) RT-PCR analysis of Hipk genes in islets isolated from a wild-type mouse. Quantitative RT-PCR using TaqMan gene expression assays was performed to validate the relative expression of Hipk genes. (b) Assays for Gapdh and (c) Pdx1 were performed in parallel as control. (d) Blots showing Hipk1, Hipk2 and Hipk3 inactivation by RNA interference (RNAi). siRNA-Lipofectamine 2000 complexes were prepared (n = 4), and suppression of glucose-stimulated insulin secretion by gene inactivation was quantified. White bars, control; black bars, Hipk1; light grey bars, Hipk2; dark grey bars, Hipk3. (e) Glucose-stimulated insulin secretion was decreased by a combination of Hipk1, Hipk2 and Hipk3 RNA interference treatment (n = 4). White bars, control; black bars, Hipk1 + Hipk2; mid-grey bars, Hipk2 + Hipk3; dark grey bars, Hipk1 + Hipk3; light grey bars, Hipk1 + Hipk2 + Hipk3. (f) Insulin promoter transcriptional activity was suppressed by Hipk1, Hipk2 and Hipk3 inactivation by RNA interference in mouse isolated islets. Islets were precultured for 24 h, and DNA-Lipofectamine 2000 complexes along with siRNA-Lipofectamine 2000 complexes were added (n = 4). Isolated islets were transfected with 0.1 μg of the rat insulin I promoter luciferase reporter plasmid pINS (−450 to 51)-pGL4 and by siRNA to Hipk1, Hipk2 and Hipk3, and control siRNA. Luciferase activity was measured 48 h after transfection. Key as above (d). (g) Endogenous insulin transcript levels were assessed by quantitative RT-PCR. Key as in (d). (h) The recruitment of HIPKs on the Ins1 and Ins2 promoter in mouse islets. ChIP assays were performed in islets using the indicated antibodies (Ab) and primer pairs. As controls, PCR reactions were performed with input DNA (lane 2), DNA immunoprecipitated by rabbit IgG (lane 3) and DNA that was immunoprecipitated in the absence of antibody (lane 1). Data represent the average results from three independent ChIP experiments. Data (a–g) are from three independent islet preparations and experiments, with measurements in duplicate, and are presented as mean ± SEM; *p < 0.05 and **p < 0.01 compared with control siRNA
Hipk family expression in the islets of mice fed a high-fat diet
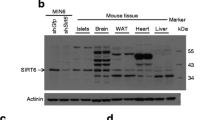
mRNA expression of Hipk1, Hipk2, and Hipk3 was increased in the extracts of islets isolated from B6 mice fed a high-fat diet for 12 weeks, compared with islets from control chow-fed B6 mice. Increases were by 44%, 78% and 109%, respectively (Fig. 2a). HIPK1, HIPK2 and HIPK3 levels were increased by 41%, 98% and 112%, respectively, in the extracts of islets isolated from B6 mice fed a high-fat diet (Fig. 2b), indicating that HIPK3 accumulation may correlate with glucose tolerance in this environmental model of type 2 diabetes.
Changes in Hipk genes mRNA expression (a) and protein (b) abundance in islets from mice fed a high-fat (black bars; white bars, control) diet (n = 4). Quantitative RT-PCR analysis of Hipk3 was done in islets isolated from wild-type mice. Protein was prepared from more than 100 islets pooled from several mice. Protein samples from lysates of isolated islets were immunoprecipitated using anti-HIPK1, -HIPK2 and -HIPK3 antibodies, and separated by SDS-PAGE. Western blot analyses were performed with the same antibodies. (c) Changes to Hipk3 mRNA expression and (d) protein levels in islets from 8-week-old (8W) and 20-week-old (20W) db/db mice (n = 4). RT-PCR analysis of Hipk3 was done in islets isolated from wild-type mice. Samples were normalised using α-tubulin. Protein samples from lysates of isolated islets were separated by SDS-PAGE and western blot analyses performed using anti-HIPK3 antibody. (e) Immunohistochemical analysis of paraffin-embedded sections from 8- and 20-week-old db/db mice using anti-insulin and anti-HIPK3 antibodies. Scale bars, 100 μm. (f) OGTT results for mice fed a high-fat diet (continuous black line; dotted line, control diet) and (g) for db/db mice (dotted line, control; thick black line, db/db 8 weeks old; thin black line, db/db 20 weeks old). Data represent means ± SEM; *p < 0.05 and **p < 0.01 compared with control
Hipk3 expression is enhanced in the islets of db/db mice
We examined Hipk3 expression in db/db mice. Hipk3 mRNA was increased by 96% and 54%, respectively, in the extracts of islets isolated from 8- and 20-week-old db/db mice compared with 8-week-old control mouse islets (Fig. 2c). HIPK3 protein was increased by 94% and 51%, respectively, in the extracts of islets isolated from 8- and 20-week-old db/db mice compared with 8-week-old control mouse islets (Fig. 2d). We also performed immunohistochemical analysis, finding that HIPK3 levels were increased in islets from db/db mice (Fig. 2e). This indicates that HIPK3 accumulation correlates with glucose tolerance in this genetic model of type 2 diabetes.
Hipk3 −/− mice had impaired insulin secretion
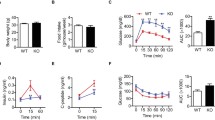
To elucidate the physiological role of HIPK3 in glucose metabolism, we examined the phenotypes of Hipk3 −/− mice. Hipk3 −/− mice showed normal weight gain on a standard and a high-fat diet (Fig. 3a). Their overall appearance at death after 12 weeks on a high-fat diet revealed no difference between Hipk3 −/− and wild-type mice on either a standard or a high-fat diet. We performed the OGTT on mice fed a standard diet. Hipk3 −/− mice fed a standard diet had significantly impaired glucose tolerance (Fig. 3b) and decreased insulin response to glucose (Fig. 3c) compared with wild-type mice. Under standard diet conditions, intraperitoneal glucose tolerance and glucose-dependent insulin secretion of Hipk3 −/− mice were similar to those of wild-type mice (Fig. 3b). Hipk3 −/− mice had significantly impaired GLP-1 secretion during an OGTT compared with that of wild-type mice (Fig. 3d). Hipk3 −/− mice fed a high-fat diet for 4 and 12 weeks had significantly impaired glucose tolerance (Fig. 3e, f) and decreased insulin response to glucose (Fig. 3g, h, k) compared with wild-type mice. The glucose-lowering effect of insulin did not differ between Hipk3 −/− and wild-type mice on a standard or a high-fat diet (Fig. 3i, j, l), supporting the hypothesis that glucose intolerance in Hipk3 −/− mice was not the result of major differences in insulin responsiveness to glucose.
Impaired glucose homeostasis and insulin secretion in Hipk3 −/− mice. (a) The body weight of Hipk3 −/− mice on a high-fat (black squares) and control (black circles) diet, and of wild-type mice on high-fat (white squares) and control (white circles) diet was similar. (b) Mice were loaded with 1.5 mg/g body weight glucose by oral (OGTT) (bold dotted line, wild-type; bold continuous line, Hipk3 −/−) or intraperitoneal (IPGTT) (thin dotted line, wild-type; thin continuous line, Hipk3 −/−) administration. The OGTT revealed impaired glucose homeostasis in Hipk3 −/− mice (n = 7). (c) The OGTT revealed impaired insulin secretion in Hipk3 −/− mice. Insulin levels were determined using a mouse insulin ELISA kit. (d) Impaired GLP-1 secretion in Hipk3 −/− mice. e Hipk3 −/− mice fed a high-fat diet for 4 or (f) 12 weeks had significantly impaired glucose tolerance. Key, as above (b). (g) The glucose intolerance observed in Hipk3 −/− mice fed a high-fat diet for 4 or (h) 12 weeks coincided with impaired insulin release. (i) The insulin tolerance test showed that wild-type (dotted line) and Hipk3 −/− (continuous line) mice fed a high-fat diet for 4 or (j) 12 weeks had similar rates of glucose clearance. (k) Insulinogenic index, defined as the ratio of insulin to glucose at 30 min after an oral glucose load in Hipk3 −/− (black bars) and wild-type (white bars) mice on a high-fat (HF) diet. (l) Values under the glucose curve and under the insulin curve during the OGTT were similar between Hipk3 −/− and wild-type mice. The insulin resistance index was calculated as AUC of insulin/AUC of glucose in the OGTT. Data represent means±SEM; *p < 0.05 compared with wild-type control mice
Loss of Hipk3 decreases glucose-induced insulin secretion
We next determined glucose-induced insulin secretion in Hipk3 −/− mice by static incubation of the same numbers of islets. Whereas insulin secretion into the medium by islets from Hipk3 −/− and wild-type mice was similar at 2.8 mmol/l glucose, insulin secretion by islets from Hipk3 −/− mice was significantly decreased at 11 and 20 mmol/l glucose compared with islets from wild-type mice (Fig. 4a). However, the secretory response in Hipk3 −/− mouse islets was restored by treatment with KCl or glibenclamide (Fig. 4b). To characterise the molecular defect in insulin secretion, we performed pancreas perifusion experiments. In Hipk3 −/− mice, the insulin secretory response to 20 mmol/l glucose was delayed and decreased (Fig. 4c). The ATP content was significantly decreased in Hipk3 −/− islets (Fig. 4d). The increase in intracellular calcium concentration in response to glucose was impaired in islets of Hipk3 −/− mice (Fig. 4e).
Characteristics of islets from Hipk3 −/− mice. (a) Insulin secretion by islets from Hipk3 −/− mice (black bars) fed a high-fat diet for 4 weeks was significantly decreased at 11 and 20 mmol/l glucose vs wild-type mice (white bars) (n = 4). (b) Insulin secretion of Hipk3 −/− and wild-type islets in the presence of KCl and glibenclamide. (c) Islets from Hipk3 −/− (continuous line) and wild-type (dashed line) mice were perifused with a medium containing 20 mmol/l glucose and 20 mmol/l KCl (n = 4). (d) Changes in ATP content in islets from wild-type and Hipk3 −/− mice after glucose stimulation for 1 h (n = 4). (e) The increase in calcium concentration in response to glucose was impaired in Hipk3 −/− islets (n = 6). Glucose-stimulated Ca2+ influx in single beta cells was measured with the Ca2+ indicator fura-2. Fura-2 post-stimulation fluorescence (F) was normalised to resting fluorescence (F0). (f) Changes in beta cell mass during consumption of a high-fat (HF) diet. The insulin content of islets was decreased in Hipk3 −/− vs wild-type mice at week 12 of an HF diet. (g) Insulin content per pancreas was compared between Hipk3 −/− and wild-type mice (n = 4). Data represent means±SEM; *p < 0.05 and **p < 0.01 compared with wild-type control mice
Loss of Hipk3 decreases beta cell proliferation and increases apoptosis
Although the insulin content of islets from Hipk3 −/− mice was not decreased at week 4 of a high-fat diet, it was significantly decreased at week 12 of a high-fat diet (Fig. 4f, g). Similarly, the proportion of islets to total area of dissected pancreas from Hipk3 −/− mice was not decreased at week 4 of a high-fat diet, but was significantly decreased at week 12 of a high-fat diet (Fig. 5a, b). We estimated beta cell proliferation on the basis of PCNA staining. During high-fat diet conditions, significantly more PCNA-positive cells were observed in the islets of wild-type mice than in Hipk3 −/− mice (Fig. 5a, c). We estimated islet cell apoptosis on the basis of TUNEL staining. During high-fat diet conditions, significantly more TUNEL-positive cells were observed in the islets of wild-type mice than in Hipk3 −/− mice (Fig. 5a, d). In summary, HIPK3 has an effect on beta cell secretory function, beta cell proliferation and apoptosis.
(a) PCNA immune staining analysis was based on the counting of 1,662 and 1,879 cells, respectively, per islet sample from wild-type (WT) (n = 4) and Hipk3 −/− (n = 4) mice fed a high-fat diet for 4 weeks, and (high-fat diet, 12 weeks) on the counting of 1,674 and 1,518 cells, respectively, per islet sample from wild-type (n = 4) and Hipk3 −/− (n = 4) mice. Tissues were processed for paraffin embedding, and 4 μm sections were cut and mounted on silanised slides. Pancreatic sections were stained with anti-insulin and anti-PCNA antibodies as indicated. We estimated islet cell apoptosis on the basis of TUNEL staining. Scale bars, 100 μm. (b) The proportion of islet to total area was decreased in Hipk3 −/− compared with wild-type mice fed a HF diet (n = 4). (c) Replication rate of beta cells from Hipk3 −/− and wild-type mice (n = 4). (d) The apoptosis rate of islet cells from Hipk3 −/− and wild-type mice (n = 4). Data represent means±SEM; *p < 0.05 compared with wild-type control mice
Decreased PDX1 and GSK3β phosphorylation in the absence of Hipk3
To identify the mechanism underlying the loss of glucose-stimulated insulin secretion and beta cell proliferation, we evaluated the mediators of HIPK3-regulated genes relative to islet function. Expression of Pdx1, a beta cell-specific transcription factor, was significantly decreased in the islets of Hipk3 −/− mice fed a high-fat diet for 12 weeks, falling by approximately 50% (Fig. 6a, m, n). Notably, we detected decreases of 34% and 30% in Gck and Slc2a2 mRNA expression in Hipk3 −/− mice, indicating that loss of HIPK3 negatively modulates the expression of these genes that encode glucose-sensing proteins (Fig. 6b, c). Moreover, we observed significantly reduced mRNA levels of Hnf4a and Tcf7l2 in Hipk3 −/− islets, suggesting that HIPK3 plays a role in influencing the expression of transcription factors. Additionally, cyclin D1 gene expression (Fig. 6d), GSK3β phosphorylation (Fig. 6m, o) and β-catenin abundance (Fig. 6m, p) were decreased in the islets of Hipk3 −/− mice, indicating a role of HIPK3 in Wnt signalling. Histological analysis showed that nuclear PDX1 abundance was significantly decreased in the islets of Hipk3 −/− mice fed a high-fat diet for 12 weeks (Fig. 6q, r). Histological analysis also showed that GSK3β phosphorylation was decreased in the islets of Hipk3 −/− mice (Fig. 6s).
(a) Quantitative RT-PCR analysis of islet-enriched transcription factors and glucose-sensing proteins Pdx1, (b) Gck, (c) Slc2a2, (d) Cyclin D1, (e) Hnf4a, (f) Tcf7l2, (g) Irs2, (h) Ins1, (i) Hnf1b, (j) Hnf1a, (k) Irs1 and (l) Ucp2. Analysis was in islets isolated from wild-type (WT) and Hipk3 −/− mice (n = 4) on a high-fat diet for 12 weeks. Samples were normalised to Gapdh. m Protein samples from lysates of isolated islets from Hipk3 −/− mice were separated by SDS-PAGE and western blot analyses performed using the antibodies as labelled (n = 4). (n) Quantification of relative PDX1, (o) pGSK3β and (p) β-catenin levels. (q) Pancreatic sections were stained with anti-PDX1 antibody. PDX1 immune staining analysis was based on the counting of 1,845 and 1,683 cells, respectively, per islet sample from wild-type (n = 4) and Hipk3 −/− (n = 4) mice fed a high-fat diet for 12 weeks, and (control diet) on the counting of 1,545 and 1,609 cells, respectively, per islet sample from wild-type (n = 4) and Hipk3 −/− (n = 4) mice. (r) Ratio of the number of nuclear PDX1-positive cells to the total number of islet cells. (s) Histological analysis of pancreatic islet phosphorylated GSK3β in wild-type and Hipk3 −/− mice. Pancreatic sections were stained with anti-phspho-GSK3β(Ser9) antibodies. Analysis was based on the counting of 1,705 and 1,692 cells, respectively, per islet sample from wild-type (n = 4) and Hipk3 −/− (n = 4) mice fed a high-fat diet for 12 weeks, and (control diet) on the counting of 1,875 and 1,586 cells, respectively, per islet sample from wild-type (n = 4) and Hipk3 −/− (n = 4) mice. Scale bars (q, s), 100 μm. Data represent means±SEM; *p < 0.05 and **p < 0.01 compared with wild-type control mice
Wnt signalling lessens the decrease in insulin secretion induced by loss of Hipk3
We investigated whether Wnt signalling can also increase glucose-stimulated insulin secretion in Hipk3 −/− islets. Wnt3a enhanced insulin secretion by 19% in Hipk3 −/− islets (Fig. 7a) upon 20 mmol/l glucose stimulation. We also found that the GSK3 inhibitor LiCl and 1-azakenpaullone treatment enhanced insulin secretion in Hipk3 −/− islets by 31% and 28%, respectively, upon 20 mmol/l glucose stimulation (Fig. 7b, c). These observations suggest that impaired Wnt signalling caused by Hipk3 deficiency leads to impaired beta cell function.
Wnt3a regulates HIPK3-induced insulin secretion. (a) Insulin secretion was decreased by Wnt3a (n = 4). Insulin secretion by islets from wild-type (white bars; light grey bars, wild-type + Wnt3a) and Hipk3 −/− (black bars; dark grey bars, Hipk3 −/− + Wnt3a) mice was measured in KRB with a glucose concentration of 2.8, 11 or 20 mmol/l. Static incubation was performed for 1 h on ten islets per tube at 37°C, after preincubation with the basal glucose concentration for 20 min. Insulin levels were determined with a mouse insulin ELISA kit. (b) The insulin secretion of isolated islets was decreased by Hipk3 deficiency and increased by the GSK3 inhibitor LiCl (n = 4). Key, as above (a), except: light grey bars, wild-type + LiCl; dark grey bars, Hipk3 −/− + LiCl. (c) Insulin secretion of isolated islets from Hipk3 −/− mice was increased by Wnt signalling activation by the GSK3 inhibitor, 1-azakenpaullone (n = 4). Key, as above (a), except: light grey bars, wild-type + 1-azakenpaullone; dark grey bars, Hipk3 −/− + 1-azakenpaullone. Data represent means ± SEM; *p < 0.05 and **p < 0.01
Discussion
It is becoming increasingly clear that the diverse biological consequences of the loss of HIPK activity include the coordinated death of cells in earlier developmental stages and the dysregulation of proper cell number in diverse tissue types. Mice that are deficient for Hipk1 and Hipk2 singly have a normal overall appearance, a finding that is probably due to a functional redundancy between HIPK1 and HIPK2 [41]. However, Hipk1 −/−; Hipk2 −/− mice are progressively lost in utero, and exhibit exencephaly and severe defects in haematopoiesis, vasculogenesis and angiogenesis. In our experiments, expression of Hipk genes was detected in adult islets, and the HIPK family additively regulated insulin secretion. Our data also indicate that high-fat diet leads most significantly to Hipk3 upregulation. The other two Hipk genes correlate with glucose tolerance and may play roles in glucose homeostasis. Given the inability of the intact Hipk3 gene to support early development in Hipk1 −/−; Hipk2 −/− mice and the contribution of HIPK3 to impaired insulin secretion, we anticipate that HIPK3 may serve unique physiological roles especially in diabetes-prone circumstances such as a high-fat diet.
We also analysed Hipk3 −/− mice. Major changes in gene phenotype were found with very modest increases in glucose levels, such as impaired glucose tolerance. Changes in islet gene expression can be explained by Hipk3 deficiency and glucose toxicity.
HIPK2 positively influences PDX1 levels and transcriptional activity, and directly phosphorylates the C-terminal portion of PDX1 [28]. HIPK2 is a potential kinase for PDX1 Ser-269, with high glucose concentrations decreasing the degree of phosphorylation on PDX1 Ser-269 in beta cells [42]. We showed that the number of PDX1-positive cells was reduced by 20% in Hipk3-knockout mice on a 12 week high-fat diet (Fig. 6r). The insulin content was reduced by 18% in the same conditions (Fig. 4f). Thus, we speculate that decreased PDX1 positivity is related to beta cell mass reduction. Considering the role of PDX1 in beta cells and non-beta cells, HIPK may affect beta cells and non-beta cells, and reduce beta cell mass. Further studies are needed to clarify the mechanisms of beta cell reduction that decreased PDX1 positivity. HIPK3 has been shown to augment transcription of androgen receptor [32], SF-1 [35] and runt-related transcription factor 2 [36]. In this study, we identified HIPK3 as a novel modulator of PDX1 abundance. Several transcription factors converge on regulatory areas within the Pdx1 promoter [43–45], including NKx2.2, a member of the NK-2 family of homeobox genes [44, 45]. HIPKs possess a homeodomain-interacting domain that is important for enhancing the repressor activities of NK homeodomain transcription factors [22], allowing the possibility that HIPK3 may affect PDX1 levels via interacting factors that promote PDX1 transcription.
We showed that insulin transcript levels are correlated with the results of luciferase assays. We used rat insulin promoter instead of mouse insulin promoter, and this difference in species may have affected our results. Concerning the effect of HIPK on insulin expression, HIPK may interact with the insulin promoter or act by modulating PDX1 levels. Further studies are needed to elucidate the mechanisms by which HIPK affects insulin expression.
This study also revealed the involvement of the Wnt–GSK3β–β-catenin cascade in the action of HIPK3. Wnt signalling influences endocrine pancreas development and modulates mature beta cell functions, including insulin secretion, beta cell survival and beta cell proliferation [6–9]. Treatment with Wnt ligands increases glucose-induced insulin secretion in wild-type islets, but not in islets deficient in the Wnt co-receptor, lipoprotein receptor-related protein 5, suggesting a deficit in glucose sensing in islets when canonical Wnt signalling is impaired [46]. β-Catenin is sufficient to induce proliferation markers and to increase beta cell mass in transgenic mice [4]. Adipocyte-derived Wnt signalling molecules induce cyclin D1 transcription, and the proliferation of beta cell lines and primary murine beta cells [5]. Beta cell-specific Gsk3b knockout prevents diabetes in Irs2-knockout [13] and high-fat diet-fed mice [14] through expanded beta cell mass with increased beta cell proliferation. In line with these reports, we have shown that beta cell proliferation is decreased in Hipk3 −/− mouse islets upon feeding a high-fat diet. In contrast to our findings, pancreas-specific depletion of β-catenin has been shown to lead to acinar cell hypoplasia, but not to change the mass and function of islets [47]. This discrepancy might be explained by β-catenin-independent signal transduction downstream of HIPK3.
Previous reports indicate that β-catenin and transcription factor 7 like 2, T cell specific, HMG box regulate proglucagon gene expression and GLP-1 synthesis in enteroendocrine cells [48, 49]. In this study, we obtained evidence that HIPK3 deficiency lowered GLP-1 concentrations and impaired glucose tolerance. It can thus be speculated that HIPK3 activates the Wnt signalling pathway for GLP-1 synthesis and secretion. Furthermore, HIPK3 may alter the response of intestinal K cells to known secretagogues, including nutrients, fatty acids and related molecules. These hypotheses deserve further examination.
In conclusion, HIPK3 has a role in insulin secretion, affecting the levels of pancreatic PDX1 and GSK3β phosphorylation. Thus, targeting HIPK3 with novel agonists could lead to new therapeutic strategies for preserving beta cell function. The present results provide new insight into the pathogenesis of type 2 diabetes.
Abbreviations
- ChIP:
-
Chromatin immunoprecipitation
- GSK:
-
Glycogen synthase kinase
- HIPK:
-
Homeodomain-interacting protein kinase
- NK:
-
Neurokinin
- PCNA:
-
Proliferating cell nuclear antigen
- PDX:
-
Pancreatic duodenal homeobox
- SF-1:
-
Splicing factor 1
- siRNA:
-
Small interfering RNA
References
Kahn CR (1994) Banting lecture. Insulin action, diabetogenes, and the cause of type II diabetes. Diabetes 43:1066–1084
Accili D (2004) Lilly lecture 2003: the struggle for mastery in insulin action: from triumvirate to republic. Diabetes 53:1633–1642
Doria A, Patti ME, Kahn CR (2008) The emerging genetic architecture of type 2 diabetes. Cell Metab 8:186–200
Rulifson IC, Karnik SK, Heiser PW et al (2007) Wnt signaling regulates pancreatic beta cell proliferation. Proc Nat Acad Sci USA 104:6247–6252
Schinner S, Ulgen F, Papewalis C et al (2008) Regulation of insulin secretion, glucokinase gene transcription and beta cell proliferation by adipocyte-derived Wnt signalling molecules. Diabetologia 51:147–154
Liu Z, Habener JF (2010) Wnt signaling in pancreatic islets. Adv Exp Med Biol 654:391–419
Welters HJ, Kulkarni RN (2008) Wnt signaling: relevance to beta-cell biology and diabetes. Trends Endocrinol Metab 19:349–355
Jin T (2008) The WNT signalling pathway and diabetes mellitus. Diabetologia 51:1771–1780
Schinner S, Willenberg HS, Schott M, Scherbaum WA (2009) Pathophysiological aspects of Wnt-signaling in endocrine disease. Eur J Endocrinol 160:731–737
Logan CY, Nusse R (2004) The Wnt signaling pathway in development and disease. Annu Rev Cell Dev Biol 20:781–810
Moon RT, Kohn AD, de Ferrari GV, Kaykas A (2004) WNT and beta-catenin signalling: diseases and therapies. Nat Rev Genet 5:691–701
Liu Z, Tanabe K, Bernal-Mizrachi E, Permutt MA (2008) Mice with beta cell overexpression of glycogen synthase kinase-3beta have reduced beta cell mass and proliferation. Diabetologia 51:623–631
Tanabe K, Liu Z, Patel S et al (2009) Genetic deficiency of glycogen synthase kinase-3beta corrects diabetes in mouse models of insulin resistance. PLoS Biol 6:e37
Liu Y, Tanabe K, Baronnier D et al (2010) Conditional ablation of Gsk-3β in islet beta cells results in expanded mass and resistance to fat feeding-induced diabetes in mice. Diabetologia 53:2600–2610
Kanei-Ishii C, Ninomiya-Tsuji J, Tanikawa J et al (2004) Wnt-1 signal induces phosphorylation and degradation of c-Myb protein via TAK1, HIPK2, and NLK. Genes Dev 18:816–829
Wei G, Ku S, Ma G et al (2007) HIPK2 represses beta-catenin-mediated transcription, epidermal stem cell expansion, and skin tumorigenesis. Proc Natl Acad Sci USA 104:13040–13045
Kim EA, Kim JE, Sung KS et al (2010) Homeodomain-interacting protein kinase 2 (HIPK2) targets beta-catenin for phosphorylation and proteasomal degradation. Biochem Biophys Res Commun 394:966–971
Swarup S, Verheyen EM (2011) Drosophila homeodomain-interacting protein kinase inhibits the Skp1-Cul1-F-box E3 ligase complex to dually promote Wingless and Hedgehog signaling. Proc Natl Acad Sci USA 108:9887–9892
Lee W, Swarup S, Chen J, Ishitani T, Verheyen EM (2009) Homeodomain-interacting protein kinases (Hipks) promote Wnt/Wg signaling through stabilization of beta-catenin/Arm and stimulation of target gene expression. Development 136:241–251
Louie SH, Yang XY, Conrad WH et al (2009) Modulation of the beta-catenin signaling pathway by the dishevelled-associated protein Hipk1. PLoS One 4:e4310
Hisaka H, Ezan J, Itoh K, Li X, Klymkowsky MW, Sokol SY (2010) Regulation of TCF3 by Wnt-dependent phosphorylation during vertebrate axis specification. Dev Cell 19:521–532
Kim YH, Choi CY, Lee SJ, Conti MA, Kim Y (1998) Homeodomain-interacting protein kinases, a novel family of co-repressors for homeodomain transcription factors. J Biol Chem 273:25875–25879
D’Orazi G, Cecchinelli B, Bruno T et al (2002) Homeodomain-interacting protein kinase-2 phosphorylates p53 at Ser 46 and mediates apoptosis. Cell Biol 4:11–19
Hofmann TG, Moller A, Sirma H et al (2002) Regulation of p53 activity by its interaction with homeodomain-interacting protein kinase-2. Nat Cell Biol 4:1–10
Kondo S, Lu Y, Debbas M et al (2003) Characterization of cells and gene-targeted mice deficient for the p53-binding kinase homeodomain-interacting protein kinase 1 (HIPK1). Proc Natl Acad Sci USA 100:5431–5436
Zhang Q, Yoshimatsu Y, Hildebrand J, Frisch SM, Goodman RH (2003) Homeodomain interacting protein kinase 2 promotes apoptosis by downregulating the transcriptional corepressor CtBP. Cell 115:177–186
Choi CY, Kim YH, Kim YO et al (2005) Phosphorylation by the DHIPK2 protein kinase modulates the corepressor activity of Groucho. J Biol Chem 280:21427–21436
Boucher M, Simoneau ML, Edlund H (2009) The homeodomain-interacting protein kinase 2 regulates insulin promoter factor-1/pancreatic duodenal homeobox-1 transcriptional activity. Endocrinology 150:87–97
Jonsson J, Carlsson L, Edlund T, Edlund H (1994) Insulin-promoter-factor 1 is required for pancreas development in mice. Nature 371:606–609
Offield MF, Jetton TL, Labosky PA et al (1996) PDX-1 is required for pancreatic outgrowth and differentiation of the rostral duodenum. Development 122:983–995
Stoffers DA, Zinkin NT, Stanojevic V, Clarke WL, Habener JF (1997) Pancreatic agenesis attributable to a single nucleotide deletion in the human IPF1 gene coding sequence. Nat Genet 15:106–110
Moilanen AM, Karvonen U, Poukka H, Janne OA, Palvimo JJ (1998) Activation of androgen receptor function by a novel nuclear protein kinase. Mol Biol Cell 9:2527–2543
Rochat-Steiner V, Becker K, Micheau O, Schneider P, Burns K, Tschopp J (2000) FIST/HIPK3: a Fas/FADD-interacting serine/threonine kinase that induces FADD phosphorylation and inhibits fas-mediated Jun NH(2)-terminal kinase activation. J Exp Med 192:1165–1174
Curtin JF, Cotter TG (2004) JNK regulates HIPK3 expression and promotes resistance to Fas-mediated apoptosis in DU 145 prostate carcinoma cells. J Biol Chem 279:17090–17100
Lan HC, Li HJ, Lin G, Lai PY, Chung BC (2007) Cyclic AMP stimulates SF-1-dependent CYP11A1 expression through homeodomain-interacting protein kinase 3-mediated Jun N-terminal kinase and c-Jun phosphorylation. Mol Cell Biol 27:2027–2036
Sierra OL, Towler DA (2010) Runx2 trans-activation mediated by the MSX2-interacting nuclear target requires homeodomain interacting protein kinase-3. Mol Endocrinol 24:1478–1497
Inoue T, Kagawa T, Inoue-Mochita M et al (2010) Involvement of the Hipk family in regulation of eyeball size, lens formation and retinal morphogenesis. FEBS Lett 584:3233–3238
Scopsi L, Wang BL, Larsson LI (1986) Nonspecific immunocytochemical reactions with certain neurohormonal peptides and basic peptide Sequences. J Histochem Cytochem 34:1469–1475
Noda M, Yamashita S, Takahashi N et al (2002) Switch to anaerobic glucose metabolism with NADH accumulation in the beta-cell model of mitochondrial diabetes. Characteristics of betaHC9 cells deficient in mitochondrial DNA transcription. J Biol Chem 277:41817–41826
Takahashi N, Hatakeyama H, Okado H et al (2004) Sequential exocytosis of insulin granules is associated with redistribution of SNAP25. J Cell Biol 165:255–262
Isono K, Nemoto K, Li Y et al (2006) Overlapping roles for homeodomain-interacting protein kinases hipk1 and hipk2 in the mediation of cell growth in response to morphogenetic and genotoxic signals. Mol Cell Biol 26:2758–2771
An R, da Silva Xavier G, Semplici F et al (2010) Pancreatic and duodenal homeobox 1 (PDX1) phosphorylation at serine-269 is HIPK2-dependent and affects PDX1 subnuclear localization. Biochem Biophys Res Commun 399:155–161
Gerrish K, Gannon M, Shih D et al (2000) Pancreatic beta cell-specific transcription of the pdx-1 gene. The role of conserved upstream control regions and their hepatic nuclear factor 3beta sites. J Biol Chem 275:3485–3492
Gerrish K, van Velkinburgh JC, Stein R (2004) Conserved transcriptional regulatory domains of the pdx-1 gene. Mol Endocrinol 18:533–548
van Velkinburgh JC, Samaras SE, Gerrish K, Artner I, Stein R (2005) Interactions between areas I and II direct pdx-1 expression specifically to islet cell types of the mature and developing pancreas. J Biol Chem 280:38438–38444
Fujino T, Asaba H, Kang MJ et al (2003) Low-density lipoprotein receptor-related protein 5 (LRP5) is essential for normal cholesterol metabolism and glucose-induced insulin secretion. Proc Natl Acad Sci USA 100:229–234
Murtaugh LC, Law AC, Dor Y, Melton DA (2005) Beta-catenin is essential for pancreatic acinar but not islet development. Development 132:4663–4674
Yi F, Brubaker PL, Jin T (2005) TCF-4 mediates cell type-specific regulation of proglucagon gene expression by beta-catenin and glycogen synthase kinase-3beta. J Biol Chem 280:1457–1464
Ni Z, Anini Y, Fang X, Mills G, Brubaker PL, Jin T (2003) Transcriptional activation of the proglucagon gene by lithium and beta-catenin in intestinal endocrine L cells. J Biol Chem 278:1380–1387
Acknowledgements
We thank M. Yamaguchi and N. Miyama (Department of Diabetes and Metabolic Disease, University of Tokyo, Tokyo, Japan) for their excellent technical assistance.
Funding
This work was supported by a grant for Translational Systems Biology and Medicine Initiative (TSBMI) from the Ministry of Education, Culture, Sports, Science and Technology of Japan (to T. Kadowaki), a Grant-in-aid for Scientific Research in Priority Areas (S) from the Ministry of Education, Culture, Sports, Science and Technology of Japan (to T. Kadowaki), Health Science Research grants (Research on Human Genome and Gene Therapy) from the Ministry of Health and Welfare (to T. Kadowaki) and a Grant-in-Aid from the Japan Society for the Promotion of Science (JSPS) (to T. Kadowaki and K. Hara).
Duality of interest
The authors declare that there is no duality of interest associated with this manuscript.
Contribution statement
NS, KH and TK planned and designed the study. NS, KH, KU and TK conceived and designed the experiments, and wrote the manuscript. HF and MH contributed to the conception and design, the analysis and interpretation of the data, and drafting the article. NT carried out the two-photon excitation imaging and contributed to drafting the article. IT, MO, HA, MN, NK and TY provided advice, and contributed to the conception and interpretation of the data and drafting the article. All authors approved the final version.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM Table 1
(PDF 17 kb)
Rights and permissions
About this article
Cite this article
Shojima, N., Hara, K., Fujita, H. et al. Depletion of homeodomain-interacting protein kinase 3 impairs insulin secretion and glucose tolerance in mice. Diabetologia 55, 3318–3330 (2012). https://doi.org/10.1007/s00125-012-2711-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-012-2711-1