Abstract.
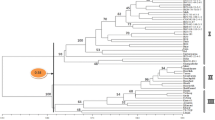
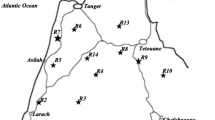
The genetic diversity produced by the amplified fragment length polymorphism (AFLP) method was studied in 94 genotypes of wild barley, Hordeum spontaneum (C. Koch) Thell., originating from ten ecologically and geographically different locations in Israel. Eight primer pairs produced 204 discernible loci of which 189 (93%) were polymorphic. Each genotype had a unique banding profile and the genetic similarity coefficient varied between 0.74 and 0.98. The phenogram generated from these similarities by the UPGMA method did not group genotypes strictly according to their geographical origin, which pattern was also seen in the principal coordinate (PCO) plot. Genetic diversity was larger within (69%) than among (31%) populations. Associations between ecogeographical variables and the mean gene diversity were found at one primer pair. The results are discussed and compared with data obtained by the simple sequence repeat (SSR) method.



Similar content being viewed by others
References
Alonso-Blanco C, Peeters AJ, Koornneef M, Lister C, Dean C (1998) Development of an AFLP based linkage of Ler, Col and Cvi Arabidopsis thaliana ecotypes and construction of a Ler/Cvi recombinant inbred line population. Plant J 14:259–271
Baum BR, Nevo E, Johnson DA, Beiles A (1997) Genetic diversity in wild barley (Hordeum spontaneum C. Koch) in the Near East: a molecular analysis using random amplified polymorphic DNA (RAPD) markers. Genet Res Crop Evol 44:147–157
Becker J, Vos M, Kuiper M, Salamini F, Heun M (1995) Combined mapping of AFLP and RFLP markers in barley. Mol Gen Genet 249:65–73
Brown AHD, Zohary D, Nevo E (1978) Outcrossing rates and heterozygosity in natural populations of Hordeum spontaneum Koch in Israel. Heredity 41:49–62
Buntjer JB (1999) Cross Checker Fingerprint analysis software v2.9. Wageningen University, The Netherlands. http://www.dpw.wau.nl/pv
Chalmers KJ, Waugh R, Watters J, Forster BP, Nevo E, Abbott RJ, Powell W (1992) Grain isozyme and ribosomal DNA variability in Hordeum spontaneum populations from Israel. Theor Appl Genet 84:313–322
Dawson IK, Chalmers KJ, Waugh R, Powell W (1993) Detection and analysis of genetic variation in Hordeum spontaneum populations from Israel using RAPD markers. Mol Ecol 2:151–159
Donini P, Elias ML, Bougourd SM, Koebner RMD (1997) AFLP fingerprinting reveals pattern differences between template DNA extracted from different plants. Genome 40:521–526
Krauss SL (1999) Complete exclusion of nonsires in an analysis of paternity in a natural plant population using amplified fragment lenght polymorphism (AFLP). Mol Ecol 8:217–226
Law JR, Donini P, Koebner RMD, Reeves JC, Cooke RJ (1998) DNA profiling and plant variety registration. III. The statistical assessment of distinctness in wheat using amplified fragment lenght polymorphisms. Euphytica 102:335–342
Miyashita NT, Kawabe A, Innan H (1999) DNA variation in the wild plant Arabidopsis thaliana revealed by amplified fragment length polymorphism analysis. Genetics 152:1723–7311
MVSP Multi Variate Statistical Program (2001) Version 3.12d. Kovach Computing Services 85,Nant-y-Felin, Pentraeth, Isle of Anglesey, LL75 8 UY, Wales, UK
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nei M, Li W-H (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76:5269–5273
Nevo E (1988) Genetic diversity in nature: patterns and theory. Evol Biol 23:217–246
Nevo E (1998a) Genetic diversity in wild cereals: regional and local studies and their bearing on conservation ex situ and in situ. Genet Res Crop Evol 45:355–370
Nevo E (1998b) Molecular evolution and ecological stress at global, regional and local scales: the Israeli perspective. J Exp Zool 282:95–119
Nevo E, Zohary D, Brown AHD, Haber M (1979) Genetic diversity and environmental associations of wild barley, Hordeum spontaneum, in Israel. Evolution 33:815–833
Nevo E, Brown AHD, Zohary D, Storch N, Beiles A (1981) Microgeographic edaphic differentiation in allozyme polymorphisms of wild barley (Hordeum spontaneum, Poaceae). Plant Syst Evol 138:287–292
Nevo E, Beiles A, Kaplan D, Golenberg EM, Olsvig-Whittaker L, Naveh Z (1986) Natural selection of allozyme polymorphism: a microsite test revealing ecological genetic differentiation in wild barley. Evolution 40:13–20
Nevo E, Apelbaum-Elkaher I, Garty J, Beiles A (1997) Natural selection causes microscale allozyme diversity in wild barley and a lichen at "Evolution Canyon" Mt. Carmel, Israel. Heredity 78:373–382
Nevo E, Baum BR, Beiles A, Johnson DA (1998) Ecological correlates of RAPD DNA diversity of wild barley, Hordeum spontaneum, in the Fertile Crescent. Genet Res Crop Evol 45:151–159
Owuor ED, Fahima T, Beiles A, Korol A, Nevo E (1997) Population genetics response to microsite ecological stress in wild barley, Hordeum spontaneum. Mol Ecol 6:1177–1187
Pakniyat H, Powell W, Baird E, Handley LL, Robinson D, Scrimgeour CM, Nevo E, Hackett CA, Caligari PDS, Forster BP (1997) AFLP variation in wild barley (Hordeum spontaneum C. Koch) with reference to salt tolerance and associated ecogeography. Genome 40:332–341
Qi X, Lindhout P (1997) Development of AFLP markers in barley. Mol Gen Genet 254:330–336
Russell JR, Fuller JD, Macaulay M, Hatz BG, Jahoor A, Powell W, Waugh R (1997) Direct comparison of levels of genetic variation among barley accessions detected by RFLPs, AFLPs, SSRs and RAPDs. Theor Appl Genet 95:714–722
Schut JW, Qi X, Stam P (1997) Association between relationship measures based on AFLP markers, pedigree data and morphological traits in barley. Theor Appl Genet 95:1161–1168
Sneath PHA, Sokal RR (1973) Numerical taxonomy. The principles and practice of numerical classification, WH Freeman and Co, San Francisco
Turpeinen T, Tenhola T, Manninen O, Nevo E, Nissilä E (2001) Microsatellite diversity associated with ecological factors in Hordeum spontaneum populations in Israel. Mol Ecol 10:1577–1591
Van der Voort JR, Wolters P, Folkertsma R, Hutten R, Van Zandvoort P, Vinke H, Kanyuka K, Bendahmane A, Jacobsen E, Janssen R, Bakker J (1997) Mapping of the cyst nematode resistance locus Gpa 2 in potato using a strategy based on comigrating AFLP markers. Theor Appl Genet 95:874–880
Van Valen L (1965) Morphological variation and width of ecological niche. Am Nat 99:377–390
Vos P, Hogers R, Bleeker M, Reijans M, Van der Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–3314
Waugh R, Bonar N, Baird E, Thomas B, Graner A, Hayes P, Powell W (1997) Homology of AFLP products in three mapping populations of barley. Mol Gen Genet 255:311–321
Weining S, Henry RJ (1995) Molecular analysis of the DNA polymorphism of wild barley (Hordeum spontaneum) germplasm using the polymerase chain reaction. Genet Res Crop Evol 42:273–281
Yeh FC, Yang RC, Boyle TBJ, Ye Z-H, Mao JX (1997) POPGENE, the user-friendly shareware for population genetic analysis. Version 1.21. Molecular Biology and Biotechnology Centre University of Alberta Canada
Zabeau M, Vos P (1993) Selective restriction fragment amplification: a general method for DNA fingerprinting. European patent application no. 92402629.7. Publ no. 0534858 A1
Acknowledgements.
We gratefully acknowledge Ms. Teija Tenhola for growing leaf material and Ms. Leena Lohermaa and Ms. Sirpa Moisander for the DNA extractions. T. Turpeinen acknowledges the Finnish Cultural Foundation for financial support. E. Nevo thanks the Ancell-Teicher Research Foundation for Genetics and Molecular Evolution.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H.F. Linskens
T. Turpeinen and T. Vanhala contributed equally to this work
An erratum to this article is available at http://dx.doi.org/10.1007/s00122-003-1286-0.
Rights and permissions
About this article
Cite this article
Turpeinen, T., Vanhala, T., Nevo, E. et al. AFLP genetic polymorphism in wild barley (Hordeum spontaneum) populations in Israel. Theor Appl Genet 106, 1333–1339 (2003). https://doi.org/10.1007/s00122-002-1151-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-002-1151-6