Abstract
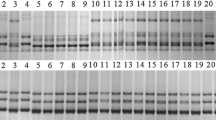
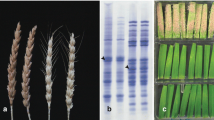
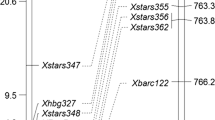
A single gene controlling powdery mildew resistance was identified in the North Carolina germplasm line NC96BGTD3 (NCD3) using genetic analysis of F2 derived lines from a NCD3 X Saluda cross. Microsatellite markers linked to this Pm gene were identified and their most likely order was Xcfd7, 10.3 cM, Xgdm43, 8.6 cM, Xcfd26, 11.9 cM, Pm gene. These markers and the Pm gene were assigned to chromosome 5DL by means of Chinese Spring Nullitetrasomic (Nulli5D-tetra5A) and ditelosomic (Dt5DL) lines. A detached leaf test showed a distinctive disease reaction to six pathogen isolates among the NCD3 Pm gene, Pm2 (5DS) and Pm34 (5DL). An allelism test showed independence between Pm34 and the NCD3 Pm gene. Together, the tests provided strong evidence for the presence of a novel Pm gene in NCD3, and this gene was designated Pm35.


Similar content being viewed by others
References
Cox TS, Raupp WJ, Wilson BS, Gill B, Leath S, Bockus WW, Browder LE (1992) Resistance to foliar diseases in a collection of Triticum tauschii germplasm. Plant Dis 76:1061–1064
Gill BS, Raupp WJ (1987) Direct gene transfers from Aegilops squarrosa L. to hexaploid wheat Crop Sci 27:445–450
Gupta PK, Varshney RK, Sharma PC, Ramesh B (1999) Molecular markers and their applications in wheat breeding. Plant Breed 118:369–390
Hsam SLK, Zeller FJ (2002) Breeding for powdery mildew resistance in common wheat (Triticum aestivum L.). In: Berlanger RR, Bushnell WR, Dik AJ, Carver TLW (eds) The powdery mildews, a comprehensive treatise. APS Press, St. Paul, pp 219–238
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Langridge P, Lagudah ES, Holton TA, Appels R, Sharp PJ, Chalmers KJ (2001) Trends in genetics and genome analyses in wheat: a review. Aust J Agric Res 52:1043–1077
Leath S, Heun M (1990) Identification of powdery mildew resistance genes in cultivars of soft red winter wheat. Plant Dis 74:747–752
Lincoln SE, Daly MJ, Lander ES (1993) Constructing linkage maps with MAPMAKER/Exp Version 3.0. A tutorial reference manual, 3rd edn. Whitehead Institute for Medical Res., Cambridge
Lutz J, Hsam SLK, Limpert E, Zeller FJ (1994) Powdery mildew resistance in Aegilops tauschii Coss. and synthetic hexaploid wheats. Genet Res Crop Evol 41:151–158
Lutz J, Hsam SLK, Limpert E, Zeller FJ (1995) Chromosomal location of powdery mildew resistance genes in Triticum aestivum L. (common wheat). 2. Genes Pm2 and Pm19 from Aegilops squarrosa L. Heredity 74:152–156
McIntosh RA, Baker EP (1970) Cytogenetic studies in wheat IV Chromosomal location and linkage studies involving the Pm2 locus for powdery mildew resistance. Euphytica 19:71–77
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating population. Proc Natl Acad Sci USA 88:9828–9832
Miranda LM, Murphy JP, Leath S, Marshall DS (2006) Pm34: a new powdery mildew resistance gene transferred from Aegilops tauschii Coss. to common wheat (Triticum aestivum L.) Theor Appl Genet 114:1497–1504
Murphy JP, Leath S, Huynh D, Navarro RA, Shi A (1998) Registration of NC96BGTD1, NC96BGTD2 and NC96BGTD3 wheat germplasm resistant to powdery mildew. Crop Sci 38:570–571
Murphy JP, Leath S, Huynh D, Navarro RA, Shi A (1999) Registration of NC97BGTD7 and NC97BGTD8 wheat germplasms resistant to powdery mildew. Crop Sci 39:884–885
Qiu YC, Sun XL, Zhou RH, Kong XY, Zhang SS, Jia JZ (2006) Identification of microsatellite markers linked to powdery mildew resistance gene Pm2 in wheat. Cereal Res Commun 34:1267–1273
Rampling LR, Harker N, Shariflou MR, Morell MK (2001) Detection and analysis systems for microsatellite markers in wheat. Aust J Agric Res 52:1131–1141
Schuelke M (2000) An economic method for fluorescent labeling of PCR fragments. Nat Biotechnol 18:233–234
Somers DJ, Isaac P, Edwards K (2004) A high density microsatellite consensus map for bread wheat (Triticum aestivum L). Theor Appl Genet 109:1105–1114
Starling TM, Roane CW, Camper HM (1986) Registration of ‘Saluda’ wheat. Crop Sci 26:200
Stein N, Herren G, Keller B (2001) A new DNA extraction method for high throughput marker analysis in a large genome species such as Triticum aestivum L. Plant Breed 120:354–356
Zadoks JC, Chang TT, Konzak CF (1974) A decimal code for the growth stages of cereals. Weed Res 14:415–421
Zhou R, Zhu Z, Kong X, Huo N, Tian Q, Li P, Jin C, Dong Y, Jia J (2005) Development of near-isogenic lines for powdery mildew resistance. Theor Appl Genet 110:640–648
Acknowledgments
The authors thank Dr. Robert McIntosh for his comprehensive review of this manuscript. We would also like to acknowledge Rene Navarro and David Wooten for their excellent help with the field experiment and Jeanette Lyerly for her technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by F. Ordon.
Rights and permissions
About this article
Cite this article
Miranda, L.M., Murphy, J.P., Marshall, D. et al. Chromosomal location of Pm35, a novel Aegilops tauschii derived powdery mildew resistance gene introgressed into common wheat (Triticum aestivum L.). Theor Appl Genet 114, 1451–1456 (2007). https://doi.org/10.1007/s00122-007-0530-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-007-0530-4